Bringing radiotherapy target segmentation into BraTS with the BraTS-MEN-RT dataset
Published in Healthcare & Nursing, Cancer, and Computational Sciences
Radiotherapy planning begins with a simple question: where is the disease we need to treat? In modern radiation oncology, that question is answered by segmentation, the careful delineation of structures on medical imaging for dose calculation and plan optimization. Over the past decade, segmentation has benefited substantially from automation, particularly for normal structures and organs at risk. Commercial and vendor-supported tools now routinely generate high-quality contours for anatomy that is relatively consistent across patients, such as the brain, brainstem, optic apparatus, and cochleae. In contrast, automated segmentation remains far less common for gross tumor volumes, and the gap is widest where the clinical need is often greatest: postoperative disease.
That imbalance was a central motivation for building the 2024 Brain Tumor Segmentation Challenge Meningioma Radiotherapy dataset, known as BraTS-MEN-RT, and for introducing a radiotherapy-focused automated segmentation challenge within the 2024 BraTS Cluster of Challenges, with continued benchmarking through the MICCAI 2025 Lighthouse Challenge. The BraTS challenges have a long history of accelerating innovation by pairing well-curated datasets with transparent evaluation. Until recently, however, much of the work in automated brain tumor segmentation has focused on diagnostic-style imaging conventions and preprocessing strategies that are well suited for research pipelines, but less aligned with radiotherapy treatment planning workflows.
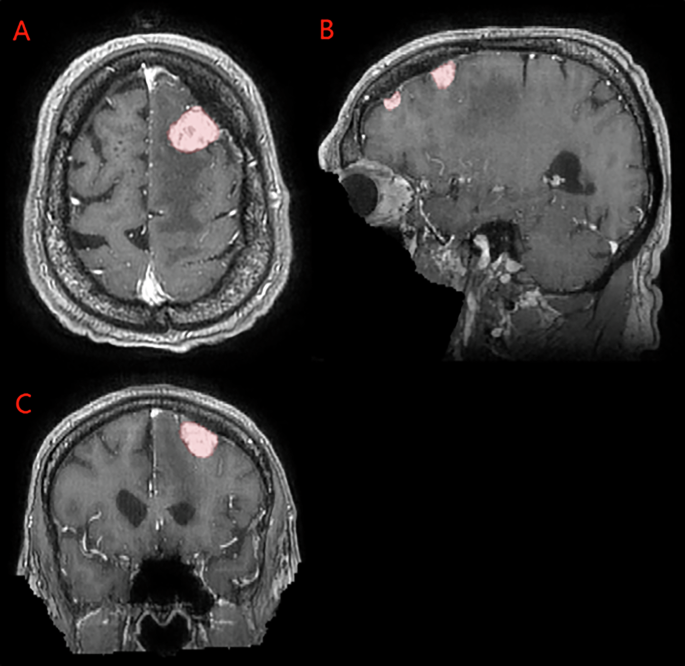
The difficulty of gross tumor volume segmentation compared with organ-at-risk contouring is not surprising. Tumor appearance varies with histology, location, size, prior interventions, and imaging technique. Postoperative target delineation adds another layer of complexity because the object of interest may have a combination of residual enhancing disease, surgical cavity, postoperative change, and regions at risk that require expert interpretation. Even when clinicians agree on general principles, postoperative findings can be heterogeneous and imaging may be distorted by surgical materials or fixation devices. In meningioma specifically, targets often include both residual enhancing tumor and postoperative tumor bed, with nuanced judgment calls regarding dural tails and enhancement at resection margins.
