How Rif1 bridles rapid embryonic DNA synthesis
Published in Cell & Molecular Biology
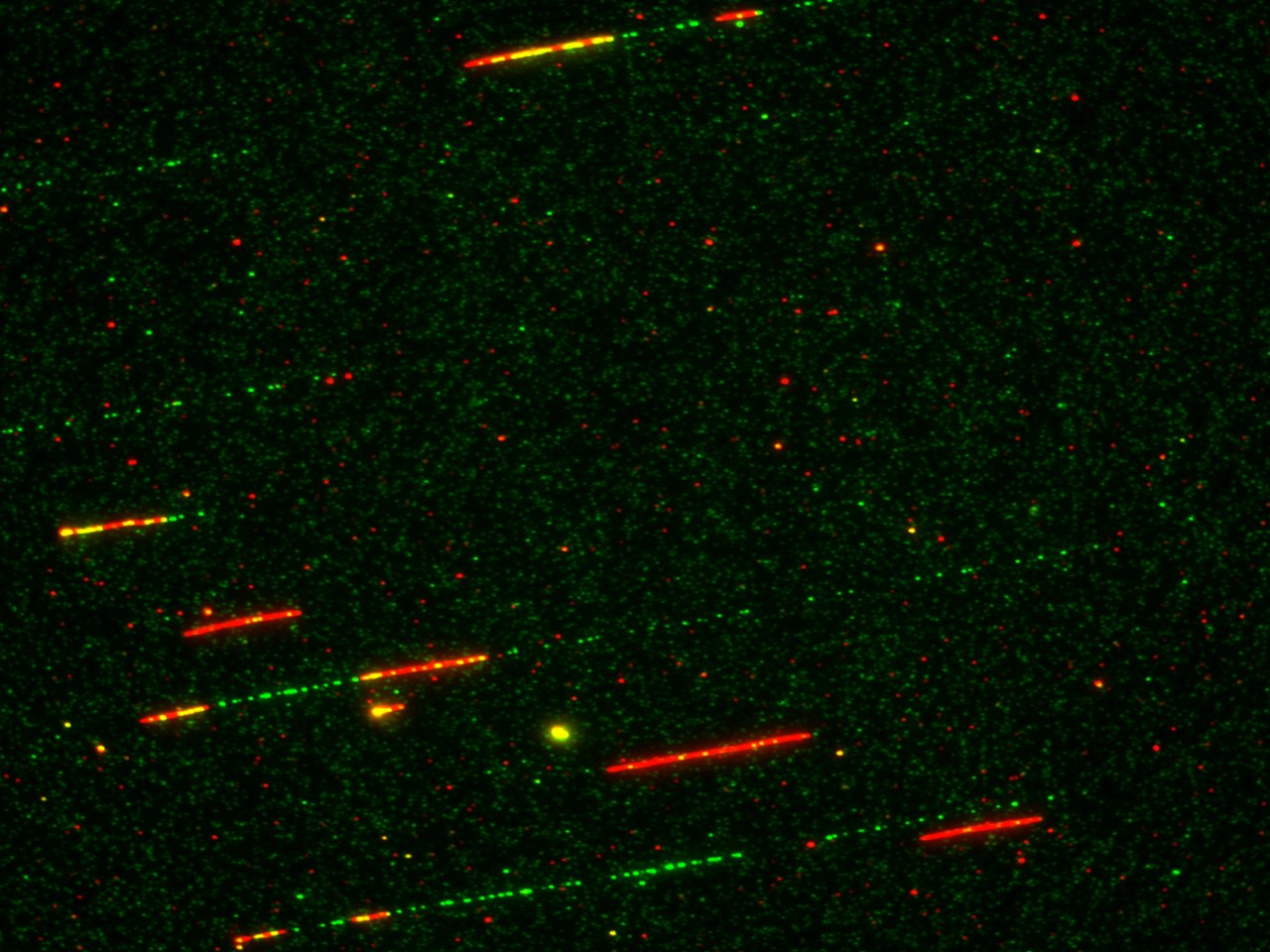
The faithful duplication of the genetic material is crucial for the cell. With several thousand active replication origins to ensure this task in vertebrates, the spatial and temporal coordination of the replication program is strictly regulated to avoid that no DNA is left unreplicated upon entry into mitosis. The temporal program of DNA replication is a conserved signature for genomes of multicellular organisms. It has been known that transcribed and euchromatin regions generally replicate early, and heterochromatin replicates late in differentiated cells. This positive correlation between transcription and early replication led to the general assumption that a regulated, temporal replication program does not exist in cell cycles where transcription and the distinction between eu and heterochromatin is absent, like in early, rapid cell cycles during the development of amphibians, fishes or insects. Before the onset of zygotic transcription, which takes place after 12 cell divisions in the model organism Xenopus laevis, cells rely only on maternally supplied factors to quickly increase cell number until the mid-blastula transition when cell cycles slow down, and the S phases lengthen. Our findings here further extend previous studies in early Xenopus and Zebrafish cell cycles and strongly suggest that regulated temporal regulation of DNA replication at the level of large chromatin domains exists before the onset of transcription. How did we come to this conclusion?
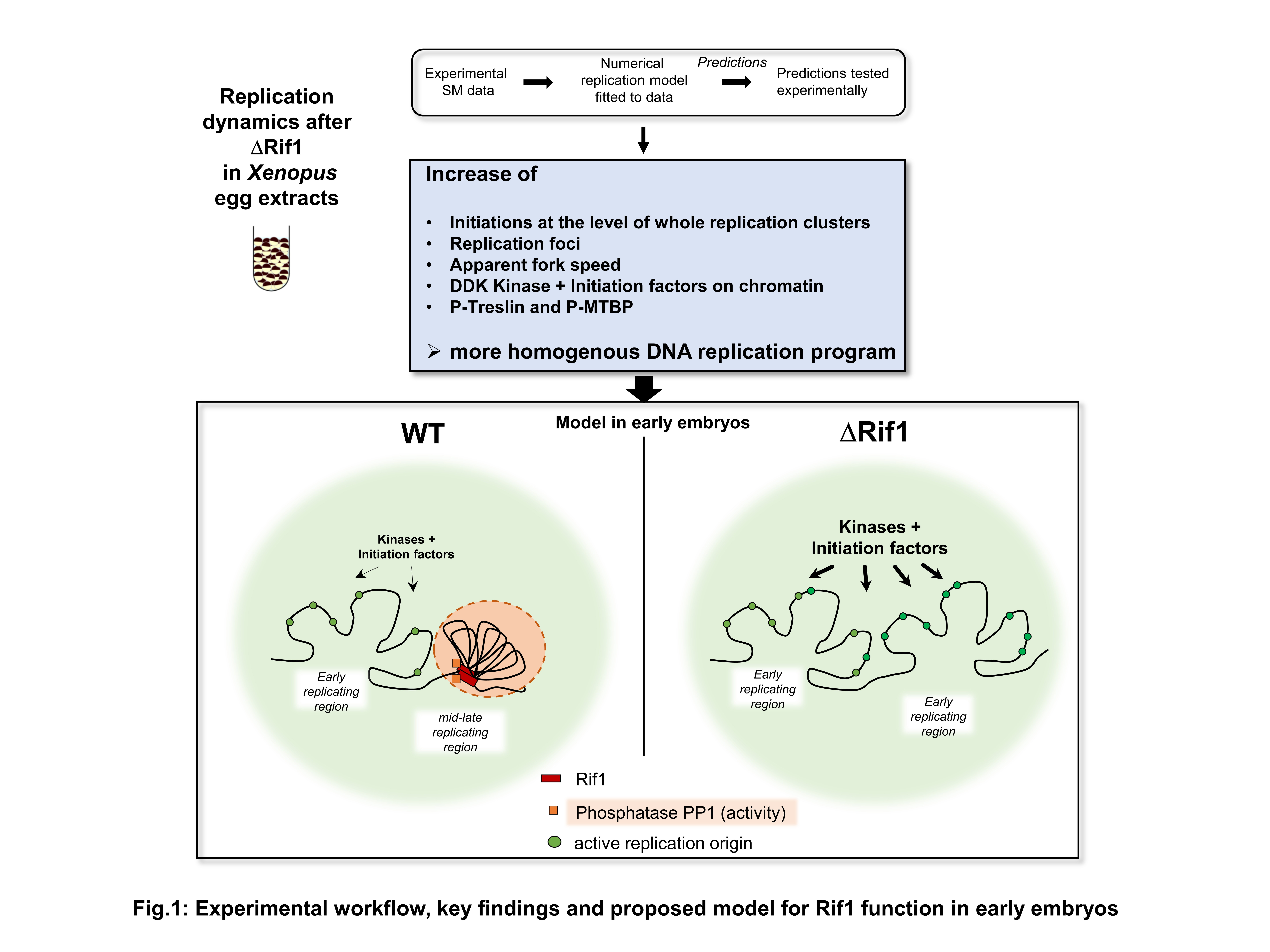
