Squeezing Cells for High-Throughput Discovery of Phase Separating Proteins
Published in Bioengineering & Biotechnology, Physics, and Protocols & Methods
Phase separation is a process similar to the separation of oil and water, generating two distinct phases from a single homogeneous mixture. This process also occurs within the cells of living systems. Life regulates various biological events such as signal transduction and gene expression in a temporally and spatially ordered manner through phase separation within cells. Studies have shown that membraneless organelles and biomolecular condensates form gradually through phase separation within cells. Abnormal phase separation processes can lead to the formation of pathological condensates, which are associated with tumor progression and neurodegenerative diseases. Therefore, the identification of phase separation proteins is of significant importance for studying fundamental life processes and identifying new therapeutic targets for diseases. 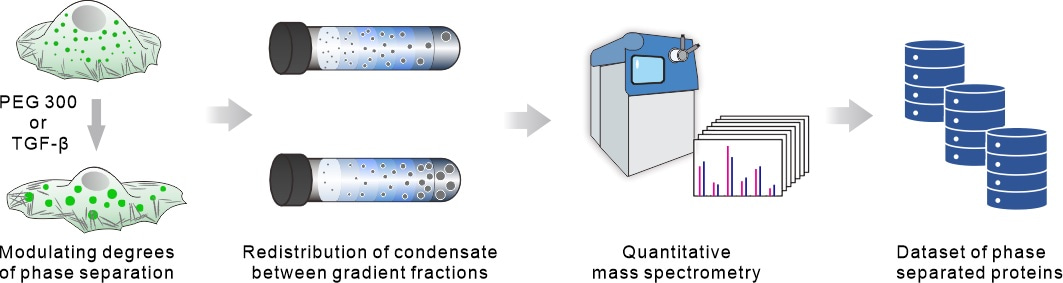
Although hundreds of phase separation proteins have been reported and thousands more have been predicted, existing achievements heavily rely on the purification of proteins in vitro and overexpression systems. Some estimates suggest that 40% of the human proteome consists of phase separation proteins, and it is theorized that any protein that supports multivalency (with three or more binding sites) can undergo phase separation. Given that phase separation behavior is widespread among different types of proteins and highly influenced by the cellular microenvironment, the key biological question lies in determining which endogenous proteins undergo phase separation in which cells, under what microenvironmental conditions, and in response to what stimuli.
To date, there is almost no feasible approach for achieving high-throughput identification of endogenous phase separation proteins at the proteomic level, nor are there effective methods for identifying phase separation proteins in response to biological processes.
The project team focuses on the fundamental behavior of protein phase separation. Due to the abundance of reversible weak physical interactions between protein molecules during phase separation, the broad spectrum of protein phase separation behavior is physically regulated by molecular concentration and molecular crowding. Building on this, the team creatively utilizes cell volume compression to induce extrusion of water from within the cell, leading to an increase in protein concentration and molecular crowding. Proteins within the cell that are undergoing phase separation or are at the edge of phase separation conditions will inevitably alter their degree of phase separation in response to these physical regulatory factors.
To efficiently capture proteins undergoing subtle changes in phase separation degree, the project team combines sucrose density gradient centrifugation and quantitative mass spectrometry techniques. They perform high-throughput identification at the proteomic level of proteins that exhibit changes in oligomerization before and after cell volume compression (1 minute), identifying these proteins as candidate phase separation proteins. Through this method, we have identified 1454 proteins that undergo phase separation under endogenous expression conditions, including 932 known phase separation proteins associated with 11 membraneless organelles, as well as 522 unreported phase separation proteins. By employing the physical cell volume compression method, we have discovered 1215 endogenous phase separation proteins that exhibit mechanical responses.
More importantly, this method can be used to identify proteins undergoing phase separation in biological signaling transduction processes or other biological events. Taking the TGF-β signaling pathway as an example, this method effectively distinguishes phase separation proteins involved in signal transduction processes (75 responsive phase separation proteins within 40 minutes of TGF-β stimulation) from those involved in long-range target gene expression during cell fate transitions (509 responsive phase separation proteins after 2 days of TGF-β stimulation). Its unbiased nature, high throughput, and sensitivity lay a solid foundation for its wide application in the fields of biology and biomedicine. The immunofluorescence staining, protein overexpression in cells, and immunoblotting results of this study all support the accuracy of this method in identifying phase separation proteins.
This research integrates multidisciplinary research approaches including biomechanics, biophysics, biomedical equipment development, and analytical chemistry to rapidly expand existing endogenous phase separation datasets. It provides a new method and perspective for the identification of endogenous phase separation proteins, the discovery of novel organelles, and the search for therapeutic targets in diseases such as cancer.
Professor Yiwei Li focuses on the fundamental biological phenomenon of cell volume and its volumetric compression, seeking to achieve application-oriented breakthroughs from basic theories. Our team pays attention to this common yet overlooked basic life phenomenon, studying its physical molecular mechanisms and its impact on life, regeneration, and therapy. In this work, we discovered that volume compression regulates large-scale protein phase separation behavior within cells (Nature Chemistry 2024). In previous work, Prof. Li reported the kinetics of gene expression regulated by volume compression, and volume compression-induced tumor single cell heterogeneity (PNAS 2021). He also discovered that volume compression stabilizes membraneless organelle formation, affects Wnt signaling pathway transduction, and regulates intestinal organ fate (Cell Stem Cell 2021; Cell Chemical Biology 2023). Additionally, they also reported volume compression's regulation of mature cell reprogramming behavior (Science Advances 2020; PNAS 2019). Based on these fundamental findings. Prof. Li's team further combines mechanics design of volume compression with molecular signal transduction to achieve the construction of functional microtissues and organoids, including fat, tumors, muscles, and intestines, leading to industrial transformation.
His team can be viewed: www.yiweililab.com
His publication profile can be found: https://scholar.google.com/citations?user=mihILDoAAAAJ&hl=en
Follow the Topic
-
Nature Chemistry
A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in