Microbiome analyses of blood and tissues suggest cancer diagnostic approach
Published in Cancer
Cancer remained a conceptual entity to me until the first year of college.
Amidst end-of-year holidays in 2012, my grandmother made a visit to the doctor for abdominal bloating and gastrointestinal distress issues, thinking they were related to her episodic diverticulitis. The diagnosis, however, was much worse, and startled the whole family. She had unresectable stage IV pancreatic cancer.
With few options for late-stage disease, her doctors tried to use erlotinib (a targeted EGFR inhibitor), without much success. Not only did the therapy fail to regress the cancer, but the clinical sequelae left my grandmother feeling even worse. Ultimately, she chose to ‘live with the cancer’ rather than spend the rest of her short life suffering from adverse drug effects. Only 33 days passed from her diagnosis to her death.
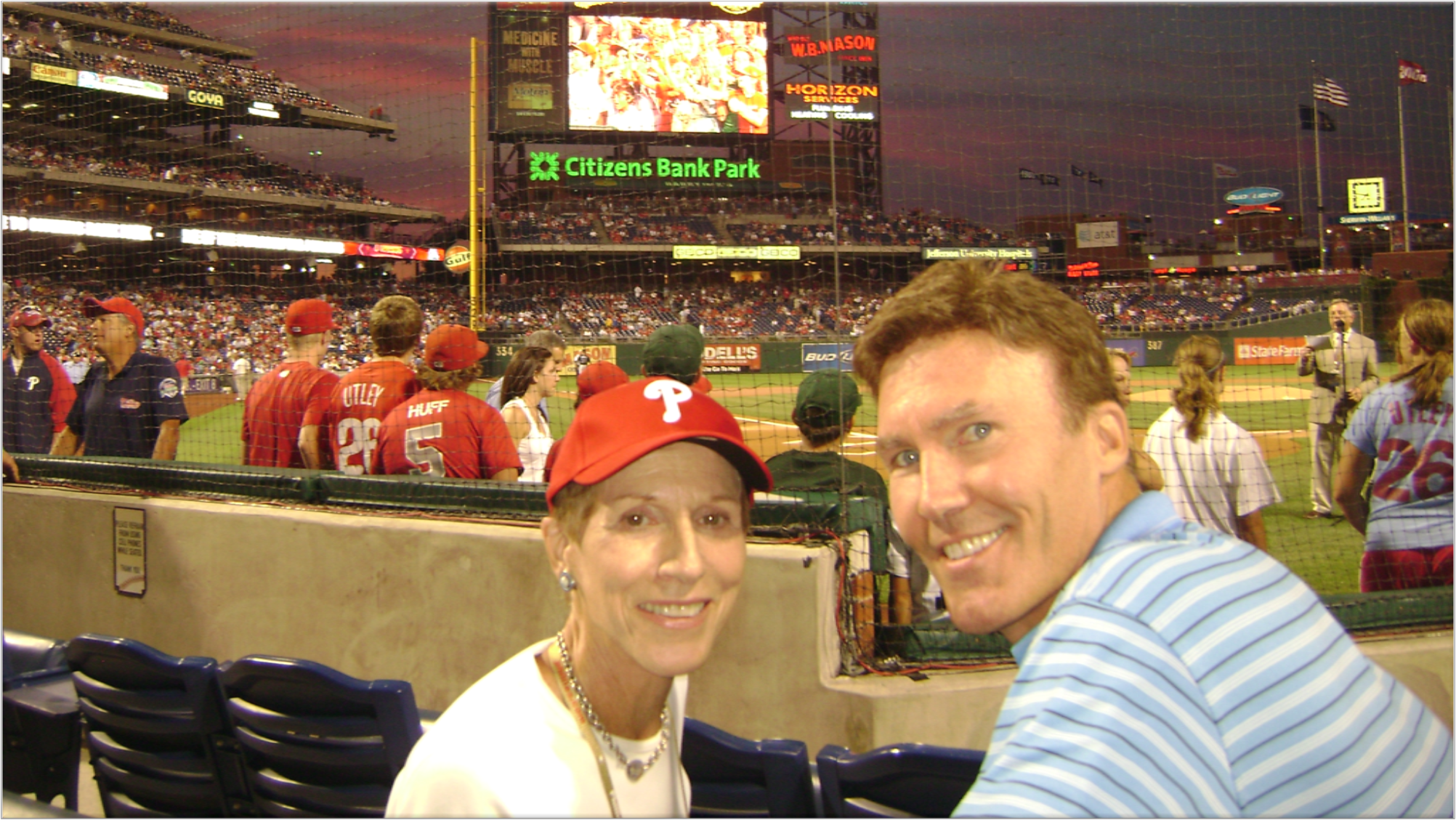

Figure 1: My grandmother, pictured with family, at a Phillies game in her last few years of life.
Without much direction, but with plenty of zeal, I now had a ‘face’ for cancer and threw myself into studying it. Many questions were on my mind: “Why did her cancer present so late? Why was it resistant to therapy? Why did it progress so quickly?”
