Genomic insights into antibiotic resistance in E. coli ST131 bacteraemia cases in Wales
Published in Ecology & Evolution, Microbiology, and Protocols & Methods
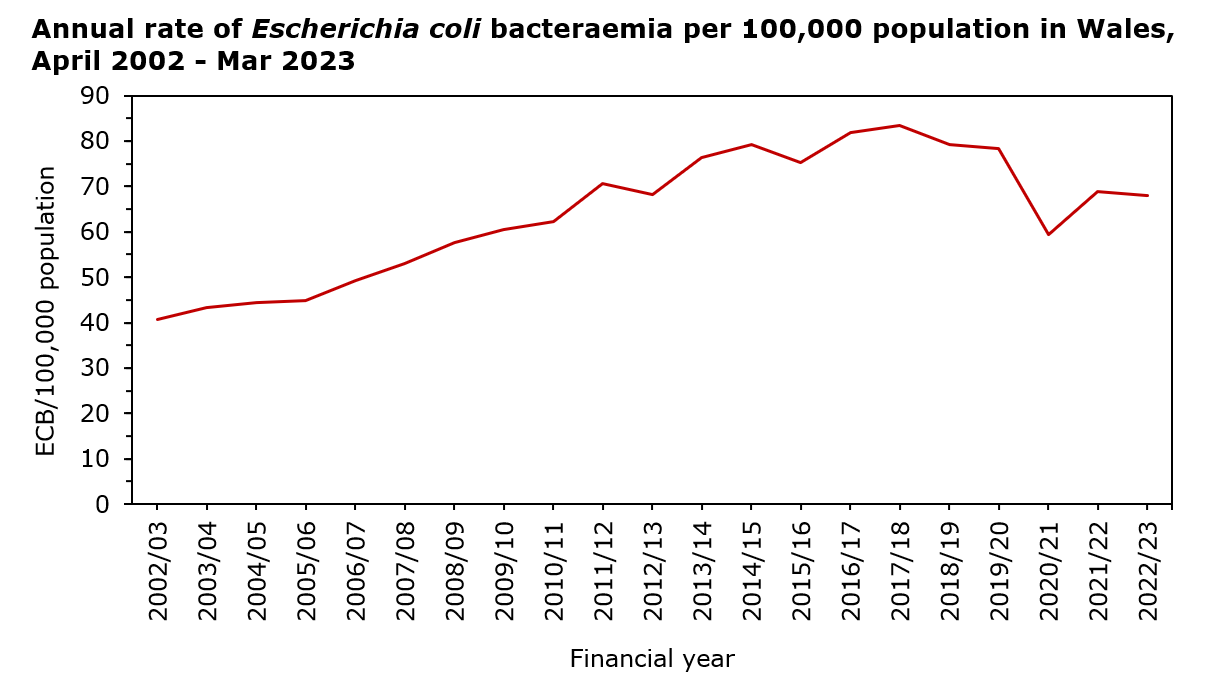
Globally, E. coli-associated bloodstream infections are on the rise. In Wales (known as Cymru in the Welsh language), the E. coli bacteraemia (ECB) rate per 1,000 hospital admissions was 5.2 in 2022/23.
Before this project was started, the annual aggregate data from our national bloodstream infection surveillance in Wales consistently showed E. coli as the primary organism responsible for bacteraemia but also revealed a concerning upward trend. In June 2011, Public Health England (replaced by the UK Health Security Agency) expanded their mandatory healthcare-associated infection surveillance program to cover ECB. The Chief Medical Officer for Wales responded to the rise in cases by seeking to understand the increase in ECB cases within the country. As a result, an in-depth investigation into ECB in Wales was requested.
The project aimed to identify modifiable risk factors contributing to the escalating ECB cases. The urgency to address this significant burden on patient care was evident to alleviate morbidity and mortality associated with ECB.
The in-depth investigation included multiple workstreams involving laboratories, infection prevention and control teams, epidemiologists and bioinformaticians from the National Health Service (NHS) and academic institutions across Wales. The main goal of one of the workstreams was to conduct retrospective and prospective whole-genome sequencing to determine whether new, more virulent or more resistant strains of E. coli were responsible for the increase in cases observed in Wales. These previous investigations using genomics and multilocus sequence typing (MLST) had shown that E. coli ST131 was disproportionately responsible for ECB cases, constituting 26% (n = 187/720). E. coli ST131 is a high-risk pandemic clone frequently associated with bacteraemia and urinary tract infections, and is a major circulating lineage in the UK (Lau et al. 2008; Harris et al. 2018). The escalating threat of antibiotic resistance emphasises the urgency of addressing it for global public health. Within this context, our study focuses on Uropathogenic E. coli (UPEC) ST131, a significant contributor to antibiotic-resistant infections, particularly bacteraemia.
By investigating the genomes of 142 E. coli ST131 strains collected from blood cultures in Wales from 2013 to 2014, we highlight the mechanisms driving third-generation cephalosporin resistance in this clinically important pathogen. This research not only sheds light on the specific challenges posed by this multidrug-resistant clone but also underscores the broader significance of genomic epidemiology in guiding targeted and timely public health interventions.
In summary, our project’s multifaceted approach, encompassing strategic human resource deployment and advanced ICT solutions, aims to inform and implement targeted interventions and contribute significantly to the broader national efforts in combating antimicrobial resistance and improving overall healthcare quality in Wales. Our study’s findings are crucial in shaping future policy and decision-making processes, supporting initiatives through increased funding and revitalising resources to combat disease burdens effectively.
We thank the Microbiology laboratories in Public Health Wales/Iechyd Cyhoeddus Cymru and across the Welsh National Health Service/Gwasanaeth Iechyd Gwladol Cymru, as well as the Specialist Antimicrobial Chemotherapy Unit at the University Hospital Wales for contributing samples to the national E. coli bacteraemia project led by the Healthcare Associated Infection, Antimicrobial Resistance & Prescribing Programme (HARP). HARP provided essential project management and epidemiological expertise, guiding the study methodology, including dataset preparation, sample selection, and transportation.
Funding for this work was provided by the Medical Research Council (grants MR/L015080/1, and MR/T030062/1), covering staff time and key computational resources for data storage and analysis. Additionally, the Wellcome Trust (project funding via the Cardiff University ISSF fund) supported sequencing and initial analysis of the samples.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in