Neanderthal feces on fire illuminates gut microbiome features
Published in Microbiology

Reference: “Components of a Neanderthal gut microbiome recovered from fecal sediments from El Salt”
Communications Biology. DOI: 10.1038/s42003-021-01689-y
Information about ancient human gut microbiomes could help answer many questions about how this extrasomatic organ impacts our physiology and intervenes on our environmental exposures 1. Some major questions are: What kinds of microbiota have persisted in our guts through the millennia, irrespective of geographic, climatic, genetic, and subsistence differences? What gamut of functions do these microbes perform for us? Are there disparities in the prevalence of quintessential microbiota among living people, and what does this imply? In short, there is a pressing desire to contextualize the attributions we place on the role of the gut microbiome today by analyzing its configurations in the past using ancient DNA (aDNA). The goal is to understand in what ways the gut microbiome evolved to integrate with our physiology, particularly in ancient times when life was precarious amidst a challenging and wild environment and during considerable ecological transformations.
Explorations with aDNA lend a picture of the nature of recent human evolution, migration, and the origins of modern human dispersals around the globe 2. Scientists have also begun to chart the evolutionary history of pathogens, including the microorganisms that cause leprosy, syphilis, smallpox, malaria, and tuberculosis 3.
Microbial genetics is entering a new age of characterizing entire microbial communities of the past, and in rare cases where samples avail, from the preserved tissues stemming from long-dead organisms. To date, only a handful of gut microbiome or metagenomic data have been recovered from coprolites (paleofeces), mummified intestinal contents, or latrine