Within a year, this pursuit led me to a sister institution of my undergraduate university named Duke-NUS Medical School (in Singapore), where I worked under Drs. Patrick Tan and Steve Rozen to study, using bioinformatics, how DNA mutations could distinguish pre-cancer from cancerous lesions in multiple cancer types [1–4]. The work powerfully transformed the way I viewed cancer diagnostics, and affirmed that cancer genomics would be a core part of my career and interests going forward.
Upon my return to Duke University in the United States, the director of precision medicine, Dr. Geoffrey Ginsburg, challenged me with a diagnostic project based on the host immune response in blood [5]. To my chagrin, the project was based on diagnosing infectious disease etiologies — rather than cancer — based on multi-omics and machine learning, but this work and concomitant skill development ended up being crucial later on for the Nature paper.
Coming to graduate school with a mixed background of cancer genomics and infectious disease genomics, it initially seemed as though they were entirely unrelated areas of study. This, however, all changed when a flurry of papers started appearing that described the involvement of microbes in cancer initiation, progression, and treatment resistance [6–11]. Catching my attention were the stories of bacteria inhabiting the majority of pancreatic cancers (76%), degrading its clinically-preferred chemotherapy (gemcitabine), and shaping short-term versus long-term patient survival — these hit very close to home [6,10]. Somehow, studying the cancer microbiome suddenly made perfect sense of my diverse set of past scientific experiences and paired it with personal passion. I decided to focus my MD/PhD work on this topic under the guidance of Professor Rob Knight at UC San Diego.
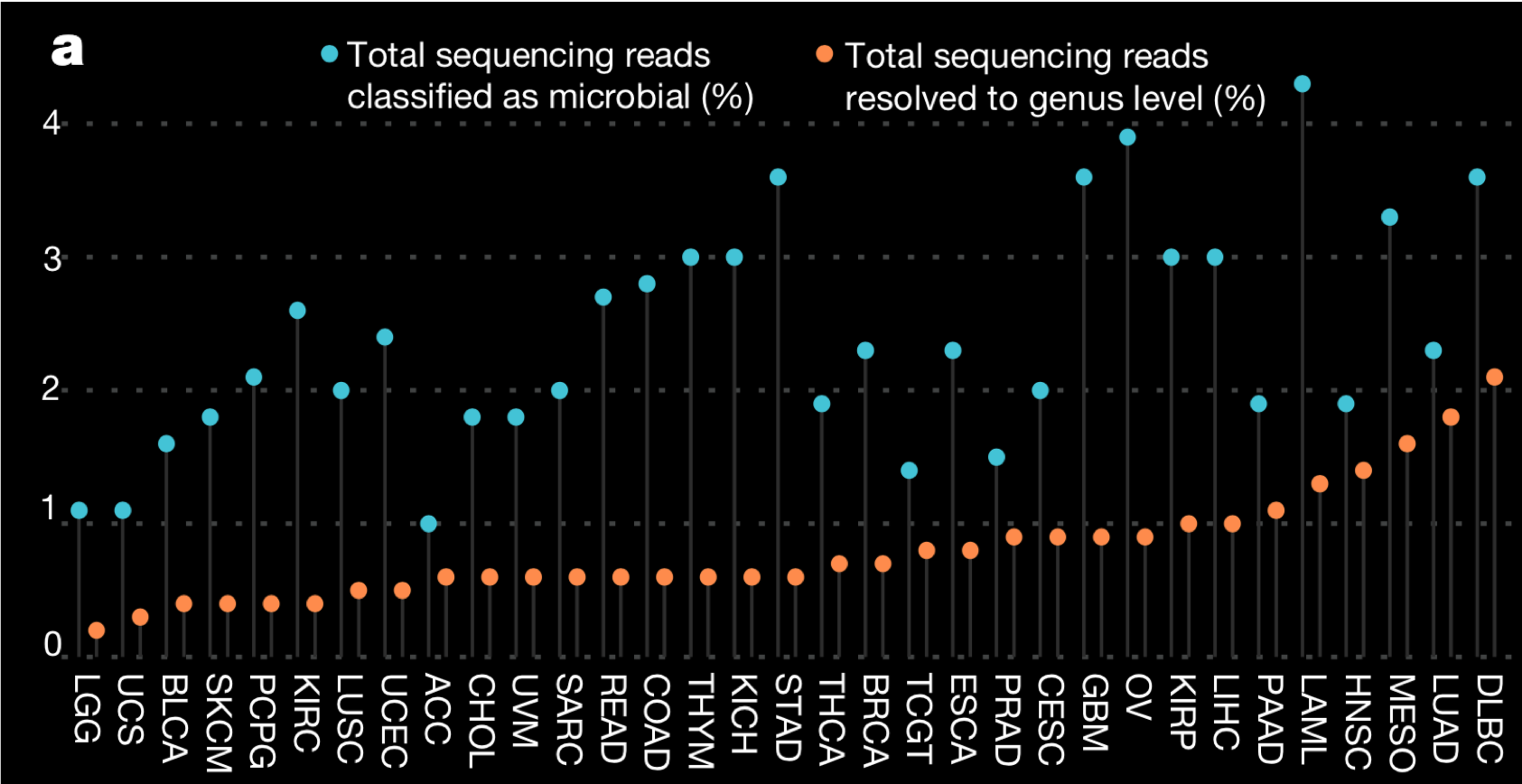
Given that there are more microbial cells (est. 39 trillion) and far more unique microbial genes (est. 2 million) in and on our bodies than their human counterparts (est. 30 trillion cells and 20,000 genes, respectively), we hypothesized that all major cancer types may harbor some degree of microbial involvement, and, if so, that this could provide valuable diagnostic information. So we started exploring The Cancer Genome Atlas (TCGA) for microbial DNA and RNA reads on an unprecedented scale. The details are contained in the paper, but an amazing discovery from this work is that the results suggest a new class of ‘oncobiome’ diagnostics (and possibly therapeutics) that could benefit future patients. Notably, and unexpectedly, we found that microbial information from a small sample of patient’s blood could be as or more effective than other currently available 'liquid biopsies' of cancer used in clinics today.
