Engineering an efficient and enantioselective enzyme for the Morita–Baylis–Hillman reaction
Published in Chemistry
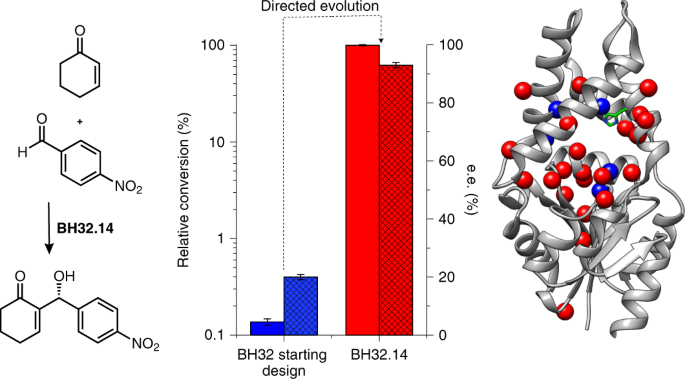
In recent years the biocatalytic toolbox has expanded significantly, allowing for a broader scope of chemical transformations. The emergence of directed evolution has played a significant role as a powerful tool to tailor natural enzymes towards desired transformations. However, there are some catalytic functions that are not found in Nature’s biosynthetic repertoire, leaving gaps in the range of chemistries available to biocatalysis. To address this limitation, computational design could be seen as an attractive and versatile alternative. This approach has provided primitive biocatalysts for a small number of well-characterised transformations that have been previously achieved with catalytic antibodies.1-3 In some cases, designs have been optimised with directed evolution affording biocatalysts approaching efficiencies of natural enzymes.4-6 For this project, we were eager to tackle more mechanistically complex and multi-step chemical transformations not previously achieved with catalytic antibodies, and for which no effective protein catalysts are known. We decided to subject the primitive de novo enzyme BH32,7 designed to promote the Morita-Baylis-Hillman (MBH) reaction (Figure 1), to directed evolution to deliver the first efficient and selective biocatalyst for this transformation.
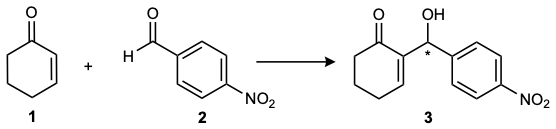
Figure 1: The Morita-Baylis-Hillman reaction. Chemical scheme of the MBH reaction between 2-cyclohexen-1-one 1 and 4-nitrobenzaldehyde 2.
The MBH reaction is a formidable transformation in organic chemistry, with common small-molecule catalysts such as DABCO and DMAP requiring high catalyst loadings and long reaction times to achieve high yields. The de novo enzyme BH32 showed low-level activity for the MBH reaction, achieving less than one turnover a day. As we began this project, the Green group was in its infancy and we did not have the infrastructure needed to screen libraries efficiently using HPLC analytics. We therefore required a screening strategy that reported on aspects of the MBH reaction in a short amount of time using equipment we had available to us. To this end, we designed an acetylated derivative of the MBH product that served as a mechanistic inhibitor and gave rise to distinct spectroscopic features upon reaction with the catalytic nucleophile of BH32. This assay enabled us to rapidly evaluate libraries of eight rounds of evolution and install 13 mutations into the BH32 scaffold, providing the substantially improved biocatalyst BH32.8. Fortunately, improvement in activity for the spectrophotometric screen seemed to correlate well with MBH activity in early rounds. However, we were aware this assay did not report on all aspects of the MBH reaction, and with an improved enzyme in hand were keen to start evaluating libraries for true catalytic turnover. With new investment secured for a high-throughput analytical suite, we performed a further six rounds of evolution using a UHPLC screen, resulting in the final variant BH32.14 which had 24 mutations when compared to BH32 (Figure 2).