deposits 4,5. No gut microbiome dataset from human-derived tissue dates to earlier than 8000 years ago, casting us in darkness about the gut microbiome features of truly prehistoric humans. People who lived sometime prior to 10,000 years ago, during the Pleistocene, experienced an entirely different geological epoch demarcating climatic, environmental, and societal differences, which assuredly means that their lifeways and subsistence regimes have long gone extinct.
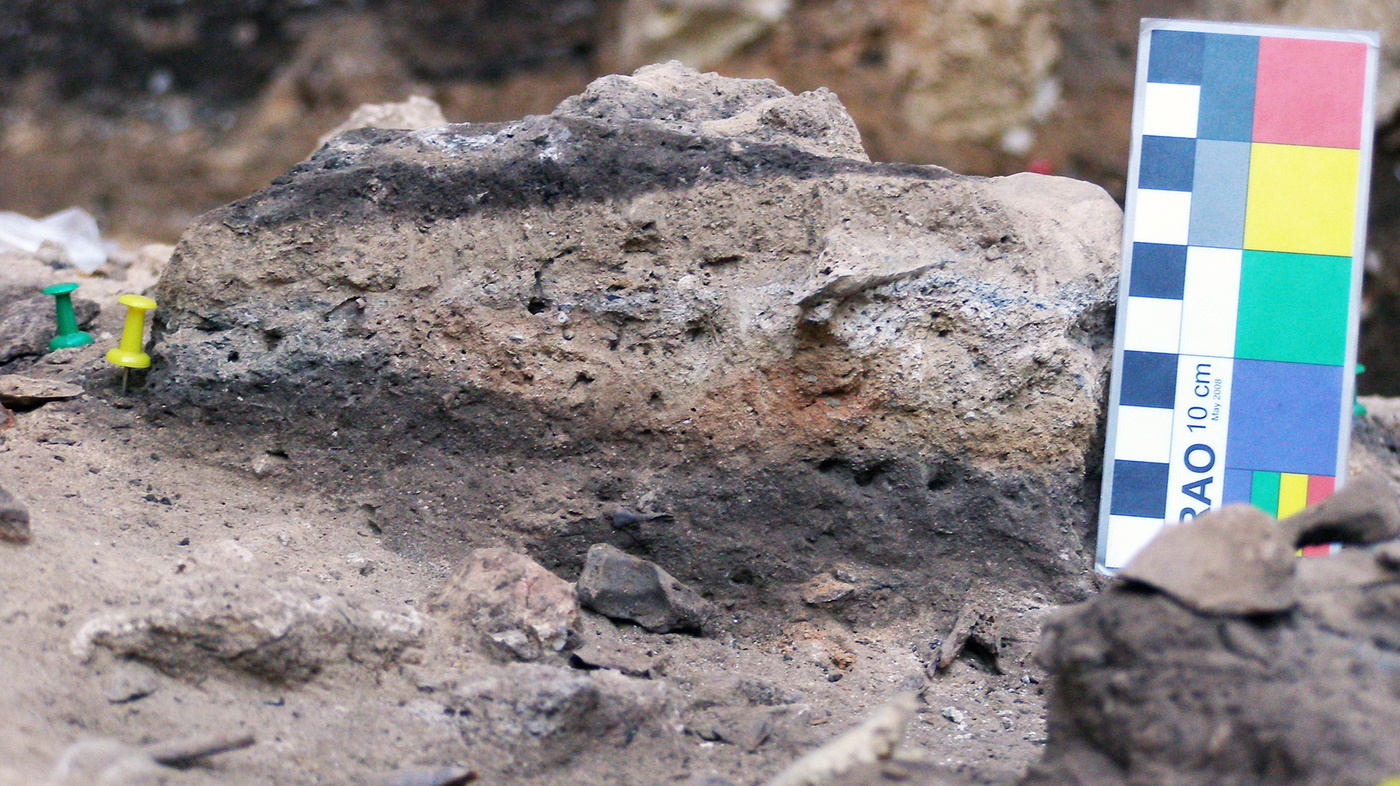
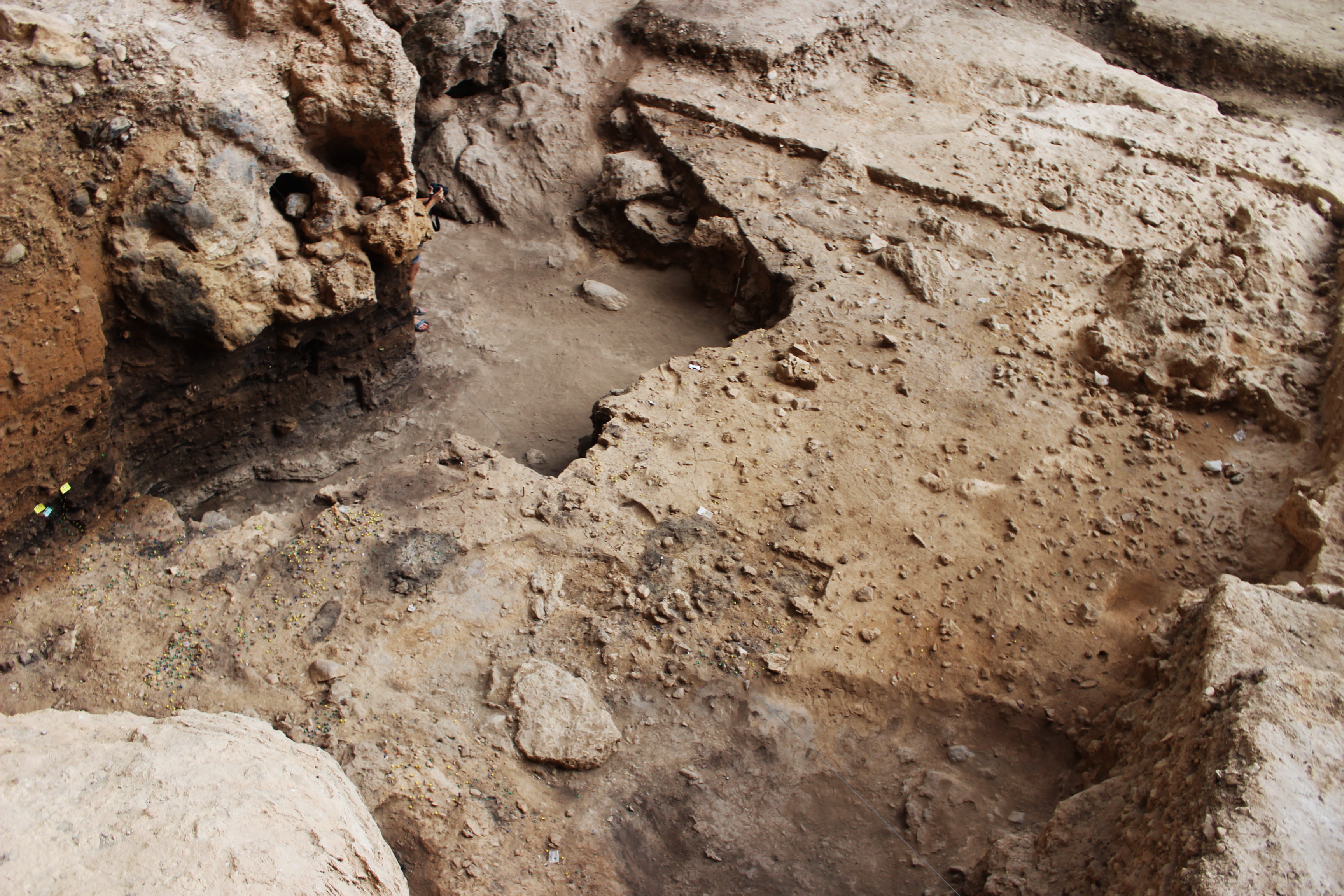
In the early summer of my first postdoctoral position, an email landed in my mailbox with a proposal to attempt to extract microbial aDNA from archaeological sediments deriving from the ash layer of an approximately 50,000 year-old combustion feature (a controlled fire), which contain fecal matter 6. These sediments, found at the open-air archaeological site of El Salt in Alicante, Spain, are curious for many reasons, beginning with the manner of their deposition. The concentration of fecal material was found on the surface of an extinguished fire, which may have been the center point of an ancient campsite. The existence of fecal material in an ancient fire pit, one that had been buried and preserved over thousands of years, prompts speculation about the customs of a people who lived in a familiar region but in nearly unimaginable circumstances.

The human origin of the fecal deposits had already been confirmed in a paper 7 demonstrating the utility of gas chromatography-mass spectrometry to detect diagnostic biomarkers in heterogeneous sediments. The sediments yielded organic residues in high concentrations, specifically coprostanol and 5β-stigmastanol, which is diagnostic of humans. The archaeological context, corresponding to the Middle Paleolithic, suggests a recurrent Neanderthal occupation. Neanderthals are our evolutionary