We studied the role of a well-known factor in multicellular organisms to control the replication timing program, Rif1, in the Xenopus in vitro system. Rif1 depletion in differentiated cells leads to many late genomic regions or domains replicating earlier and some early regions replicating later. Mechanistically, Rif1 inhibits origin firing by recruiting a phosphatase (protein phosphatase 1, PP1) to chromatin, counteracting the S phase kinase DDK and the activation of the replicative helicase in all organisms. It was known that during early Xenopus laevis developmental stages, Rif1 depletion accelerated DNA synthesis in the vitro system1,2 and early cell cycles in vivo2. Having these few pieces of the puzzle, we still needed the whole picture. We hypothesized that Rif1 could regulate the replication program similarly to differentiated cells, meaning globally at the level of entire chromatin domains. The still poor Xenopus laevis genome assembly makes whole NGS approaches, used in differentiated cells to map genome-wide replication domains, challenging. Therefore, we went down a different way and combined experimental DNA fiber stretching and biochemical approaches in time course experiments with numerical simulations (Fig.1).
Using a DNA combing approach, we found that Rif1 depletion in the in vitro system led to a surprisingly substantial increase in origin activation without strongly decreasing distances between neighboring origins on DNA fibers. Other observations further suggested that Rif1 depletion simply leads to accelerating the replication program without changing local replication parameters or, in other words, to