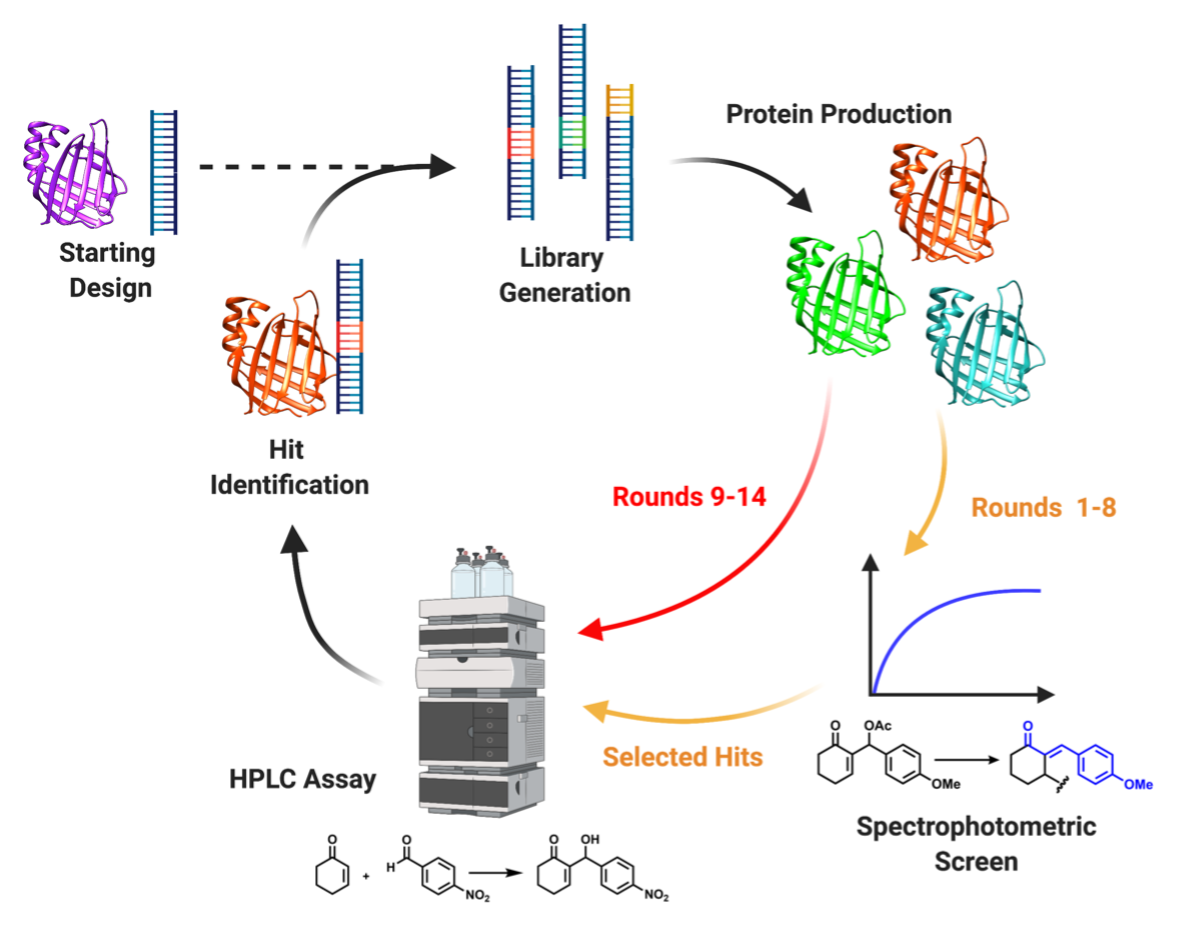
Figure 2: Directed Evolution Workflow. Schematic diagram showing the evolution workflow used to generate and evaluate libraries. Rounds 1-6 used a spectrophotometric screen to evaluate libraries, selected hits were then purified and screened for MBH activity using a HPLC assay. Rounds 7-10 evaluated libraries for MBH activity in lysate using a HPLC assay, selected hits were then purified and screened.
