Host Cell Metabolism Goes Viral: Unveiling How HEV Hijacks Your Cells
Published in Chemistry, Microbiology, and Cell & Molecular Biology
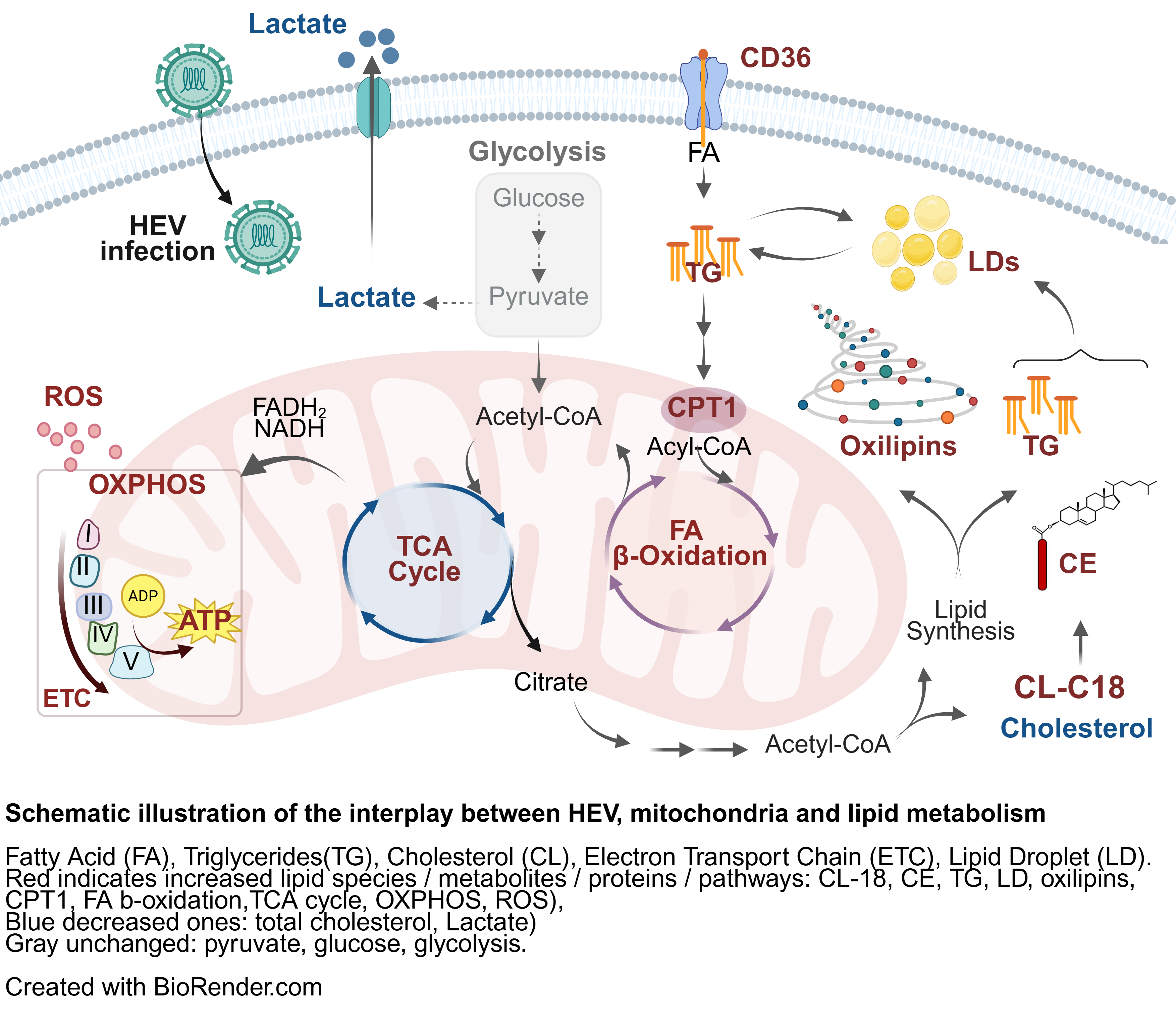
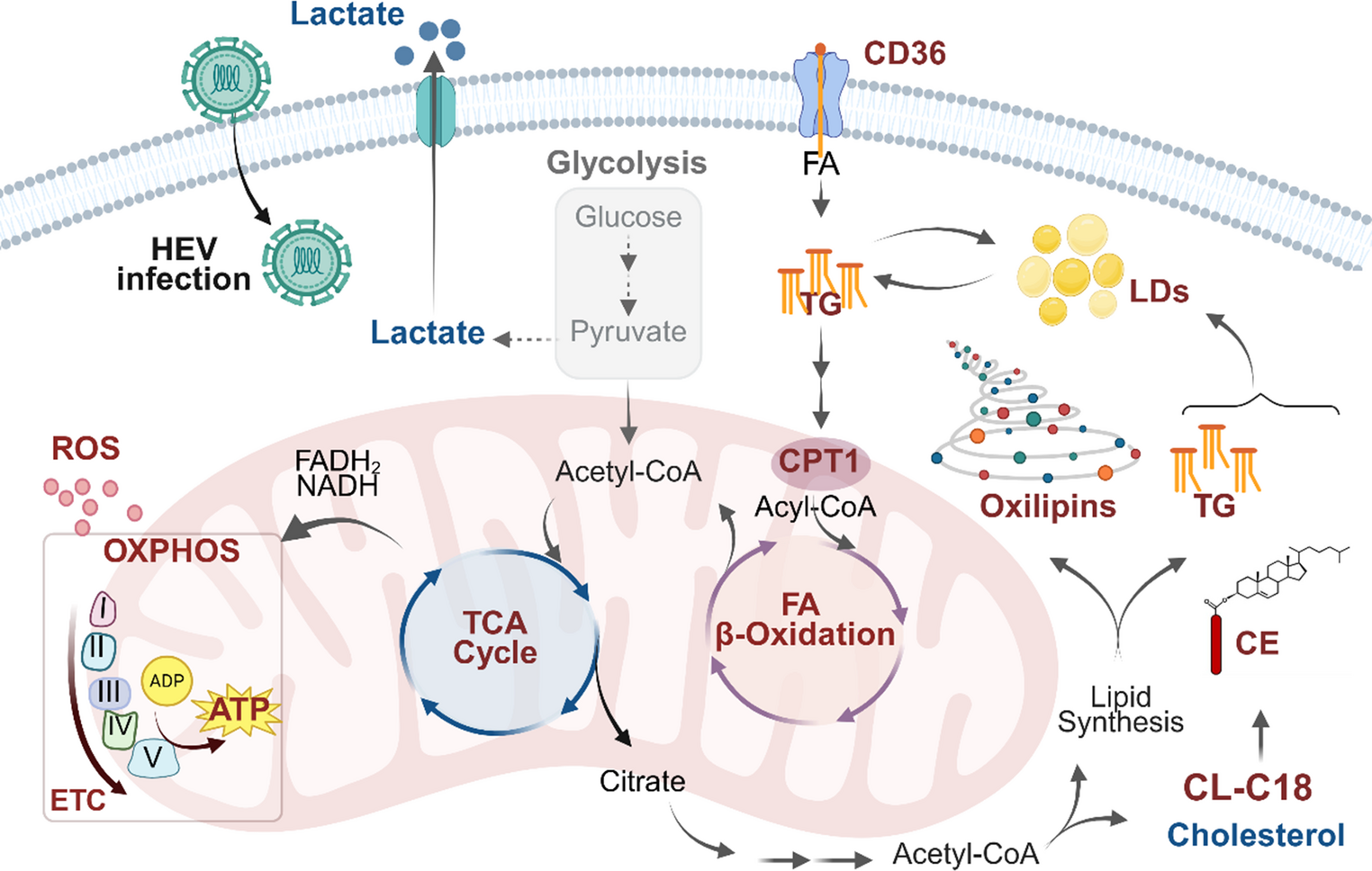
Our latest study (Glaziou et al., 2026, Mol Cell Life Sci) reveals that HEV hijacks host cell metabolism to fuel its replication:
- TCA cycle, mitochondrial oxidative phosphorylation (OXPHOS), fatty acid metabolism, and β-oxidation, are extensively rewired.
- Lipid storage organelles expand, long-chain neutral lipids accumulate, and pro-inflammatory oxylipins surge.
- HEV relies on lipid-fuelled OXPHOS rather than glycolysis.
- Distinct metabolic programs for HEV-1 versus HEV-3, aligning with their differential pathogenicity.
These findings highlight critical host metabolic vulnerabilities exploited by HEV and provide a roadmap for targeting metabolic hubs as therapy.
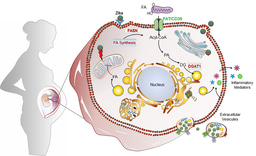
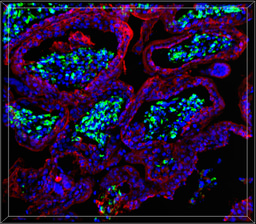
Interestingly, while our previous work in placental models (Gouilly et al., 2018, Nat Commun) showed HEV-1 triggers inflammatory damage at the maternal-fetal interface, the placenta is a highly active metabolic organ, and how HEV reshapes its metabolism remains an exciting question for future studies.
Takeaway: HEV doesn’t just infect—it reprograms, fuels, and manipulates host metabolism to thrive. Understanding these strategies could point to novel therapeutic interventions.
Follow the Topic
-
Cellular and Molecular Life Sciences
Cellular and Molecular Life Sciences (CMLS) is a multidisciplinary Open Access journal covering the latest aspects of biological and biomedical research.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in