How the closest relatives of eukaryotes grew a larger genome
Published in Ecology & Evolution, Microbiology, and Genetics & Genomics
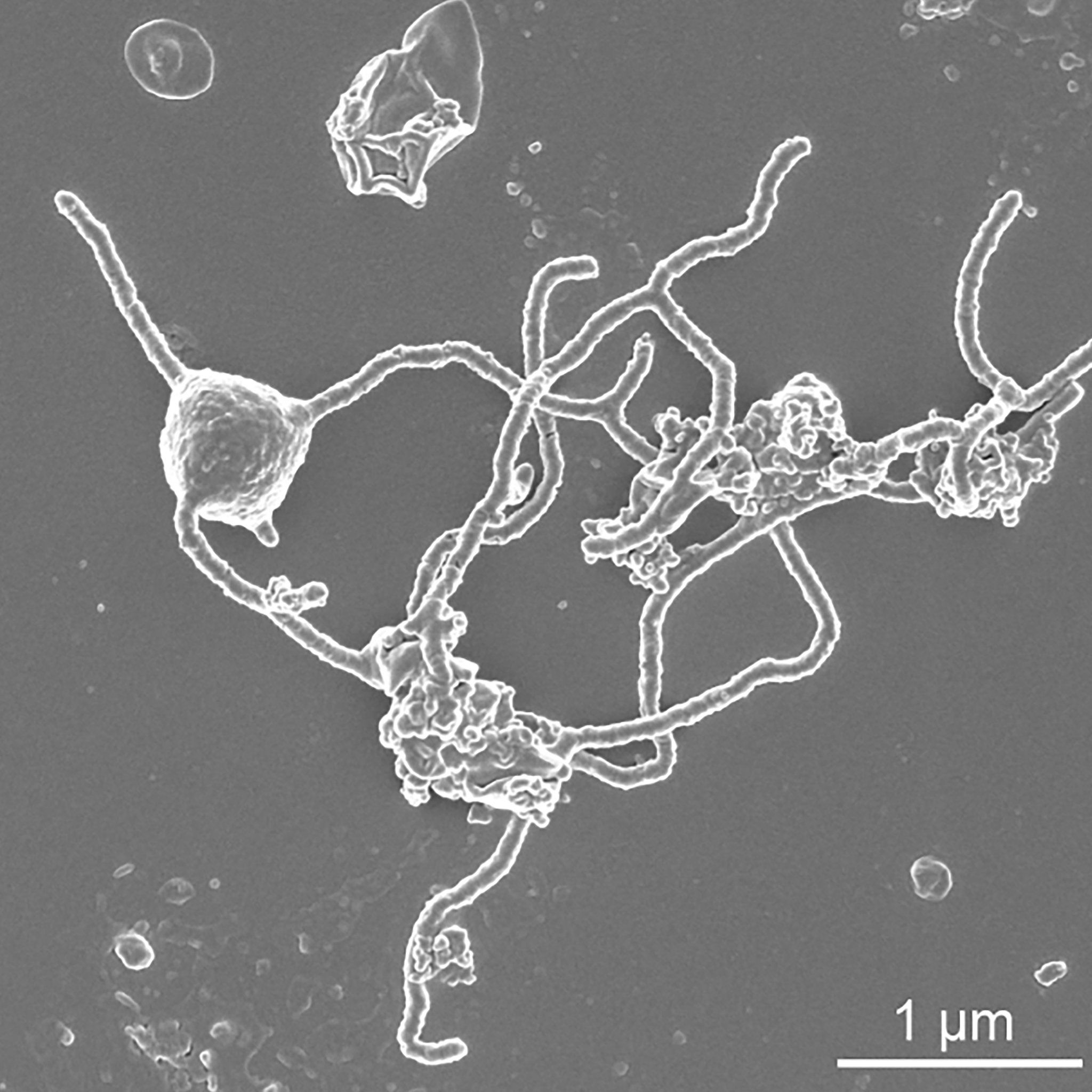

The origin of the eukaryotic cell is one of the most puzzling events in the history of life on Earth. The predominant view is that eukaryotes emerged through symbiotic interactions between (at least) an alpha-proteobacterial endosymbiont that later become the mitochondrion and an archaeal host. The phylogenetic affiliation of the latter remained elusive until about 10 years ago, when metagenomic sampling of the deep sea revealed an archaeon with remarkably eukaryote-like features hidden in its genome. Since then, this archaeon and its relatives (collectively named Asgard archaea, or phylum Asgardarchaeota) have become a cornerstone of eukaryogenesis research, offering us an unprecedented view into the nature of one of the two key partners of the symbiosis that paved the way for plants, fungi, and even us.
Our group has long been studying eukaryogenesis from a phylogenomics perspective, in particular through the characterization of the Last Eukaryotic Common Ancestor (LECA) and the ancestry of its gene families. Asgard archaea have been shown to encode gene families previously thought to be eukaryote-specific (Eukaryotic Signature Proteins, or ESPs), and display other remarkable eukaryote-like features, such as a dynamic cytoskeleton, and a significantly larger genome than their closer archaeal relatives. These recent discoveries mean that some of the traits and cellular structures that we considered eukaryotic hallmarks may have originated much before LECA, wanting us to discern how they originated within the Asgard archaeal lineages.
The recent years have seen a boom in the number of publicly available genomes of the Asgard clade as well as improvements in the phylogenomic methodology, which posed an unique opportunity for us to explore the genome dynamics of this intriguing clade with a particular focus on what it could tell us about the origin of eukaryotic traits.