After a total of fourteen rounds of evolution, the evolved MBHase BH32.14 shows a 160-fold improvement in turnover number when compared to the starting design BH32. Although no explicit selection pressure was applied to selectivity, enzyme selectivity for the R-enantiomer increased from 20% to 93% e.e. for BH32 and BH32.14 respectively. Using low catalyst loadings of BH32.14 we can achieve >94% yield, and comparison to small molecule catalysts such as DABCO and DMAP highlights the efficiency of BH32.14. Obtaining crystal structures of evolved variants proved challenging, eventually we crystallised the advanced variant BH32.12 which revealed several interesting features. As well as the designed His23 nucleophile, the Arg124 mutation installed during evolution was shown to be of critical importance for MBH catalysis. DFT analysis showed that Arg124 stabilises multiple intermediates across the reaction coordinate (Figure 3). This mechanism shows similarities to methods developed by organic chemists, where nucleophilic catalysts are paired with hydrogen-bond donors such as thioureas.
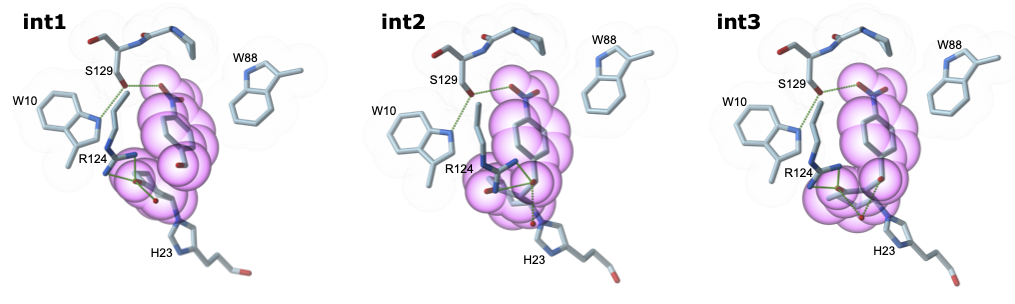 Figure 3: Proposed catalytic mechanism of BH32.14. DFT states for intermediates 1, 2 and 3 are shown in an all atom coloured stick representation.
Figure 3: Proposed catalytic mechanism of BH32.14. DFT states for intermediates 1, 2 and 3 are shown in an all atom coloured stick representation.
We elected to investigate the substrate scope of the engineered MBHase, and to our delight revealed that that BH32.14 accepts a broad range of aldehyde and enone coupling partners, including notoriously challenging substrates such as unsaturated lactones. Interestingly, BH32.14 also promotes the selective monofunctionalisation of dialdehydes. The broad substrate scope of BH32.14 means that it should provide an ideal starting point for the development of MBH biocatalysts with interesting activities. This work demonstrates that the combination of computational design and directed evolution can provide new biocatalysts for important chemical transformations not found in nature.
For more information, please read our article in Nature Chemistry via: https://www.nature.com/articles/s41557-021-00833-9
References
- Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008).
- Jiang, L. et al. De novo computational design of retro-aldol enzymes. Science 319, 1387–1391 (2008).
- Siegel, J. B. et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels-Alder reaction. Science 329, 309–313 (2010).
- Eiben, C. B. et al. Increased Diels–alderase activity through backbone remodelling guided by Foldit players. Nat. Biotechnol. 30, 190–192 (2012).
- Blomberg, R. et al. Precision is essential for efficient catalysis in an evolved Kemp eliminase. Nature 503, 418–421 (2013).
- Obexer, R. et al. Emergence of a catalytic tetrad during evolution of a highly active artificial aldolase. Nat. Chem. 9, 50–56 (2017).
- Bjelic, S. et al. Computational design of enone-binding proteins with catalytic activity for the Morita-Baylis-Hillman reaction. ACS Chem. Biol. 8, 749–757 (2013).
Follow the Topic
-
Nature Chemistry

A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in