Greater than the sum of their parts: finding relationships between histone modifications in single cells
Published in Bioengineering & Biotechnology

Towards understanding chromatin state regulation at single-cell resolution.
Detecting histone modifications in single cells by sequencing is still in its infancy, but has the potential to unlock the full spectrum of different chromatin states in the genome of individual cells. More established single-cell sequencing technologies, such as mRNA-seq and ATAC-seq, interrogate only a tiny fraction of the genome. Expanding beyond mRNA abundances and open chromatin is challenging because profiling chromatin states involves the interactions between a large number of different histone modifications and proteins that decorate the entire genome. Progress therefore hinges on both new measurement methods that can map multiple epigenetic marks in single cells as well as new analysis methods that connect different modalities together.
From solving technical pain points to addressing broader gaps.
The personal motivation for mapping and analyzing multiple histone modifications in single cells came from pain points in analyzing different histone modifications separately [1]. Many basic analyses that were straightforward in conventional datasets (e.g. single-cell RNA- and ATAC-seq) were difficult in single-cell histone modification data. For example, the entire procedure of cell typing assumes that there is a literature of known marker genes to annotate clusters in the data to known cell types. However, these marker genes that were derived from mRNA abundance data often have no signal in certain histone modifications (e.g. in H3K9me3).
After some reflection, we realized that these seemingly technical issues were actually signs of a broader gap. We lacked quantitative understanding of the relationships between different chromatin states, and especially how those relationships change across cell types and during differentiation. We therefore focused on elucidating these relationships in single cells.
scChIX-seq: an integrated experimental and computational framework
Most methods that detect histone modifications in single cells use a “single-incubation” strategy. We incubate cells with an antibody specific to a histone modification. The antibody acts as a beacon, telling us where to cut the DNA (Figure 1). The upside of this single-incubation strategy is that assigning cuts to a histone modification is trivial: every cut belongs to the same modification. However, single-incubation profiles only one histone modification per cell.
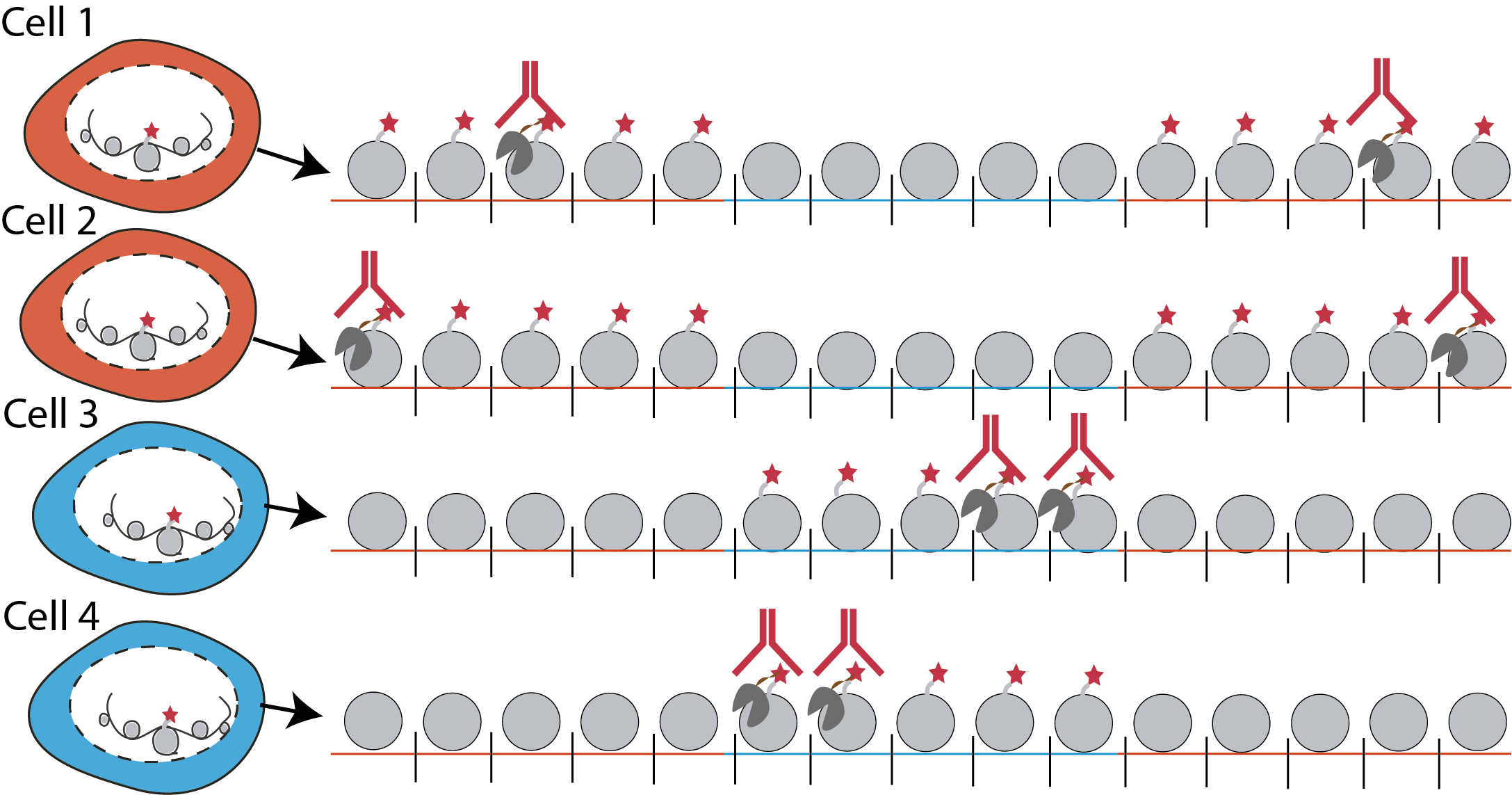
What if we placed two beacons instead to measure two modifications per cell? In this naive “double-incubation” strategy, the data generated would be more complicated. The beacon would now tell us genomic regions that have either one of the two modifications, but not which one exactly. The challenge would now be to assign each cut fragment back to its modification-of-origin. We found that by training a statistical model from single-incubated data, we could solve this deconvolution problem (Figure 2). We called this integrated experimental and computational method, scChIX-seq (single-cell chromatin immunocleavage and unmixing followed by sequencing) [2]. Of note, there have recently been a number of experimental efforts to address this cut assignment problem [3, 4, 5, 6]. We can now generate multimodal histone modification data in single cells.
