How a Cellular “Gatekeeper” Helps Viruses Thrive: Exploring NUP153 in Flavivirus Infections
Published in Microbiology
If you’ve ever looked up the definition of a virus, you may have seen the phrase “viruses are obligate intracellular parasites”. In simple terms, this means viruses can only make more of themselves inside living cells. They don’t have the tools they need to reproduce on their own, they instead must hijack the cell’s machinery.
In our research group, led by Prof. Anna Överby, we study a family of viruses called orthoflaviviruses. This group includes well‑known viruses such as Zika (ZIKV), Dengue (DENV), and tick-borne encephalitis virus (TBEV). You may have heard of these in the news, as they can cause severe disease and in some cases death, affecting millions of people worldwide.
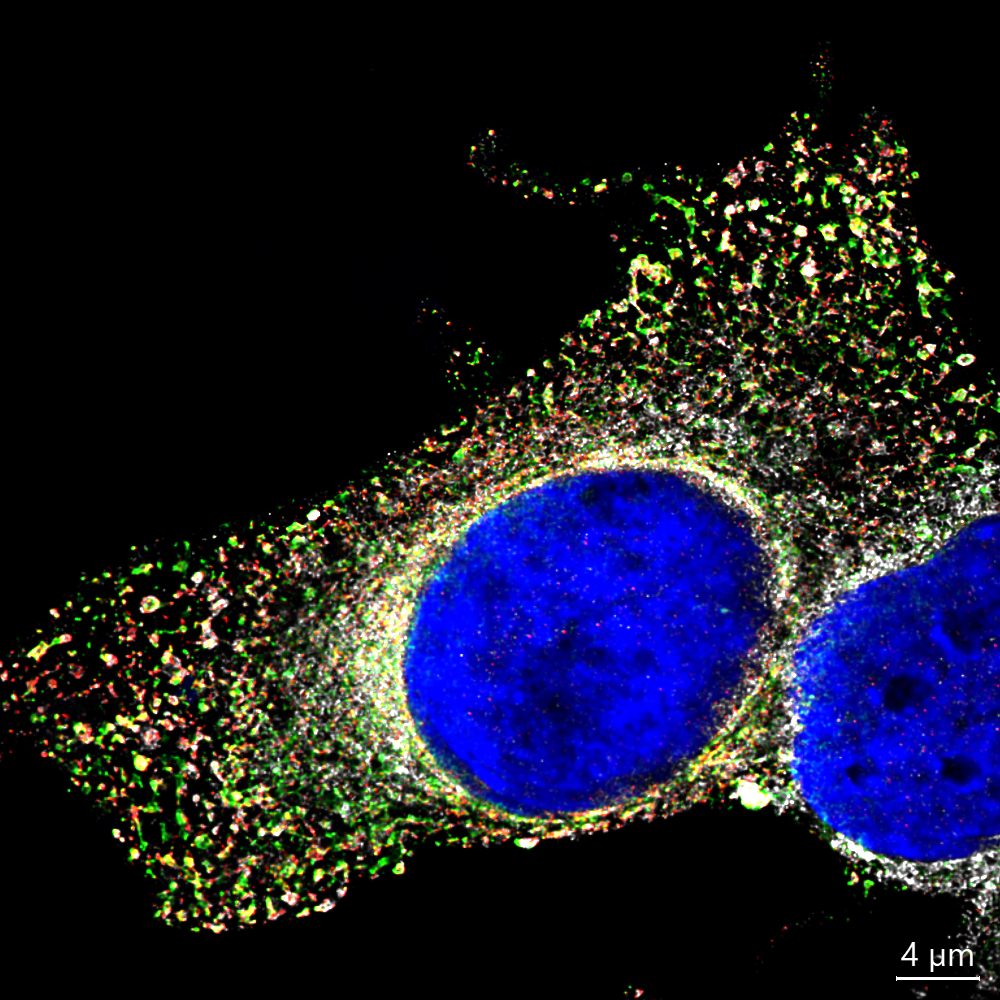
Orthoflaviviruses create small compartments in the cell’s cytosol (the fluid outside the nucleus). Inside these compartments, they copy their RNA genome, which can then be used to produce the proteins they need to assemble new virus particles. Most of their life cycle happens in the cytosol, but interestingly, we still don’t fully understand how proteins from the nucleus influence infection. This knowledge gap led us to focus on a family of nuclear proteins called nucleoporins or NUPs.
