Validated workflows for preparing and characterizing core‑stained and surface‑labeled fluorescent polymer particles with simple commercial automation tools
Published in Bioengineering & Biotechnology, Chemistry, and Materials
Explore the Research
Validated workflows for preparing and characterizing core-stained and surface-labeled fluorescent polymer particles with simple commercial automation tools
The problem: A pipetting bottleneck in nanomaterial preparation
Nanometer- and micrometer sized fluorescent polymer particles (PSP) are workhorses of modern life sciences. They serve as reporters in bioanalytical assays, enable the sensitive detection of neutral and ionic analytes in sensing platforms, and are used as calibration standards for fluorescence microscopes and flowcytometry. Nowadays, many fluorescent PSPs are still prepared by manual swelling of premanufactured PSP with organic solvents, which is time-consuming and labor-intensive, and can provide operator dependent results.
The routine workflow for the preparation of fluorescent PSP involves the following steps: add PSP and organic swelling solvent, add hydrophobic dye, wait for about 1h, centrifuge the PSP off, wash the PSP with ethanol to remove excess dye, and repeat these centrifugation-washing cycles 3-5 times. In addition to being time-consuming and labor-intensive, this procedure can provide operator dependent results. In our lab, we kept running into the same drawbacks: two researchers performing exactly the same protocol and ending up with fluorescent PSP batches that could substantially differ in dye loading efficiency, fluorescence brightness, and particles recovery. When comparing the results across four operators, PSP dye loading efficiencies varied by almost a factor of two. This batch-to-batch variability can hamper the comparability and reproducibility of intended fluorescent PSP applications.
Our approach: A pipetting robot off the bench
To ease fluorescent PSP reproducibility and accelerate fluorescent PSP preparation, we developed a simple automation workflow. Rather than building an expensive customized automated platform, we asked a simple question: Could we use a conventional, commercial, self-programmable, pipetting robot, utilized in many bioanalytical and analytical laboratories, to automate the key liquid handling steps of particle staining and surface labeling?