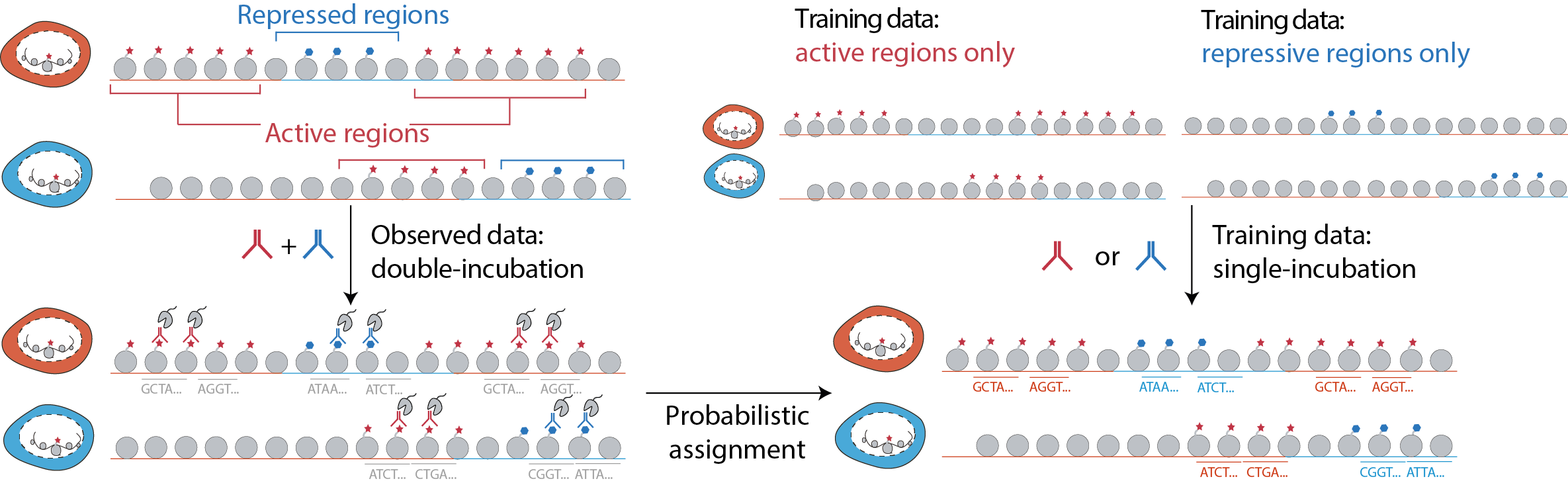
Greater than the sum of their parts: analysis methods that connect between modalities
Simply having multimodal data does not automatically reveal the relationships between the modalities. We therefore developed new analysis methods tailored for this new data type, focusing on revealing insights that would otherwise be missed from conventional unimodal data. For example, we found many-to-one relationships where different cell types could share a similar heterochromatin landscape. We also found temporal relationships where changes in one histone modification preceded the changes in another. Conceptually, what we can get from multimodal data is the joint distribution between histone modifications, whereas unimodal data would only have marginal distributions (Figure 3). Overall, these new multimodal methods revealed dynamic relationships between histone modifications.
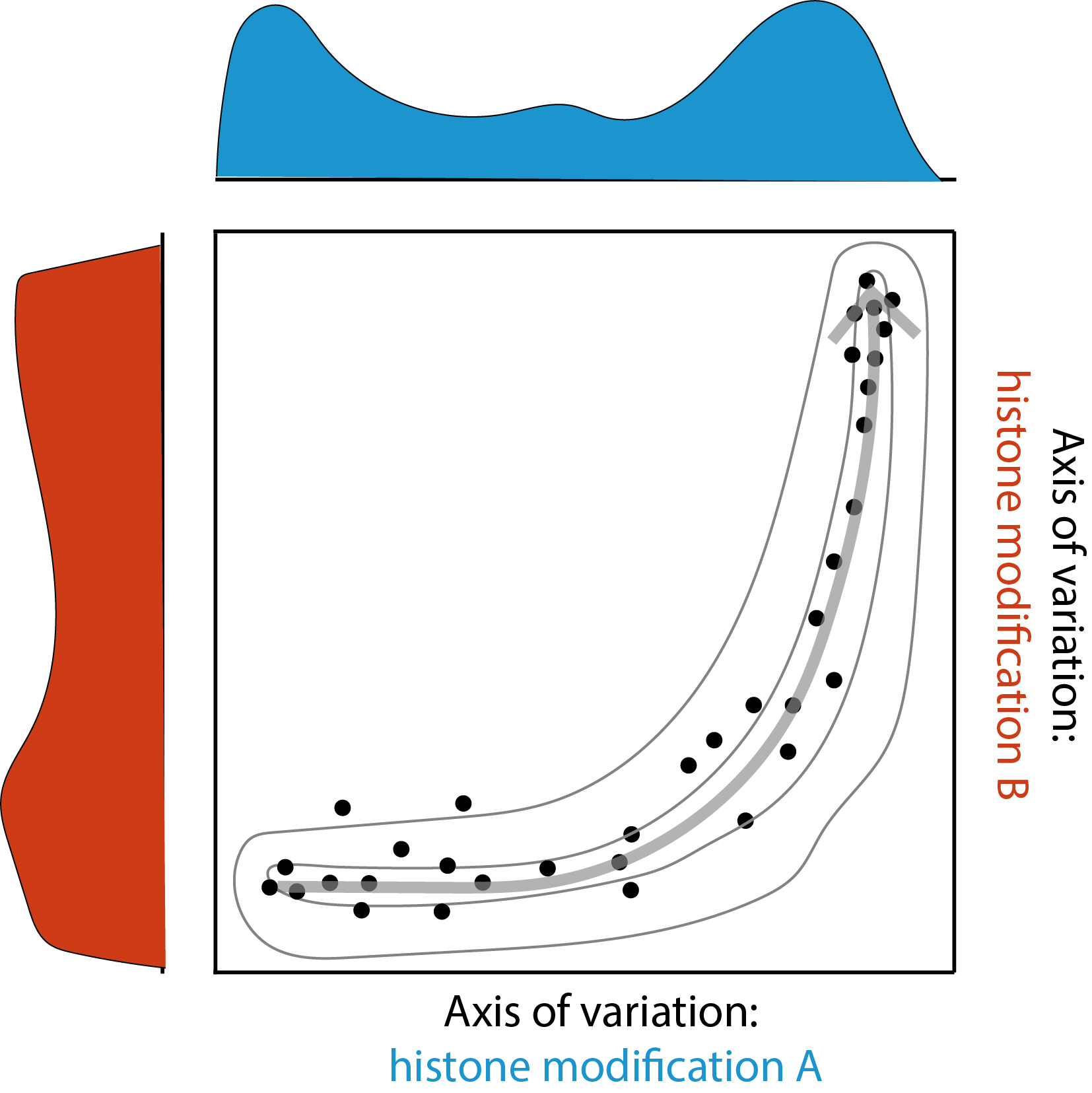
Future perspectives
Multimodal histone modification and transcription factor binding data is increasingly more prevalent. Exciting opportunities lay ahead in mapping and modeling how cells read the different histone modifications, transcription factors, and other regulators to modify or maintain their cell type-specific functions. Since there are so many different types of histone modifications and transcription factors, we will likely encounter basic experimental design questions. Which epigenomic combinations give the most “bang for sequencing buck”? Finally, perturbation experiments with multimodal epigenomic readouts may help disentangle the cause and consequence of different factors that decorate the chromatin.
