The way scientists unravel biomedical questions has been fundamentally transformed by the CRISPR/Cas9 genome editing system. Consequently, there is a lot of excitement about the new opportunities that CRISPR editing brings for biomedical research and molecular medicine. In the PaquetLab, we apply this technique for precise editing of induced pluripotent stem cells (iPSCs) to generate novel in vitro models of human neurodegenerative and neurovascular diseases. We believe that with the help of CRISPR, such models will not only allow to better understand human disease but also to develop much-needed drugs for treatments.
Despite the great potential of the CRISPR system, it is important to acknowledge that genome editing may be accompanied by collateral damage1,2. If editing with CRISPR goes awry, the consequences could be massive, ranging from wasted funding over misleading research results to incorrectly edited cells for replacement therapies. With this in mind, we examined our successfully edited cell lines thoroughly and were surprised to see that many were affected by inadvertent alterations at the targeted locus3. These so-called on-target effects (OnTE) had been identified previously in genome-edited mice 4–7, but it was unclear if clinically relevant human cells, such as iPSCs, would also be affected. Furthermore, it had not been investigated if OnTE also occur in cells edited by the homology-directed-repair (HDR) pathway, which is widely exploited in the field to insert specific mutations or correct mutant cells back to wildtype. Both aspects also have important implications for CRISPR therapies in patients.
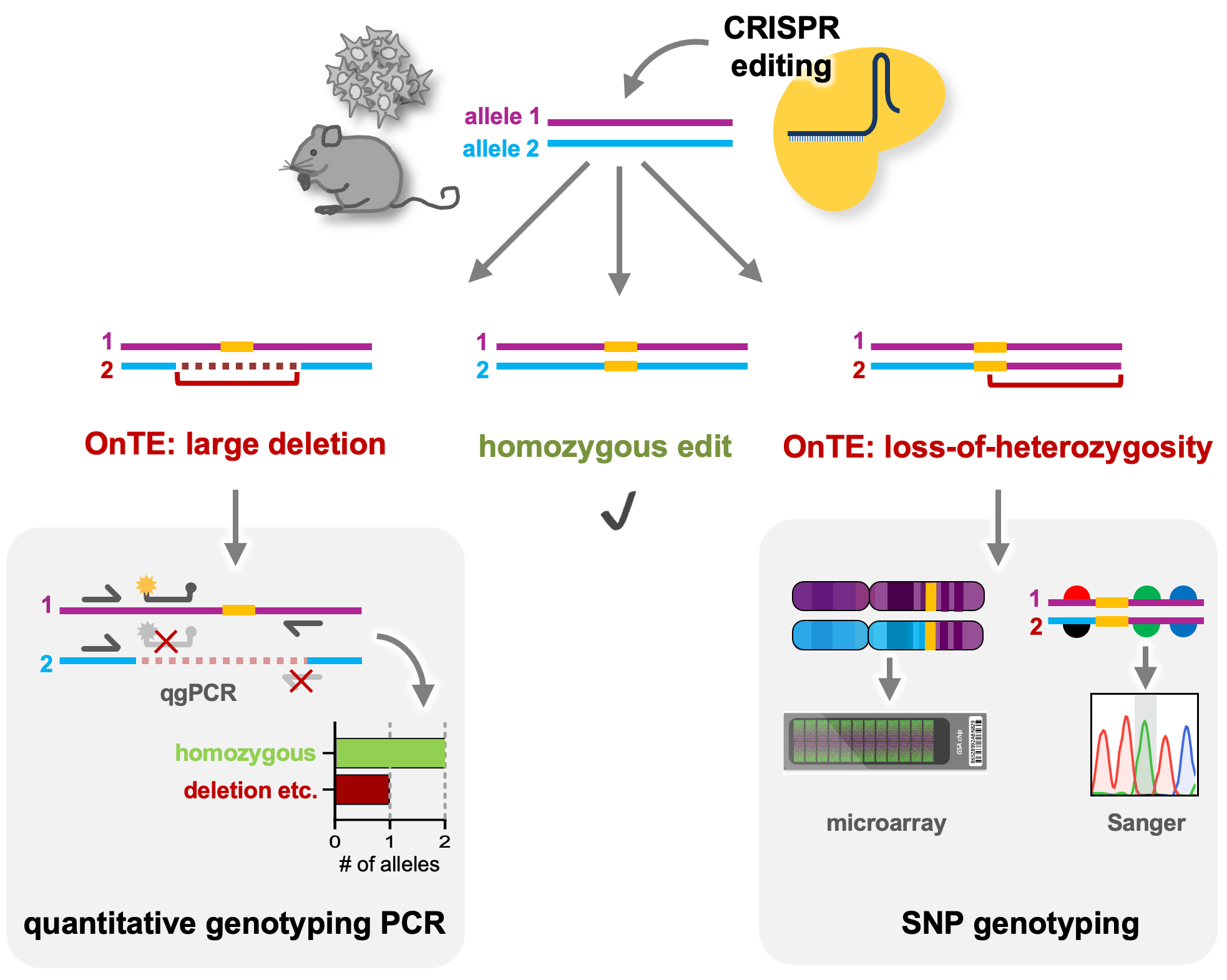
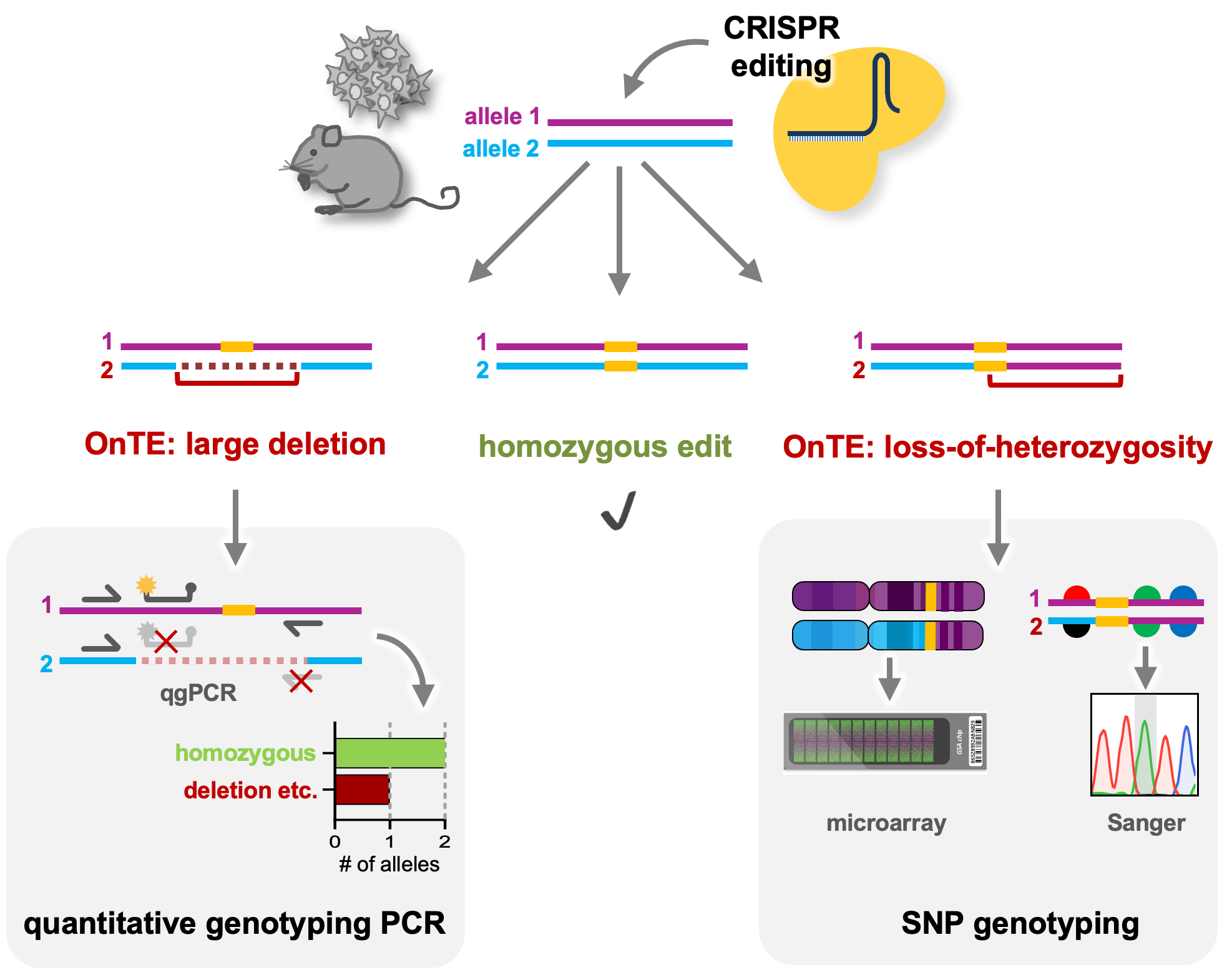