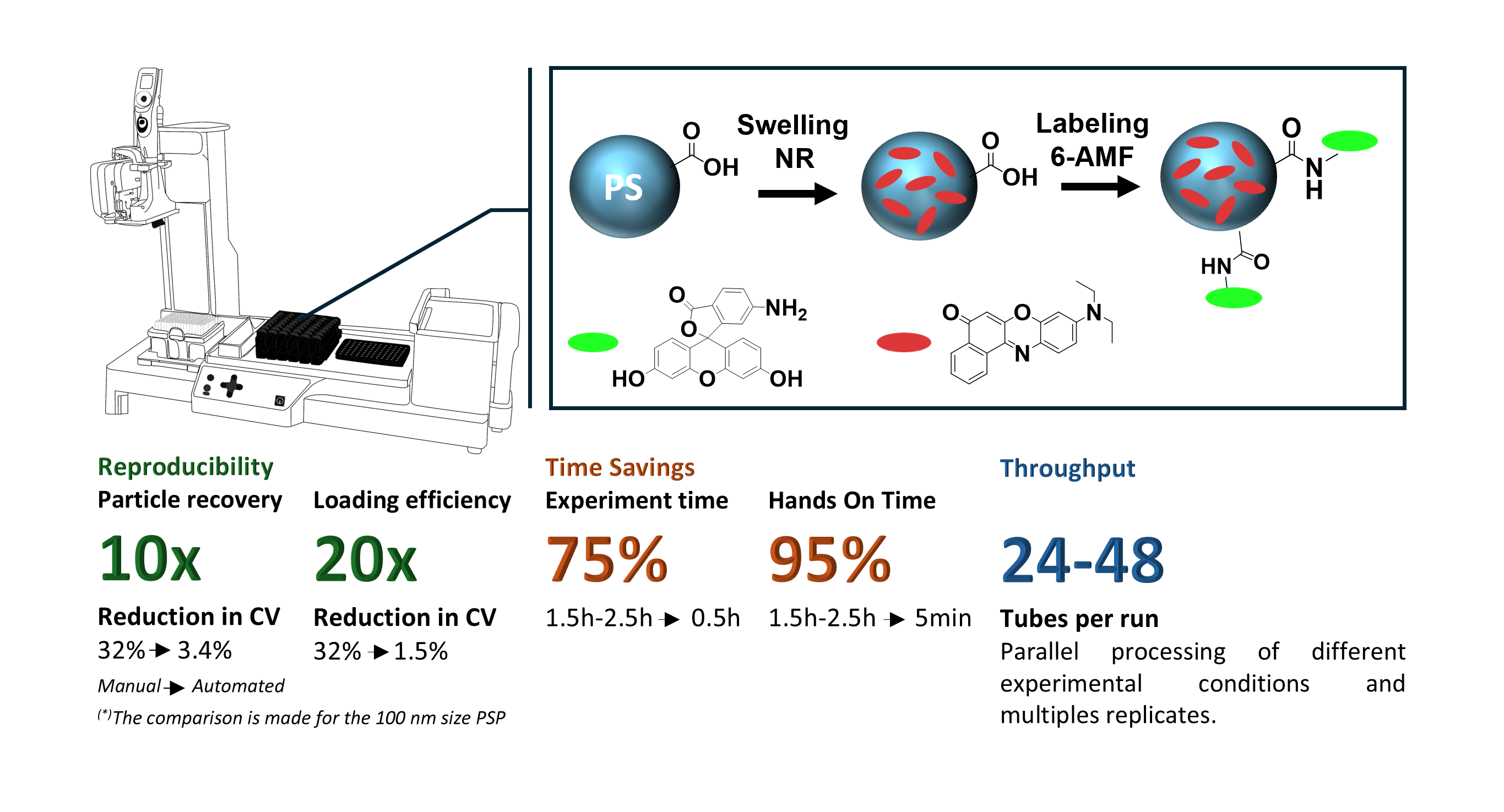
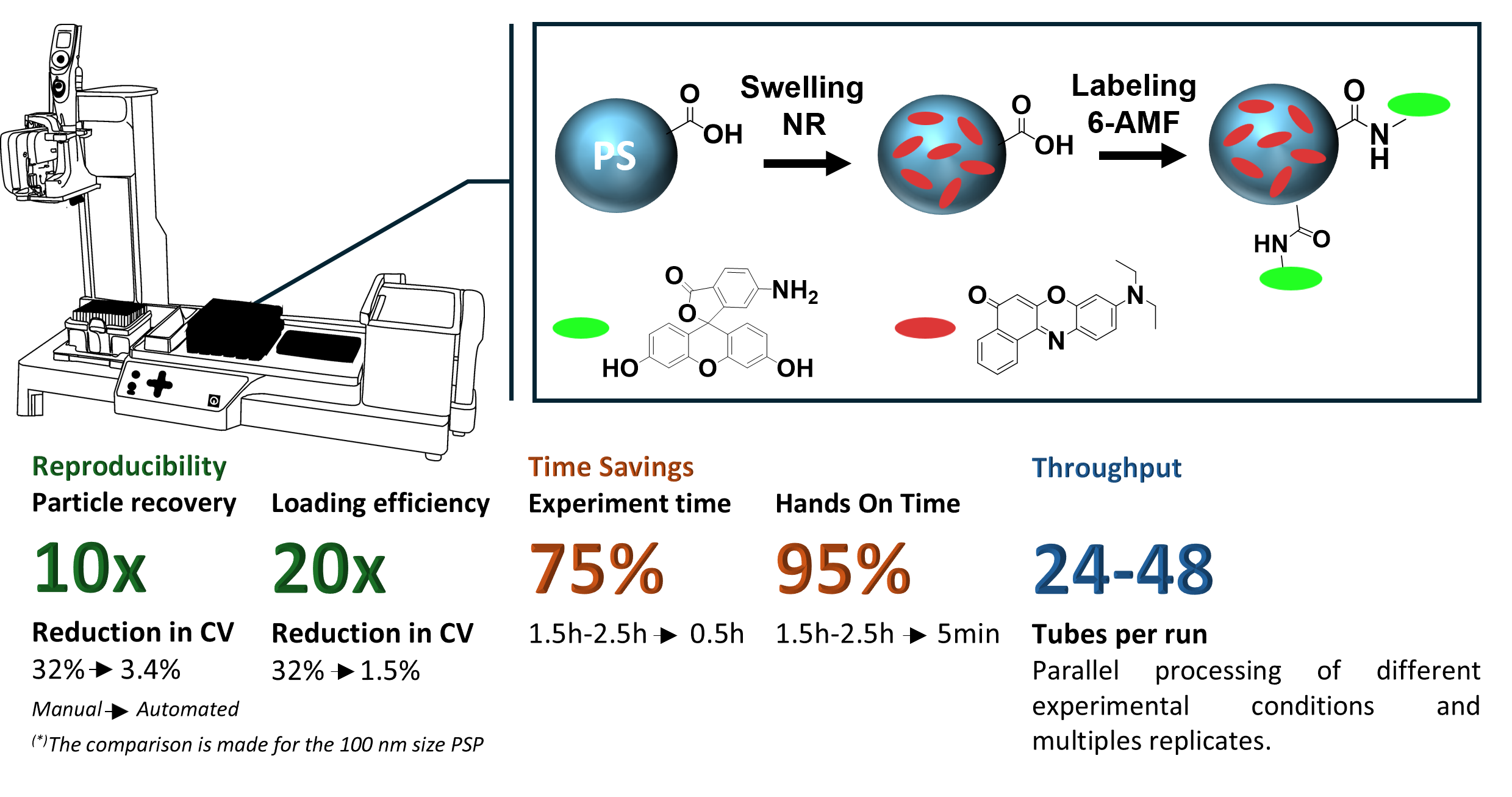
We started with Nile Red (NR) as our dye workhorse and loaded this hydrophobic dye into carboxylated PSP via an already established swelling procedure. First, we optimized the loading workflow for 200 nm PSP and then adapted this workflow and validated for 100 nm and 1000 nm PSP. Thereby, the pipetting robot was programmed to dispense water, add an aqueous PSP dispersion, introduce the dye dissolved in a tetrahydrofuran (THF): N,N-dimethylformamide (DMF) mixture, and add water to terminate the swelling process after incubation, with all steps being performed with precisely controlled timing, volumes, and speed, followed by washing and redispersion steps.
We then expanded this workflow to PSP surface labeling, shown exemplary by covalently attaching pH-sensitive 6-aminofluorescein (6-AMF) onto carboxylated PSP and fluorescein isothiocyanate (FITC) onto aminated PSP, both previously loaded with Nile Red. This allowed us to fabricate ratiometric pH-sensor beads in a single automated workflow, core-stained with the red emissive and pH-inert reference dye Nile Red and surface-labeled with a green, fluorescent pH-sensor dye.
 What we found
What we found
The results of our automated fluorescent PSP preparation workflows are striking. The variation of the particle recovery rate was reduced tenfold from 32% (manual) to just 3.4%. The dye loading efficiency, reached 66% ± 2% using automation, compared to 32% ± 12% for the manual approach. Equally compelling was the time saving and the reduced hands-on-time. The automated pipetting steps for the loading of NR into the PSP were completed in less than 30 minutes compared to 1.5 to 2.5 hours for the manually performed steps even by well trained staff.
