Highlights of the BMC Series – April 2025
Published in Healthcare & Nursing, Computational Sciences, and Genetics & Genomics
BMC Genomics - Growth resilience to weather variation in commercial free-ranging chickens in Ethiopia

Chicken farming is a growing industry in Ethiopia and demand is expected to increase by over 200% in the next 25 years. The majority of flocks are made up of indigenous breeds that are adapted to local conditions. However, a growing portion is made up of imported high-productivity breeds that fail to thrive due to increasing extremes of temperature and other climate-change induced environmental stressors.
This paper aimed to investigate the effects of changing weather conditions on the growth of commercially raised chickens by looking at both growth resistance phenotypes and associated genomic architecture. Over 1,500 cross-bred chickens were weighed weekly and were genotyped for this study. Weekly growth was compared to weather patterns to determine resilience phenotypes and genetic markers were identified for these phenotypes.
The authors found that growth was significantly impacted by temperature, humidity, and precipitation. They also found significant genomic variance between resilience phenotypes and identified specific candidate genes associated with lipid metabolism and adipocyte homeostasis. These results could be used to breed chickens that are both highly productive and adapted to local weather conditions.

Anemia in women of reproductive age is a prevalent problem worldwide, and India has one of the highest rates at 57%. Efforts to reduce the prevalence of anemia have focused on ways to increase the availability and distribution of supplements to prevent and treat anemia. However, there has not been much focus on interventions to increase consumption of supplements and understanding of medical guidelines.
In this longitudinal cluster randomized controlled trial, an intervention package that focused on changing social norms was tested as a way to increase iron folic acid consumption among women of reproductive age. The intervention package comprised community-based education sessions, health communication videos, and hemoglobin testing. Over 4,000 women were recruited for the trial and were randomized by geographical cluster to either the intervention or the control.
The results showed that the intervention improved social norms around supplement consumption compared with the control as well as increased consumption more in intervention communities. This highlights a potentially useful way to increase uptake of important micronutrient supplements and reduce the burden of anemia.
BMC Primary Care - Exploring the relationship between cultural and structural workforce issues and retention of nurses in general practice (GenRet): a qualitative interview study

