Learning the rules of metabolic innovation in yeast evolution
Published in Ecology & Evolution, Microbiology, and Genetics & Genomics
We often hear stories where basic science discoveries end up having important applications. The research in this paper is an example of a project with an initially applied objective turning into a more basic science story. I was a postdoctoral research fellow at the Great Lakes Bioenergy Research Center (GLBRC) and one of my objectives was to identify new yeast species that could be promising for biofuel production. Since optimizing the rate of biofuel production is important to making the process more environmentally friendly and economically feasible, we were looking for yeasts with naturally high rates of metabolism that could rapidly convert sugars to alcohol.
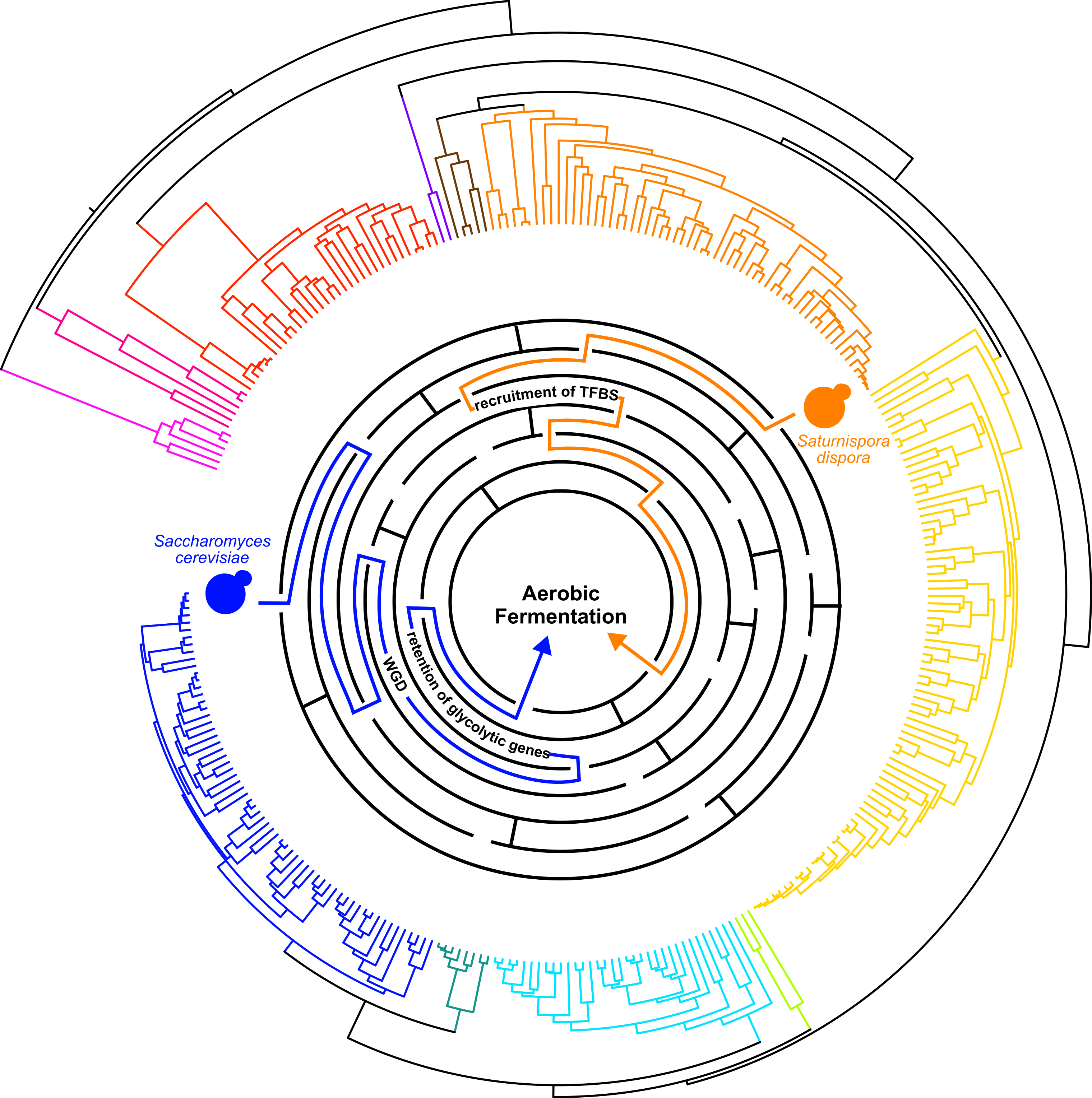
To measure the rate of glycolysis, a central metabolic pathway, I optimized an extracellular acidification rate assay (ECAR) that would be scalable across hundreds of species. After measuring ECAR in nearly 300 different species, I noticed a cluster of rapid ECAR yeasts in the genus Saturnispora. I wanted to make sure that our screening was identifying yeast with rapid glycolysis, so I grew a few species in the genus Saturnispora in larger volumes tracking the growth and collecting the media to measure glucose and ethanol levels over time. This follow up showed that these yeasts consumed glucose rapidly, produced ethanol aerobically, and produced less cells than closely related species with low ECAR. These are the hallmarks of not only rapid glycolysis but also an interesting metabolic trait in yeast, aerobic fermentation, which is when yeasts produce ethanol even when there is oxygen around. This set us off on the task of identifying a new mechanism of aerobic fermentation.