siblings, but went extinct roughly 40,000 years ago. They were not ancestors to our species, but rather they were our contemporaries for a brief time, and sometimes they were even mates. Learning about their biology and society is relevant for us because it helps inform about the evolutionary origins of our own species traits. If traits are shared, then they likely stem from a common ancestor, whereas if traits are unique, then they may indicate a divergence in selective pressures. That email initiated a large collaborative effort among scientists within the disciplines of anthropology, archaeology, and microbiology to probe for the quintessence of the human gut microbiome.
Why would sediment from a Neanderthal “hearth” contain traces of their feces? One scenario is that they burned their waste. It works simultaneously as a source of fuel and a waste management solution, so expedient that we might marvel at this display of intelligent practicality. However, the fecal deposits do not show signs of burning. The sterols (analyzed in the earlier paper) were unaltered, and the nucleic acids (analyzed in the present work) are intact. One marvelous hypothesis suggests that the defecation signifies the abandonment of the fire while keeping the site “preserved” for later occupation – a territorial marking. Evidence of these behaviors are seen at other Middle Paleolithic sites and indicate a shifting and ephemeral use of the space. The fire may have been the central activity point at one time, but in the next occupation, the central area could move to a different space and a new fire established 8. Experiments performed at El Salt suggest that the Pinus nigra wood most often used by Neanderthals reaches a high temperature of ≈ 700º C very quickly, but cannot be maintained for long 9. At that temperature, sterols and DNA disappear, thus the feces were likely deposited after the fire was lit.