Nurses are a vital part of healthcare services, but current shortfalls in the nursing workforce are expected to worsen in the coming decade. In England, the shortage of nurses is especially acute in general practice, with up to 28% of nurses considering leaving general practice within one year.
In order to understand the factors that both encourage and discourage nurses to stay in general practice, the authors of this paper conducted a qualitative study. They interviewed 41 current and former general practice nurses and nurse leaders in England and Wales to explore how general practice culture and structure support or challenge workforce retention.
The findings indicate that a range of cultural and structural issues affect nurses’ intention to remain in general practice in both positive and negative ways. These issues include recognition of the value of nurses in general practice, limited input into decision-making, and access to professional development and representation. These insights suggest ways that stakeholders, including policy makers, employers, and professional organizations, can support nurses to remain and flourish in general practice.
BMC Gastroenterology - An artificial intelligence model utilizing endoscopic ultrasonography for differentiating small and micro gastric stromal tumors from gastric leiomyomas
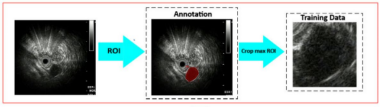
Gastric stromal tumors (GSTs) are a subtype of gastrointestinal submucosal tumors (SMTs) that have high malignant potential and often need aggressive surgical and pharmaceutical intervention. Gastric leiomyomas (GLs) are a less-common subtype of SMTs that are usually benign and require treatment only if they grow or cause other symptoms. Differentiating between these two subtypes is crucial during the diagnostic process, but accurately distinguishing them using endoscopic ultrasonography is challenging, especially for smaller tumors (<2.0 cm).
To address this challenge, the authors of this paper created an artificial intelligence (AI) model trained on images of confirmed GSTs and GLs. The model was then validated on a different set of images.
The AI model was able to accurately differentiate between GSTs and GLs smaller than 2.0 cm. When multiple images of each tumor were used in the AI model, its diagnostic efficiency was better than that of clinical prediction models and endoscopists. This model could offer valuable support to clinicians to accurately diagnose and treat small SMTs.
Follow the Topic
-
BMC Primary Care
Previously known as BMC Family Practice, this is an open access, peer-reviewed journal that considers articles on all aspects of primary health care research.
-
BMC Nutrition
BMC Nutrition is an open access, peer-reviewed journal that considers articles on all aspects of nutritional sciences.
-
BMC Gastroenterology
This is an open access, peer-reviewed journal that considers articles on all aspects of the prevention, diagnosis and management of gastrointestinal and hepatobiliary disorders, as well as related molecular genetics, pathophysiology, and epidemiology.
-
BMC Genomics
This is an open access, peer-reviewed journal that considers articles on all aspects of genetics, genomics and proteomics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics of human pathogens
For many years now, the study of pathogens’ genomes has enabled more accurate and timely diagnostics while allowing the tracking of specific strains or isolates. This can help discover the source of a specific epidemic and how it spreads, which in turn informs the best way to contain it. However, recent advances in third-generation sequencing techniques and the bioinformatics tools linked with them have transformed practice. It is now possible to sequence individual genomes much faster and much cheaper, which is opening new avenues for pathogen tracking, even in areas with limited funding such as the global South.
Advances in genomic technologies will also facilitate the emergence of personalized medicine approaches tailored to specific infections—such as genome-guided antiretroviral therapy for HIV or drug-resistance profiling in tuberculosis. Real-time genomic surveillance, exemplified by platforms like GISAID or other repositories during the COVID-19 pandemic, enables rapid detection and monitoring of emerging variants, thus improving public health responses.
This Collection invites submissions looking at human pathogens genomes and data sets to build on the knowledge already accumulated. We will also consider rapid diagnostics datasets and new techniques in genomic studies of human pathogens.
If you have a data note to submit, please submit to our sister collection Genomics of human pathogens data notes in BMC Genomic Data.
Topics of interest include, but are not limited to:
- Genomic characterization of bacterial pathogens
- New genomes of pathogens or strains
- Viral genomics and infection dynamics
- Comparative genomics of human parasites
- Microbial resistance mechanisms in pathogens
- Genomic epidemiology of infectious diseases
- Host-pathogen genomic interactions
- Metagenomic approaches to pathogen discovery
- Evolutionary genomics of emerging infections
- Functional genomics and regulatory networks in pathogens
This Collection supports and amplifies research related to SDG 3, Good Health & Well-Being.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Jul 20, 2026
Cattle genomics
BMC Genomics invites researchers to contribute to our Collection on Cattle genomics focusing on understanding the genetic makeup of bovine species, which is essential for improving livestock breeding and health. Advances in genomic technologies, such as next-generation sequencing and RNA sequencing, have enabled researchers to reveal insights into traits such as growth, meat quality, milk production, disease resistance, reproductive fitness, and overall adaptability in bovine genomes. This Collection aims to highlight the latest research developments in cattle genomics, encompassing both genomic and transcriptomic studies that contribute to the understanding of bovine biology.
Recent breakthroughs in genomic selection and precision breeding techniques have already shown promise in increasing efficiency in cattle production. The use of CRISPR-Cas genome editing, for example, has allowed for precise modifications to the cattle genome, introducing beneficial genetic variations without the linkage drag associated with traditional breeding methods. Additionally, the integration of omics technologies is paving the way for a holistic understanding of cattle biology, allowing for more effective management and breeding strategies. Studying the rumen microbiome using genomics, transcriptomics, proteomics, and metabolomics has revealed how microbial communities contribute to feed efficiency and nutrient absorption. This comprehensive approach enables targeted nutritional strategies that improve cattle health and productivity while reducing environmental impact. Such integrative studies facilitate the selection of cattle with optimal microbiome compositions, leading to more sustainable and efficient cattle production systems.
As research in cattle genomics progresses, we can anticipate the development of more sophisticated genomic tools that will enable precise manipulation of genetic traits in bovine populations. This may lead to enhanced resilience against diseases, improved reproductive performance, and better adaptation to changing environmental conditions. Ultimately, continued innovation in this field holds the potential to reform cattle production systems, ensuring sustainable livestock farming for future generations.
- Genomic selection in cattle breeding
- Transcriptomic analysis of bovine traits
- Pathogenicity and disease resistance genomics
- Advances in RNA-Seq applications for cattle
- Omics approaches to cattle health and productivity
- Genetic mapping of economically important traits
- Gene editing
- Metagenomics of the bovine gut microbiome
- Epigenetic regulation of growth and reproduction
- Comparative genomics of cattle and other livestock species
This Collection supports and amplifies research related to SDG 2, Zero Hunger.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Aug 26, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in