We reasoned that by comparing gene content between species with aerobic fermentation and closely related species without aerobic fermentation, we would be able to identify the genes responsible for our phenotype. To test our predictions, I optimized a method to genetically modify one of the species with aerobic fermentation, Saturnispora dispora, to systematically delete candidate genes which were absent in the low ECAR species. However, for many months, I would make a mutant and measure its ECAR, only to find no impact on the phenotype. This was a very frustrating stage, but I was determined to figure it out. I decided to sequence the transcriptomes of two rapid and two low ECAR species, hoping that this would narrow down the candidates by determining which genes were actually being expressed in the species with aerobic fermentation. When I analyzed the data, I immediately noticed a striking pattern, in which many of the genes encoding glycolytic enzymes, especially those at the end of the pathway, have much higher expression in the species with aerobic fermentation.
At this point, I shifted my focus to the transcription factors that might be influencing the expression of glycolytic genes. I systematically identified putative transcription factor binding sites in the promoters of glycolytic genes across our four focal species. I noticed more Gal4p-binding sites in the promoters of rapid ECAR species. These additional binding sites were consistent in the promoters upstream of the genes encoding the glycolytic enzymes: hexokinase, glyceraldehyde-3-phosphate dehydrogenase, and enolase. These results directly connected to two papers that I had been reading and re-reading throughout this project.
First, a 2007 paper modeled that these three enzymes, all retained as duplicates in the model yeast, Saccharomyces cerevisiae, had the biggest influences on glycolytic flux. And second, a 2018 paper showing that, in Komagataella phaffii (a species which can’t grow on galactose), overexpression of a Gal4-like transcription factor induces aerobic fermentation. It finally felt like all the pieces of the puzzle were coming together! I anxiously made a deletion mutant lacking this transcription factor to test if Gal4p was truly impacting the ECAR phenotype. When I tested out the new strains, the mutants had lost the rapid ECAR phenotype, consumed glucose slowly, and stopped producing ethanol, all supporting our hypothesis that Gal4p was required for the aerobic fermentation phenotype!
As I was writing up the manuscript, the parallels between the duplicated genes in Saccharomyces and upregulated genes in Saturnispora really stuck with me. The more we discussed this, the more we thought these data were hinting at an evolutionary constraint where increased abundance of these enzymes is key to increased glycolytic flux and aerobic fermentation. And here we have examples of two different molecular mechanisms (Fig. 1), gene duplication and transcription factor binding site recruitment, both acting on the same genes to increase the enzyme’s abundance. This interpretation also broadens the applicability of these findings to explore genomic data and identify new candidates for rapid glycolytic flux based on sequence data alone. We also know that convergent evolution is observed throughout the tree of life, but in many cases, we don’t yet know the genomic basis underlying the convergence. I expect that, in the future, comparing genomes and phenotypes in non-model species to what we already know about model organisms will help us identify more mechanisms and constraints that influence trait evolution.
