Rare cells have their own circles: how we came to see them differently
Published in Protocols & Methods and Anatomy & Physiology
One of the central goals of single-cell analysis is to achieve a comprehensive and fine-grained understanding of cellular composition within tissues. For years, this task has relied heavily on dimensionality reduction and clustering. These approaches perform remarkably well in identifying major cell types, but they often run into a dilemma when dealing with rare cell populations: increasing resolution may help separate small groups of cells, but at the cost of unstable over-partitioning; maintaining global structure, on the other hand, tends to obscure truly meaningful rare populations within noise.
The turning point came from an unexpected analogy.
During a discussion about cell–cell relationships, we casually asked: What if we think of cellular neighborhoods as “social circles”? In a tightly connected social circle, one's friends are often also friends with one another—a “friend-of-a-friend is also a friend” pattern, which in network terms correspond to a highly connected local clique. This led us to wonder: do similar structures exist in cellular neighborhood networks?

When we began to examine this question systematically, the answer became surprisingly clear. Some small groups of cells, even when globally sparse or located close to major populations, exhibited strikingly tight local connectivity. Their neighbors were not only connected to the central cell, but also to each other, forming compact and cohesive “circles”. This observation led us to rethink what defines a rare cell population: perhaps rarity is not about spatial isolation, but about how cells are locally organized.
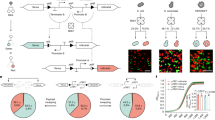
Based on this insight, we defined a metric, Q, for each cell to quantify the cohesiveness of its local neighborhood. Cells with high Q values reside in highly “cliquish” local structures. Building on this idea and incorporating network diffusion, we developed the RareQ method. Across a wide range of datasets, RareQ demonstrated stable performance in both sensitivity and accuracy. Notably, due to its simplicity, the method is computationally efficient and scales well to very large datasets. Its performance also generalizes consistently to multimodal single-cell and spatial transcriptomics data.
A reviewer’s comment provided an additional layer of insight. He pointed out that the local topological features captured by RareQ bear a strong conceptual resemblance to those described in small-world network theory. While we had not explicitly framed our work in this context, the connection is indeed natural: systems that appear sparse at a global scale may exhibit high connectivity locally. Signals that seem weak or noisy from a global perspective may, when viewed through a local structural lens, reveal meaningful biological organization.
Looking back, the most valuable outcome of this work is not the algorithm itself, but a shift in perspective. Rare cells may not be isolated entities; instead, they can exist as highly organized units embedded within their own “circles”.
More broadly, this perspective reminds us that in complex biological systems, “rare” does not necessarily mean random or noisy. Rather, signals that appear insignificant at a global scale may exhibit clear and stable patterns when examined locally. In other words, what we often overlook is not the signal itself, but the way we choose to observe it.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in