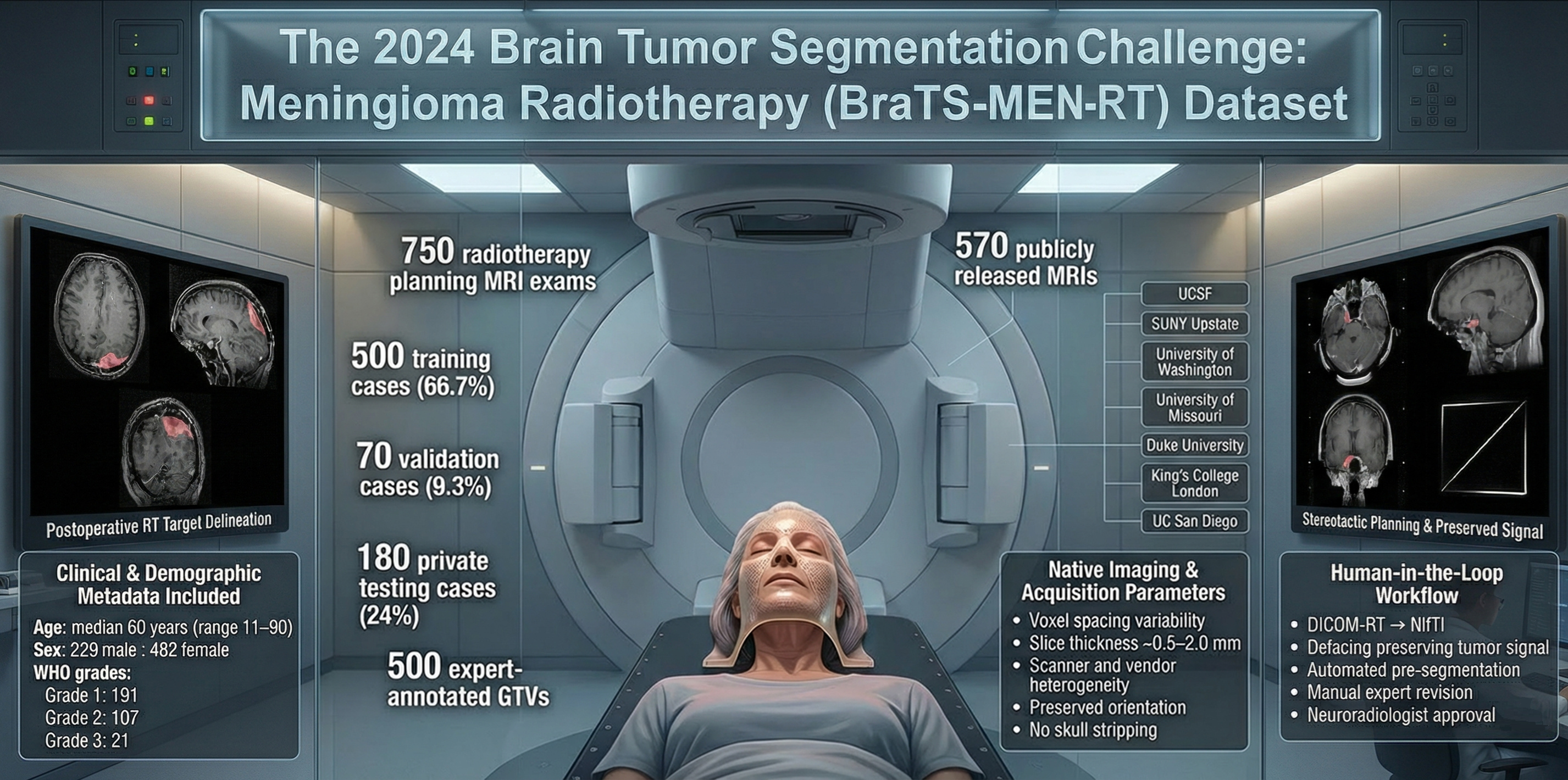
This is precisely where convenient research preprocessing can become a liability. Skull stripping, rigid atlas registration, and isotropic resampling simplify many computer vision tasks, but they can remove contextual information that radiotherapy depends on. For stereotactic radiosurgery cases, the presence of headframes and localizer fiducials is not a nuisance artifact; it is part of the clinical imaging environment and can meaningfully affect both image appearance and the usability of any deployed model. One of the core design choices of BraTS-MEN-RT was therefore to preserve the native spatial resolution and orientation of radiotherapy planning 3D T1-weighted post-contrast MRIs. Skull stripping was intentionally avoided to retain clinically relevant extracranial anatomy, while defacing was applied to remove identifying facial features without compromising the planning-relevant volume.
Building the dataset required assembling enough high-quality, consistently defined target volumes to support meaningful training and benchmarking. BraTS-MEN-RT includes radiotherapy planning MRIs from seven academic centers in the United States and the United Kingdom, including Duke University, UC San Francisco, SUNY Upstate, the University of Washington, the University of Missouri, King’s College London, and UC San Diego. In total, 750 planning MRI exams were collected and divided into training, validation, and a private testing partition. Of these, 570 cases form the publicly accessible image set, and 500 include expert gross tumor volume segmentations for model development. The private test set was deliberately retained to enable unbiased evaluation as this radiotherapy track continues in future challenges.