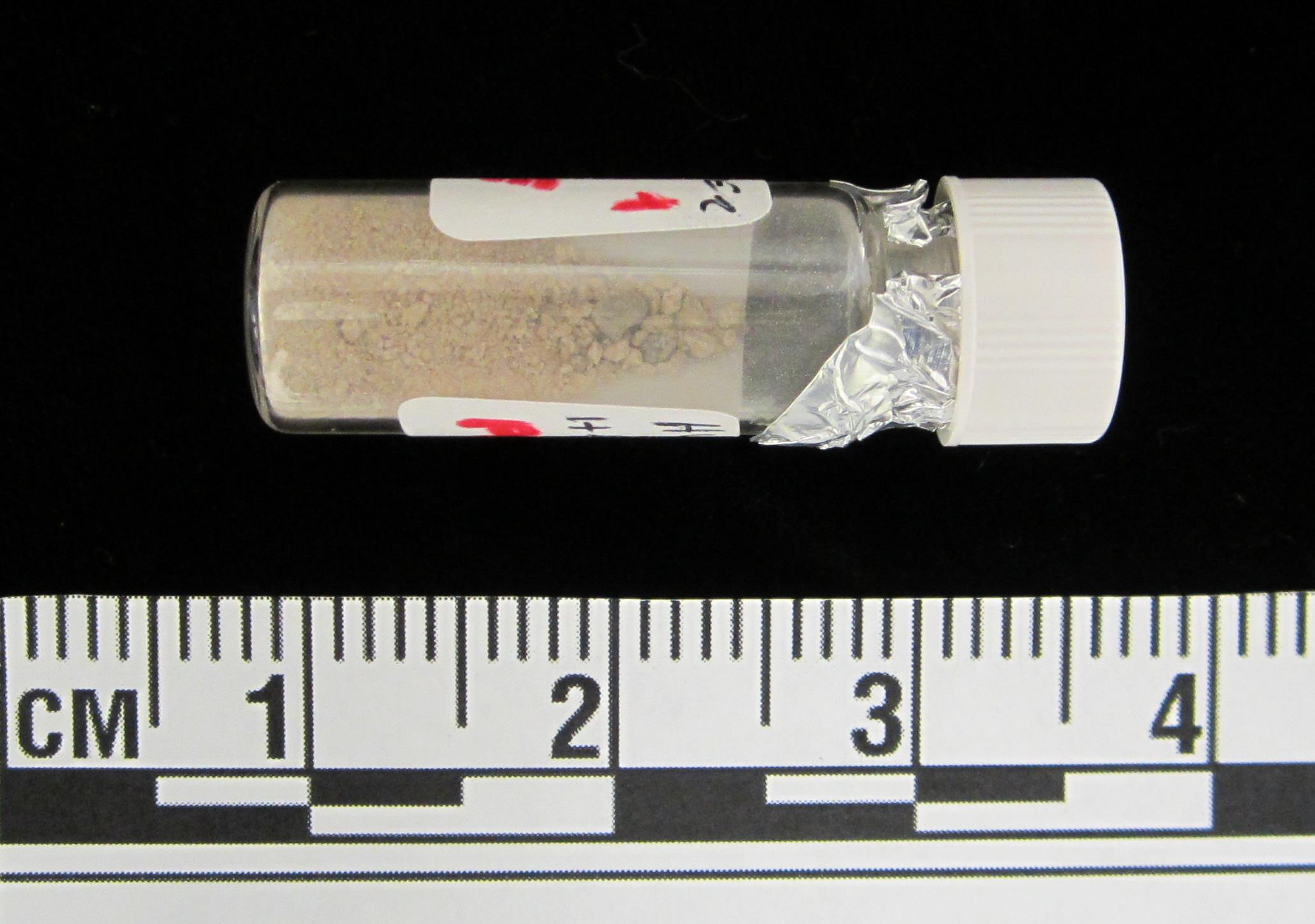

Here was a perfect opportunity to investigate what remained in these fecal time capsules and what more we might learn about these people’s lives. Perhaps a genetic signature of the original microbiome community could be recovered or perhaps other elements referent to their health, diet, and environment were accessible. One unique feature about human fecal matter is that, due to such a high phosphate content, these materials autofluoresce with a green emission when energized by blue light 10. The fecal sediments from El Salt had already demonstrated this property in the earlier publication, but upon receipt of the sample sediments at the Laboratories of Molecular Anthropology and Microbiome Research at the University of Oklahoma, I could not resist to see the autofluorescent propensity for myself. Using the microscope in the clean inner-room of the ancient lab, I mounted a few milligrams of sediment from each sample vial, representing a different layer of the combustion feature (referred to as ES1-ES7 in the report). There, with the blue light and filter applied, the samples fluoresced with an encouraging brilliant green glow.
