From values to network reconfiguration in single-cell perturbations
Published in Cancer and Protocols & Methods
Single-cell transcriptomics has transformed how we study biological systems, enabling high-resolution profiling across diverse conditions at scale. Yet despite this advance, the analytical logic of most comparative studies remains largely unchanged: quantify cell abundance, identify differentially expressed genes, perform pathway enrichment, and infer upstream regulators.
This workflow is powerful—but it rests on an implicit assumption: that biologically meaningful perturbations manifest as changes in gene expression levels. When this assumption holds, the framework performs remarkably well. When it does not, important signals can go undetected—not because they are weak, but because expression is not the only dimension being altered.
Autosomal dominant combined immunodeficiency caused by IRF4 mutations makes this limitation particularly clear. The pathogenic variants do not reduce IRF4 expression; instead, they alter its DNA-binding specificity. IRF4 remains present and transcriptionally active, yet its regulatory interactions with target genes are systematically rewired. The perturbation resides not in expression magnitude, but in the architecture of gene regulation itself.
This observation led us to reconsider what a “perturbation” actually means. Genes do not function in isolation—their roles emerge through relationships: coordination, dependency, co-regulation, and context-specific coupling. A perturbation can therefore reshape the architecture of these relationships without necessarily producing large expression changes. What changes is not simply signal intensity, but how genes relate to and depend on one another.
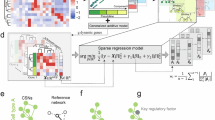
scDNS was developed from this perspective. Instead of asking which genes change their expression, we ask which genes change their dependency relationships across conditions—and crucially, in which cells these changes occur. This reframing also addresses a persistent disconnect in standard analyses, where “which genes are perturbed” and “which cells are affected” are often treated as separate questions. Mechanistic understanding requires linking the two: a perturbation becomes interpretable only when we can trace where in the cellular landscape it exerts its effects.

We applied scDNS across several biological contexts. In IFN-β–treated PBMCs, network analysis highlighted upstream regulators—including the IFN-β receptor itself—with minimal transcriptional change, capturing regulatory signals not readily detected by differential expression analysis. In COVID-19 lung tissue, where cellular susceptibility is highly heterogeneous, scDNS identified perturbation-associated genes with therapeutic relevance that were missed by expression-based prioritization. In gemcitabine-treated pancreatic cancer cells, TIMM44 emerged as a functionally perturbed node despite stable expression, pointing to a mitochondria-associated vulnerability with potential clinical implications. Across these cases, the shared pattern is a set of biologically meaningful signals that fall outside the reach of expression-based analyses.
Looking back, gene expression is one projection of biological activity, while system-level function is also encoded in the relationships among components. Measuring a single dimension captures part of the picture; an entire class of information instead resides in the structure of those relationships.
As datasets grow in scale and complexity, the key limitation may no longer be what we can measure, but how we choose to ask questions. Some of the most important signals may already be present in our data—waiting not for better measurement, but for a more appropriate framework.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Ask the Editor – Inflammation, Metastasis, Cancer Microenvironment and Tumour Immunology
Got a question for the editor about inflammation, metastasis, or tumour immunology? Ask it here!
Continue reading announcementRelated Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in