The 6-AMF surface labeling workflow was equally well reproducible. The mean labeling concentration across five independently prepared sensor bead batches was 31 ± 4 nmol, confirming that the automated two-step workflow (core staining followed by surface conjugation), can be reliably executed in a single programmed sequence. The same concept was demonstrated for FITC labeling of aminated PSP, showing that the automation approach can be utilized for different common conjugation chemistries.
Why it matters
This work is deliberately designed for being easy to follow. Therefore, we utilized a broadly accessible pipetting robot increasingly employed in many labs instead of an expensive custom build robotic platform. All tools are commercially available, affordable, and can be easily programmed even by non-expert users.
The broader impact of the presented automated and validated workflows for the preparation of fluorophore-loaded and labeled PSP extends well beyond our laboratory. Reproducible, well characterized fluorescent particles are required for fluorescent assays, bioimaging, sensing studies and calibration tools for fluorescence methods in the life and material sciences.
What comes next
These workflows are not limited to polymer particles such as polystyrene particles or the exemplary assessed dyes. The underlying chemistry of swelling based dye encapsulation and reactive surface conjugation is widely applicable to organic and inorganic particles of different composition and different surface functionalities. We are currently exploring extensions of easily integrated automation concepts in workflows involved in the production and characterization of functional nanomaterials for photonics, imaging, and sensing applications.
We hope this work encourages more researchers to take the step towards automation, not as a distant aspiration, but as a practical, accessible improvement to everyday nanomaterial workflows.
Follow the Topic
-
Analytical and Bioanalytical Chemistry
Analytical and Bioanalytical Chemistry (ABC) is the only journal with global visibility and the mission to rapidly publish excellent and high-impact contributions on fundamental and applied topics of (bio-)analytical research that is supported by a large group of learned societies around the world.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in