References
[1] Zeller, P., Yeung, J., Viñas Gaza, H. et al. Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis. Nat Genet (2022). https://doi.org/10.1038/s41588-022-01260-3
[2] Yeung, J., Florescu, M., Zeller, P. et al. scChIX-seq infers dynamic relationships between histone modifications in single cells. Nat Biotechnol (2023). https://doi.org/10.1038/s41587-022-01560-3
[3] Gopalan, Sneha, et al. "Simultaneous profiling of multiple chromatin proteins in the same cells." Molecular Cell 81.22 (2021): 4736-4746. https://doi.org/10.1016/j.molcel.2021.09.019
[4] Meers, M.P., Llagas, G., Janssens, D.H. et al. Multifactorial profiling of epigenetic landscapes at single-cell resolution using MulTI-Tag. Nat Biotechnol (2022). https://doi.org/10.1038/s41587-022-01522-9
[5] Bartosovic, M., Castelo-Branco, G. Multimodal chromatin profiling using nanobody-based single-cell CUT&Tag. Nat Biotechnol (2022). https://doi.org/10.1038/s41587-022-01535-4
[6] Stuart, T., Hao, S., Zhang, B. et al. Nanobody-tethered transposition enables multifactorial chromatin profiling at single-cell resolution. Nat Biotechnol (2022). https://doi.org/10.1038/s41587-022-01588-5
Acknowledgements
We thank Scott Uminga for creating the cover art (https://www.scottuminga.com/).
Follow the Topic
-
Nature Biotechnology
A monthly journal covering the science and business of biotechnology, with new concepts in technology/methodology of relevance to the biological, biomedical, agricultural and environmental sciences.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in