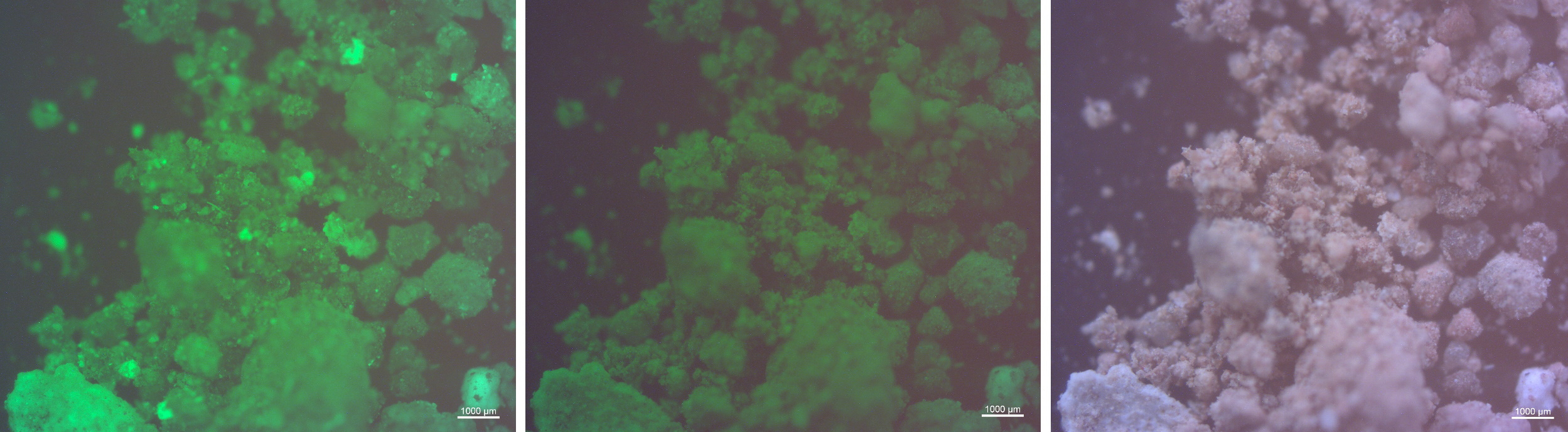

Each benchmark quantification during aDNA recovery – extraction concentrations, amplification cycles, fragment size distribution, and clean blanks – held increasing promise. Furthermore, sediment controls that came from other areas of the archaeological site were largely sterile. It seemed plausible that, despite their age and dispersed heterogeneous context, the tiny fecal deposits in layers ES1-ES7 still contained endogenous genetic material from the gut microbiome of a Neanderthal. After completing bioinformatic analyses, and the meticulous assessments of the degradation patterns that authenticate the ancient nature of genetic sequences, the microbial profiles for the seven sediments presented a “core” hominid gut microbiome (found both in modern human and the African great apes) 11. The other sediment “control” samples from other areas of the excavation site contained only aDNA of taxa well-known in environmental and soil contexts, but never in host mammalian guts. Based on these results, we can anticipate the symbiotic time-depth between humans and some co-resident microbes to be at least one million years. This implies a fixed physiological relationship that is linked to normal development and health in longevity across both humans and Neanderthals as an ancient legacy.