Figure 1: Schematic of OnTE detection after CRISPR editing: quantitative genotyping PCR can detect large deletions, insertions or complex rearrangements by identifying the number of intact alleles at the edited locus. SNP genotyping either by nearby SNP sequencing or genome-wide analysis by SNP microarrays, can identify regions of loss-of-heterozygosity.
The OnTE we identified in our cell lines included mono-allelic large deletions, insertions, complex rearrangements or regions of loss-of-heterozygosity (LOH) around the target site. Although such changes can be very drastic, they are often invisible in standard quality controls such as Sanger genotyping. If undetected, such OnTE can have deleterious effects on gene expression, leading to data misinterpretation with the abovementioned negative consequences on study reliability etc. To understand the extent of the problem, we investigated the occurrence of OnTE at several loci and in several cell lines, which revealed that OnTE are a universal problem in human iPSCs3. This clearly highlighted the need for thorough on-target quality control analysis after genome editing. However, most current CRISPR-based studies lack such measures, which is likely because accessible detection tools were missing. We therefore developed broadly applicable tools for on-target analysis. We believe these will be a valuable resource for the genome editing field for several reasons:
Our protocol
- is available to all CRISPR users as it can be implemented solely based on standard molecular biology techniques such as PCR amplification, gel electrophoresis, Sanger sequencing and quantitative PCR. We also provide an alternative method for LOH detection that enables genome-wide analysis but requires specialized equipment.