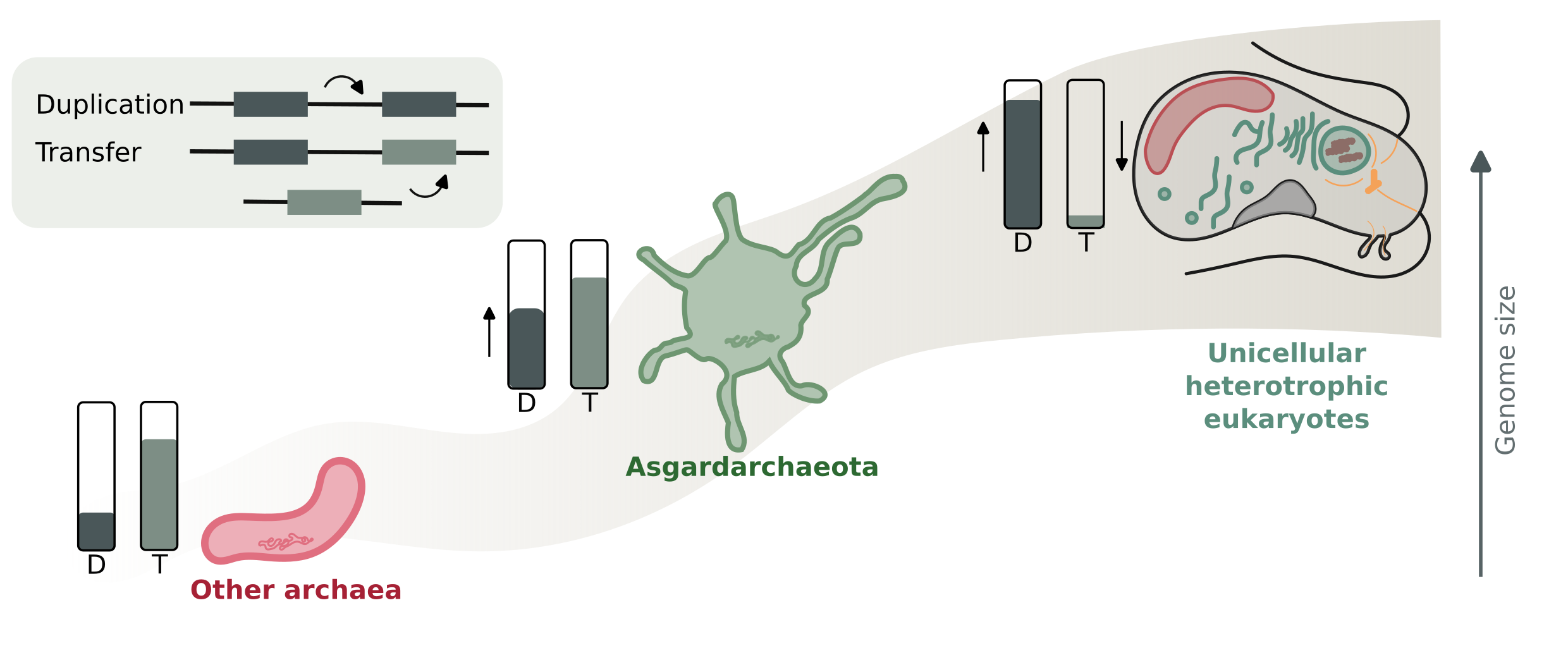
Figure: Schematic view on the processes governing genome size in non-Asgard archaea (exemplified by Thermoproteota), Asgardarchaeota, and unicellular eukaryotes. D= Duplication rates, T= Transfer rates. Asgardarchaeota occupy an intermediate position in terms of the D/T balance between unicellular eukaryotes and other archaea.
We first explored the evolution of genome size in Asgard archaea against the backdrop of its closest prokaryotic relatives, Thermoproteota, to gain insights on whether this process was driven via Horizontal Gene Transfer (HGT) or duplications, as each of these processes dominate genome dynamics in prokaryotes and eukaryotes, respectively.
We reconstructed the evolutionary histories of about 4,700 gene families across a representative set of Asgard archaea, Thermoproteota and other prokaryotes, and compared these gene histories to the phylogeny of the species to infer gene duplication, loss and HGT across all considered lineages. Our results indicated that gene duplication and not HGT was the major process behind the increased genome size in Asgard archaea relative to Thermoproteota. This high duplication rate was atypical for a prokaryote and situate Asgard archaea in a sort of intermediate position between prokaryotes and unicellular eukaryotes, highlighting the ‘stepping stone’ nature of the Asgard clade and raising the intriguing possibility that the closest Asgard archaeal ancestor of eukaryotes was somewhat ‘primed’ for further genome expansions during eukaryogenesis. Moreover, a detailed analysis of the ESPs displayed an increase in intra-Asgard HGT relative to other gene families, which could explain their patchy distribution within current Asgard genomes despite their inferred presence in LAECA.
Furthermore, an in-depth analysis of HGT from bacteria into Asgard archaea revealed a predominance of metabolic genes, generally acquired individually rather than as whole metabolic modules, and which were peripheral within the organism’s protein-protein interaction network. HGT from bacteria into Asgards occurred continuously, from diverse partners and in a lineage-specific manner, but we found no evidence for any of these transferred genes to have made it to the Last Eukaryotic Common Ancestor (LECA). This argues against the idea that the ‘surplus’ of gene families with bacterial origin in LECA result from accumulated ancient HGT into the archaeal ancestor of eukaryotes. Alternative explanations, including HGT to LECA from additional bacterial partners, are needed to explain this discrepancy.
In summary, our work traced the evolution of Asgard archaeal genomes, and revealed a new trait in which Asgard archaea seem to be at an intermediate position between prokaryotes and eukaryotes: duplication-driven genome expansion.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in