The ability to interrogate the ancient microbiome data relies on the expectations about the human microbiome that are derived from work with modern human and animals gut microbiomes. We can only look for what we already expect to find in a human gut, otherwise we run the risk to over-interpret contamination, but we also lose the chance to identify what may be true novelty in an ancient microbiome. This limitation will remain until more data can be confidently assigned to an ancient host-microbiome. However, it is essential that we put some stakes in the ground and track the human gut microbiome signal where we see it, and then methodically expand the line to incorporate repeatedly associated taxa. The same procedures are needed for demarcating core gene functions. From El Salt we can identify 210 hominid-gut-specific bacterial symbionts in the sedimentary layers associated with Neanderthal occupation, all at the highest detection rate among the combustion feature sediments, ES1-ES7, and all source tracked as primarily gut-microbiome features.
It cannot be ruled out that taxa identified in the El Salt sediments are in fact environmental contaminants, nor can we fully determine their taxonomic assignment due to physical diagenetic processes and limitations with virtual reference data that pertain only to modern bacterial genomes. It may also be that feces from other animals were supplied by the Neanderthals for their fire, since surely these were enterprising and resourceful people. Though, on the basis of the convening evidence – good preservation, lipid biomarker patterns, archaeological context, sediment micromorphology, and now molecular genetic information – the parsimonious picture is that these are indeed a first glimpse of components from a Neanderthal gut microbiome. With new evidence or further analyses, this picture will be open for expansion or revision, but it serves now as a declarative and a comparative data set with which we can forge new intuitions, plan new analyses, and most importantly, look for more opportunities to further explore the ancient human gut microbiome.
Reference: “Components of a Neanderthal gut microbiome recovered from fecal sediments from El Salt” Communications Biology. DOI: 10.1038/s42003-021-01689-y
Works cited:
- Schnorr, S. L., Sankaranarayanan, K., Lewis, C. M. & Warinner, C. Insights into human evolution from ancient and contemporary microbiome studies. Curr. Opin. Genet. Dev. 41, 14–26 (2016).
- Perri, A. R. et al. Dog domestication and the dual dispersal of people and dogs into the Americas. Proc. Natl. Acad. Sci. 118, e2010083118 (2021).
- Stone, A. C. & Ozga, A. T. Ancient DNA in the study of ancient disease. in Ortner’s Identification of Pathological Conditions in Human Skeletal Remains 183–210 (Elsevier, 2019). doi:https://doi.org/10.1016/B978-0-12-809738-0.00008-9
- Tito, R. Y. et al. Insights from characterizing extinct human gut microbiomes. PLoS One 7, (2012).
- Sabin, S. et al. Estimating molecular preservation of the intestinal microbiome via metagenomic analyses of latrine sediments from two medieval cities. Philos. Trans. R. Soc. B Biol. Sci. 375, 20190576 (2020).
- Galván, B. et al. New evidence of early Neanderthal disappearance in the Iberian Peninsula. J. Hum. Evol. 75, 16–27 (2014).
- Sistiaga, A., Mallol, C., Galván, B. & Summons, R. E. The Neanderthal Meal: A New Perspective Using Faecal Biomarkers. PLoS One 9, e101045 (2014).
- Sistiaga, A. Omnivory and human evolution: a new perspective on paleodiets using the faecal biomarker approach. (Universidad de La Laguna, 2015).
- Mallol, C. et al. Human actions performed on simple combustion structures: An experimental approach to the study of Middle Palaeolithic fire. Quat. Int. 315, 3–15 (2013).
- Altemüller, H.-J. & Van Vliet-Lanoe, B. Soil Thin Section Fluorescence Microscopy. in Soil Micro-Morphology: A Basic and Applied Science (ed. Douglas, L. A. B. T.-D. in S. S.) 19, 565–579 (Elsevier, 1990).
- Nishida, A. H. & Ochman, H. A great-ape view of the gut microbiome. Nat. Rev. Genet. 20, 195–206 (2019).
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in