- is applicable to a variety of cell lines and edited organisms.
- works for NHEJ- as well as HDR-edited cells/organisms.
Taken together, OnTEs occur frequently after CRISPR editing and their exclusion is highly important to ensure reliability of findings obtained with edited lines or organisms. We hope that our detailed workflow will allow simple and reliable OnTE detection for many people in the field, and that these additional measures will improve the reliability of CRISPR-based research and its application in molecular therapies.
References
- Thomas, M., Burgio, G., Adams, D. J. & Iyer, V. Collateral damage and CRISPR genome editing. PLoS Genetics 15, (2019).
- Burgio, G. & Teboul, L. Anticipating and Identifying Collateral Damage in Genome Editing. Trends in Genetics vol. 36 905–914 (2020).
- Weisheit, I. et al. Detection of Deleterious On-Target Effects after HDR-Mediated CRISPR Editing. Cell Reports 31, (2020).
- Adikusuma, F. et al. Large deletions induced by Cas9 cleavage. Nature 560, (2018).
- Owens, D. D. G. et al. Microhomologies are prevalent at Cas9-induced larger deletions. Nucleic acids research 47, 7402–7417 (2019).
- Shin, H. Y. et al. CRISPR/Cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. Nature Communications 8, (2017).
- Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nature Biotechnology 36, (2018).
Lab webpage: https://www.isd-research.de/paquetlab
Lab Twitter: https://twitter.com/PaquetLab
Follow the Topic
-
Nature Protocols
This journal publishes secondary research articles and covers new techniques and technologies, as well as established methods, used in all fields of the biological, chemical and clinical sciences.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in