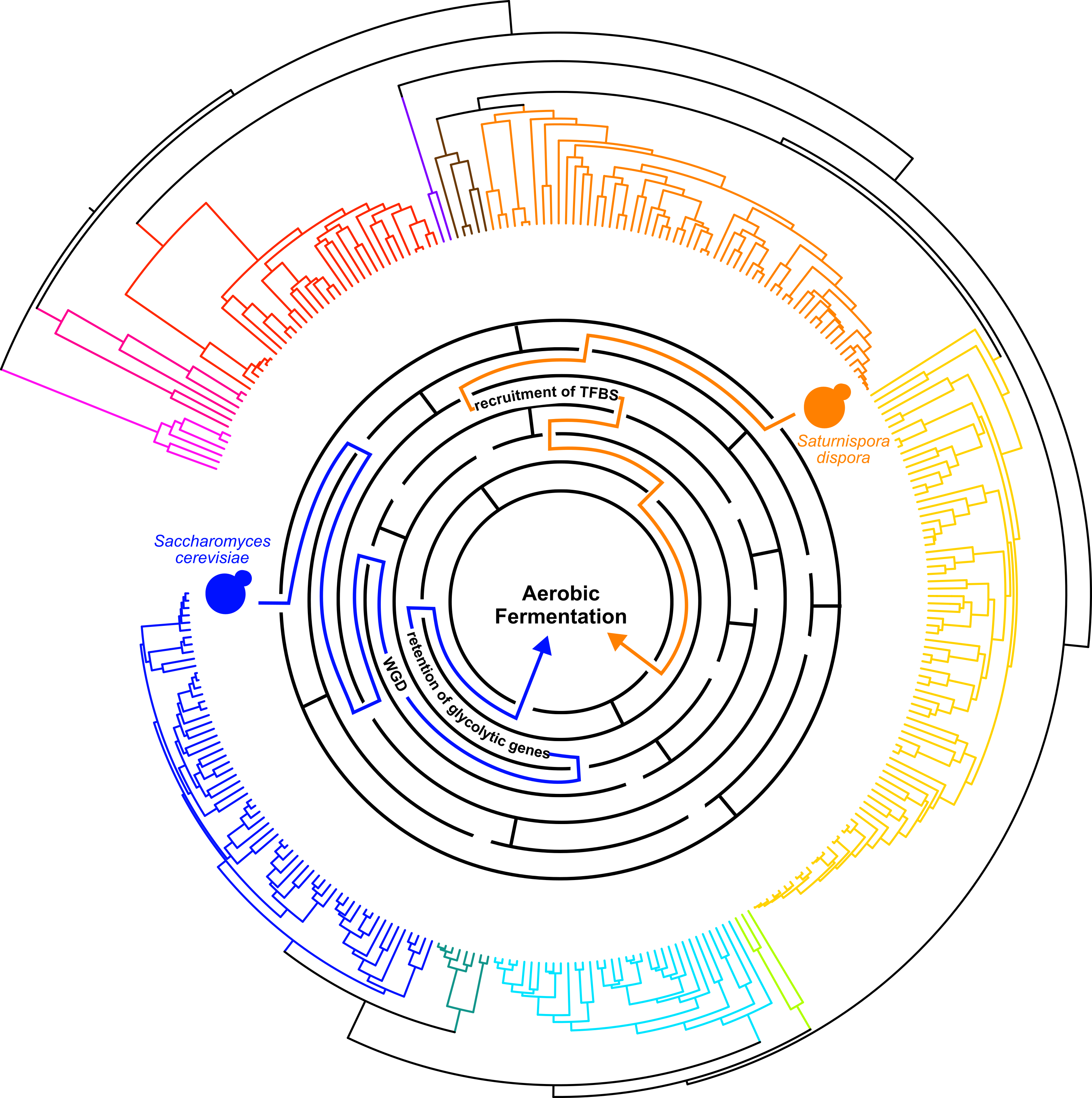
Figure 1. Convergent evolution of aerobic fermentation in Saccharomyces cerevisiae and Saturnispora dispora through different molecular mechanisms represented as different paths through a maze.
Follow the Topic
-
The EMBO Journal
Launched in 1982, The EMBO Journal is EMBO’s flagship publication, recognised worldwide for its originality, breadth, and enduring contributions to molecular biology.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in