While the images and labels are the most visible product, the real story lies in the human effort behind them. The contours in BraTS-MEN-RT are grounded in clinical care. They represent the cumulative work of radiation oncologists, neuroradiologists, and collaborating clinicians across institutions, performed under the time constraints and complexity of routine practice. Collectively, this foundational work amounts to thousands of hours spent delineating gross tumor volumes in intact and postoperative meningioma cases over many years. Transforming those clinical contours into a standardized, research-ready resource then required hundreds of additional hours devoted to harmonizing definitions, performing meticulous slice-by-slice review, correcting inconsistencies, and ensuring adherence to a shared radiotherapy-focused protocol. Small differences in target definition can meaningfully alter treatment volumes, and those differences matter not only for clinical interpretation and treatment delivery, but also for training and evaluating automated models.
Quality assurance was equally time intensive, particularly around de-identification. In neuroimaging, defacing is a common strategy to reduce re-identification risk while preserving intracranial anatomy. In radiotherapy planning, the stakes are higher because tumors can extend toward the skull base or extracranially, and aggressive anonymization risks removing clinically relevant tissue. For this reason, every defaced volume underwent careful manual review to ensure that intracranial anatomy and tumor signal were preserved. When partial tumor removal occurred, cases were either manually corrected or excluded if preservation was not feasible. These decisions rarely make headlines, but they are decisive for whether a shared dataset is truly usable.
An equally important but less visible component of the dataset is its metadata. BraTS-MEN-RT includes clinical and demographic information such as patient age, sex, and WHO tumor grade when available, along with imaging metadata describing acquisition parameters and native resolution. Imaging parameters vary widely across scanners, vendors, and institutions, and full documentation was not mandated as a condition of data contribution. This reflects the reality of multi-institutional radiotherapy practice and lowers barriers to data sharing. From a modeling perspective, the metadata enables researchers to study how algorithm performance varies with resolution, slice thickness, and patient or tumor characteristics, while also forcing models to generalize across heterogeneous acquisition conditions rather than relying on narrowly standardized inputs. For radiotherapy, where robustness across institutions is essential, this variability is a feature rather than a flaw.
Another critical part of the story is governance. Multi-institutional data sharing is not just a technical challenge, but a legal and ethical one. Institutional review boards, privacy officers, compliance teams, and legal groups all contribute to evaluating pathways for de-identification and public release. Data use agreements and repository requirements must align to ensure that the scientific goal of open sharing is met without compromising patient privacy. Many individuals involved in this process are never listed as authors, yet the dataset would not exist without their work.
Why does all of this matter? Because automated segmentation for radiotherapy targets will not improve without datasets that faithfully represent radiotherapy practice. Industry efforts have understandably focused on organs at risk, where ground truth is easier to define and variability is limited. Yet the clinical bottleneck often remains the target itself, particularly in postoperative settings where interpretation is nuanced and variability is high. If we want models that meaningfully assist radiation oncologists, we need resources that capture postoperative complexity, acquisition heterogeneity, and real planning context. The BraTS challenge framework provides a natural home for this kind of benchmarking, and the addition of a radiotherapy-focused meningioma segmentation track is a step toward making radiotherapy targets a more central focus in medical image analysis. When data and metadata are shared, researchers can reproduce results, compare methods, and build on each other’s work rather than repeatedly solving access problems in parallel. Over time, this collaborative approach can produce tools that reduce avoidable manual burden, improve consistency, and allow clinicians to focus their expertise where it is most needed.
The scientific data manuscript, “The 2024 Brain Tumor Segmentation Challenge Meningioma Radiotherapy (BraTS-MEN-RT) dataset,” (https://doi.org/10.1038/s41597-026-06649-x) details the cohort, annotation protocol, metadata, and quality control pipeline, and the dataset is publicly available via Synapse (https://doi.org/10.7303/syn59059779), with a private held-out test set supporting ongoing challenge-based evaluation.
Follow the Topic
-
Scientific Data
A peer-reviewed, open-access journal for descriptions of datasets, and research that advances the sharing and reuse of scientific data.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics in freshwater and marine science
Publishing Model: Open Access
Deadline: Jul 23, 2026
Genomes of endangered species
Publishing Model: Open Access
Deadline: Jul 01, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in