NUPs are the building blocks of the nuclear pore complex. Think of this structure as a highly selective doorway in the nuclear membrane that controls what moves in and out of the nucleus: such as RNA, proteins, and other essential molecules.
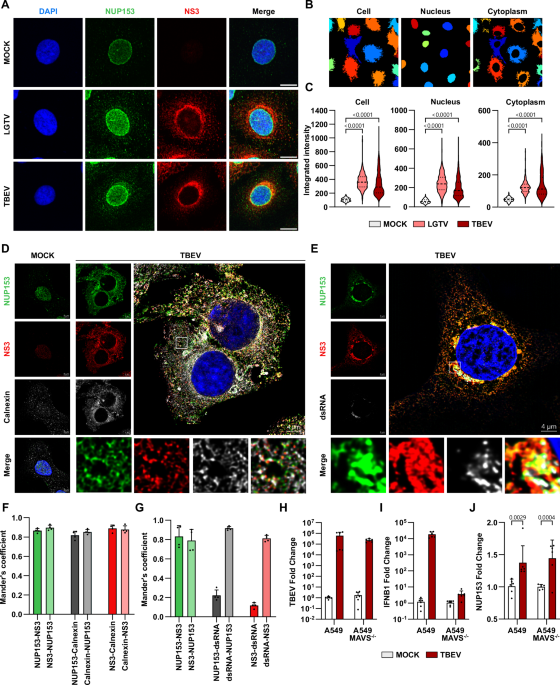
When we examined the location of NUPs during infection, we were surprised to find that some of them were no longer confined to the nuclear membrane. Instead, they appeared in the cytosol right next to viral components. Among these, one NUP stood out: NUP153.
To understand the function of NUP153 during infection, we needed to remove it from cells and see what happens. But there was a challenge: NUP153 is a very stable, essential protein. Traditional methods to “knock it out” often cause the cells to die. Luckily, we obtained special cells where NUP153 can be rapidly degraded using the auxin‑inducible system without harming the cells themselves. Using this tool, we discovered that when NUP153 is removed, orthoflavivirus infection decreases. This suggests that NUP153 normally “helps” the virus replicate.
NUP153 is known to bind RNA, and since these viruses have RNA genomes, we wanted to see if there was a direct interaction. Indeed, NUP153 bound to viral RNA and even more interestingly, it bound at a very specific location on it: the region between the genes encoding structural and non‑structural proteins.
We also found that during infection, NUP153 began interacting with viral proteins and different sets of cellular proteins than usual, including several involved in translation—the process that turns RNA into proteins. Viruses rely heavily on hijacking this machinery.
Putting these findings together, we examined how viral protein production changed in cells lacking NUP153. We saw that the ratio of structural to non‑structural viral proteins shifted. Structural proteins form the virus particle, while non‑structural proteins help replicate the genome and evade the immune system. Altering this balance disrupts the virus’s ability to efficiently produce new particles. As a result, virus production slows down, leading to delayed spread.
Why does this matter?
Although vaccines exist for some orthoflaviviruses, there are currently no antiviral drugs that directly target them. To develop effective treatments, we need a detailed understanding of how these viruses interact with the host cell.
Our study uncovers a new role for NUP153 in infection and highlights how viruses repurpose even essential cellular proteins for their own benefit. While this is only one piece of a much larger puzzle, it brings us closer to identifying potential targets for future antiviral therapies.
Much remains to be explored but every new insight moves us one step closer to developing tools that can protect people from these dangerous pathogens.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in