To accelerate the translation of this work to the clinic, several of us have started a company around microbial-based (“Micro-”) cancer (“-noma”) diagnostics (“Micronoma”). The many questions that remain to be explored in the field, both scientific and clinical, mean that there are many future discoveries to be made. We hope that this paper will benefit researchers, clinicians, and biotech alike in the days to come, and we invite you to solve them together for the benefit of future cancer patients, so that no one would have to go through the same surprise and pain that my grandmother experienced.
Written by Greg Poore with edits from Sandrine Miller-Montgomery and Rob Knight.
References
1. Lim WK, Ong CK, Tan J, Thike AA, Ng CCY, Rajasegaran V, et al. Exome sequencing identifies highly recurrent MED12 somatic mutations in breast fibroadenoma. Nat Genet. 2014;46: 877–880. doi:10.1038/ng.3037
2. Tan J, Ong CK, Lim WK, Ng CCY, Thike AA, Ng LM, et al. Genomic landscapes of breast fibroepithelial tumors. Nat Genet. 2015;47: 1341–1345. doi:10.1038/ng.3409
3. Nairismägi M-L, Tan J, Lim JQ, Nagarajan S, Ng CCY, Rajasegaran V, et al. JAK-STAT and G-protein-coupled receptor signaling pathways are frequently altered in epitheliotropic intestinal T-cell lymphoma. Leukemia. 2016;30: 1311–1319. doi:10.1038/leu.2016.13
4. Vettore AL, Ramnarayanan K, Poore G, Lim K, Ong CK, Huang KK, et al. Mutational landscapes of tongue carcinoma reveal recurrent mutations in genes of therapeutic and prognostic relevance. Genome Med. 2015;7: 98. doi:10.1186/s13073-015-0219-2
5. Poore GD, Ko ER, Valente A, Henao R, Sumner K, Hong C, et al. A miRNA Host Response Signature Accurately Discriminates Acute Respiratory Infection Etiologies. Front Microbiol. 2018;9: 2957. doi:10.3389/fmicb.2018.02957
6. Geller LT, Barzily-Rokni M, Danino T, Jonas OH, Shental N, Nejman D, et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science. 2017;357: 1156–1160. doi:10.1126/science.aah5043
7. Bullman S, Pedamallu CS, Sicinska E, Clancy TE, Zhang X, Cai D, et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science. 2017;358: 1443–1448. doi:10.1126/science.aal5240
8. Jin C, Lagoudas GK, Zhao C, Bullman S, Bhutkar A, Hu B, et al. Commensal Microbiota Promote Lung Cancer Development via γδ T Cells. Cell. 2019;176: 998–1013.e16. doi:10.1016/j.cell.2018.12.040
9. Meisel M, Hinterleitner R, Pacis A, Chen L, Earley ZM, Mayassi T, et al. Microbial signals drive pre-leukaemic myeloproliferation in a Tet2-deficient host. Nature. 2018;557: 580–584. doi:10.1038/s41586-018-0125-z
10. Riquelme E, Zhang Y, Zhang L, Montiel M, Zoltan M, Dong W, et al. Tumor Microbiome Diversity and Composition Influence Pancreatic Cancer Outcomes. Cell. 2019;178: 795–806.e12. doi:10.1016/j.cell.2019.07.008
11. Aykut B, Pushalkar S, Chen R, Li Q, Abengozar R, Kim JI, et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature. 2019;574: 264–267. doi:10.1038/s41586-019-1608-2
Follow the Topic
-
Nature
A weekly international journal publishing the finest peer-reviewed research in all fields of science and technology on the basis of its originality, importance, interdisciplinary interest, timeliness, accessibility, elegance and surprising conclusions.
Related Collections
With Collections, you can get published faster and increase your visibility.
Carbon Dioxide Removal
Publishing Model: Hybrid
Deadline: Jan 16, 2027




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Great work and a touching story!
So fruitful to read the paper for the students and researchers,Thanks George poore for this excellent research and provide all of us useful information.