Towards a better description of the interaction of enterotoxigenic Escherichia coli with intestinal mucus
Published in Microbiology
Context : At the interface between the human digestive lumen and the host, the mucus layer which is secreted by goblet cells, constitutes both a physical barrier against the entry of pathogens and the reservoir of a specific commensal microbiota1,2. Notably, enteric pathogens have to interact with and penetrate this line of defense in order to colonize the intestinal epithelium. Among enteric pathogens, enterotoxigenic Escherichia coli (ETEC), the main agent responsible for travelers’ diarrhea, is equipped with an arsenal of adhesins and mucinases, allowing respectively adhesion to intestinal epithelial cells and mucus degradation3.
Objectives and methods: The interactions between intestinal mucus, human gut microbiota and ETEC in the virulence of the pathogen remain poorly studied. Using a set of complementary in vitro approaches simulating the human digestive lumen or the intestinal epithelium, this work describes how the intestinal mucus compartment can shape the survival of the reference strain ETEC H10407, its adhesion to intestinal cells, the expression of its virulence genes, the induction of a pro-inflammatory response (interleukin-8) and its interaction with a gut microbiota of human origin.
A focus on the newly developed TIM-1 model to assess ETEC survival and virulence. Regionalized and dynamic in vitro gut models which capture the complexity of the human gastrointestinal physiology are valuable tools to investigate ETEC behavior in human digestive conditions. Among the available systems, the multi-compartmental and computer-controlled TNO gastrointestinal model (TIM-1) is currently considered as the most complete simulator of the upper gastrointestinal tract by simulating the main physicochemical parameters of the human digestion4,5 (Figure 1). In the study from Sauvaitre et al., this model was adapted for the first time to simulate the mucus environment by two means: addition of mucus secretion (final concentration of 3 g.L-1 throughout the digestive tract to mimic mucus shedding) and addition of two polyester pockets containing 40 beads of mucin- alginate beads placed in each of the four compartments of TIM-1 to provide physical surface.
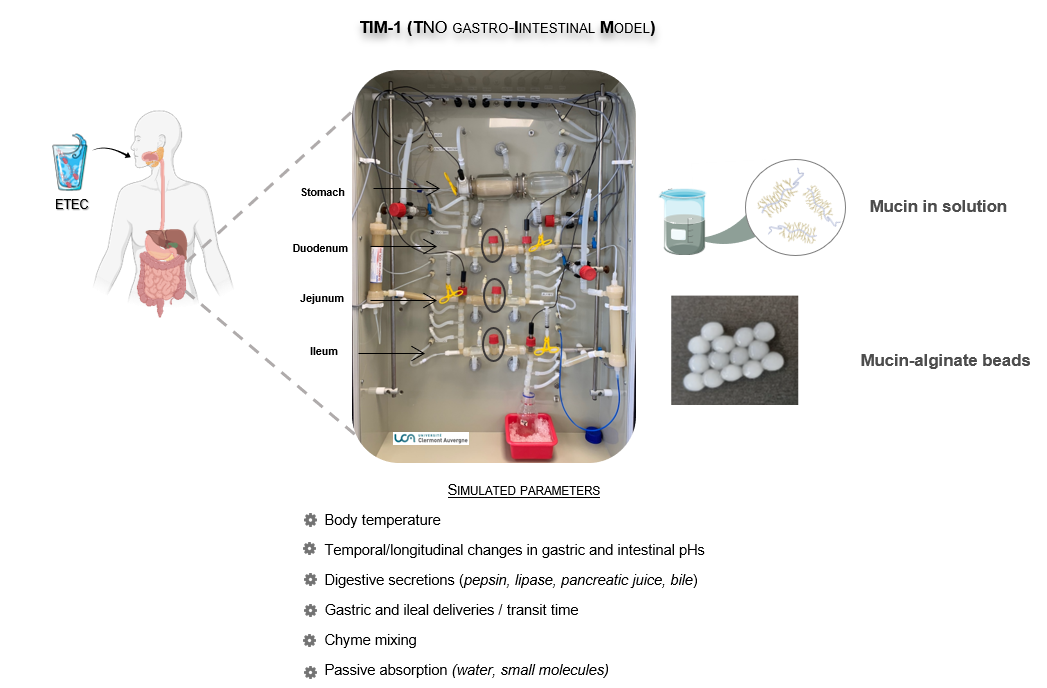
Figure 1. Overview of TIM-1 system (TNO gastro-Intestinal tract Model 1) used to investigate ETEC strain H10407 survival and regulation of virulence genes under human simulated digestive conditions.
This model enables to reproduce the physicochemical conditions encountered in the lumen of the human stomach and 3 parts of the small intestine (duodenum, jejunum and ileum). In this study, mucin in solution (as mucus secretion) and mucin-alginate beads (as a physical mucus surface) were added for the first time.
Results. Using the TIM-1, we demonstrate that the mucus compartment favors the survival of ETEC H10407, probably by providing a niche allowing the pathogen to maintain itself under stringent human digestive conditions. By integrating the host part through the use of the Caco2 (enterocytes) / HT29-MTX (mucus cells) co-culture model, we demonstrate that the presence of mucus promotes ETEC adhesion and modulate the expression of main virulence genes. When incorporating a highly complex gut microbiota from human origin in batch fermentation experiments, we demonstrate that the addition of mucus does not promote ETEC colonization.
Using this set of complementary in vitro approaches, our results suggest that the presence of a mucus niche promotes the survival of ETEC in the lumen of the human upper digestive tract and their adhesion to intestinal epithelial cells, suggesting an increased colonization capacity. This study highlights the major role played by mucus in the pathophysiology of ETEC infections, thus paving the way for the development of new strategies against these infections specifically targeting the interaction with mucus.
REFERENCES
- Sauvaitre, T. et al. Tripartite relationship between gut microbiota, intestinal mucus and dietary fibers: towards preventive strategies against enteric infections. FEMS Microbiol Rev 45, fuaa052 (2021).
- Etienne-Mesmin, L. et al. Experimental models to study intestinal microbes-mucus interactions in health and disease. FEMS Microbiol Rev 43, 457–489 (2019).
- Roussel, C., Sivignon, A., de Wiele, T. V. & Blanquet-Diot, S. Foodborne enterotoxigenic Escherichia coli: from gut pathogenesis to new preventive strategies involving probiotics. Future Microbiol 12, 73–93 (2017).
- Roussel, C. et al. Multi-targeted properties of the probiotic Saccharomyces cerevisiae CNCM I-3856 against enterotoxigenic Escherichia coli (ETEC) H10407 pathogenesis across human gut models. Gut Microbes 13, 1953246.
- Cordonnier, C. et al. Dynamic In Vitro Models of the Human Gastrointestinal Tract as Relevant Tools to Assess the Survival of Probiotic Strains and Their Interactions with Gut Microbiota. Microorganisms 3, 725–745 (2015).
Cover illustration : Josefien Van Landuyt
Follow the Topic
-
npj Biofilms and Microbiomes
The aim of this journal is to serve as a comprehensive platform to promote biofilms and microbiomes research across a wide spectrum of scientific disciplines.
Related Collections
With Collections, you can get published faster and increase your visibility.
Microbial endocrinology
Publishing Model: Open Access
Deadline: Oct 21, 2026
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in