a more homogenous, temporally compressed program. This was not a big surprise to us since we hypothesized that Rif1 regulates at a level above the observed DNA fiber length (100-200 kb). Consistent with this, we also observed a strong increase in the replication foci number at the beginning of the S phase after Rif1 depletion. This suggests that Rif1 slows down DNA replication at the level of replication clusters (50-150 kb) or replication domains (200kb-1Mb).
We next fitted our established numerical model3 to our experimental DNA combing data to gain further insights. Our model assumed that the Xenopus genome could be divided into regions where origins are activated with a high probability or, in other words, early replicating regions and into regions where origins fire with a low probability or late replicating regions. This is similar to observations and models in differentiated cells. The numerical model fitted well with the control and Rif1 depleted data, and we analyzed the changes in our seven model replication parameters. The results predicted that Rif1 depletion would intrinsically increase only one parameter, which defines the accessibility of limiting replication factors. Can experiments validate this prediction, and what could be these factors?
By quantitative western blotting of chromatin fractions, we found that Rif1 depletion led to an increase of the S phase kinase Drf1/Cdc7 and several key initiation factors (Treslin/MTBP, RecQL4) involved in the activation of the replicative helicase complex. In addition, we found that the DDK-dependent phosphorylation of Treslin and MTBP increased, which could be directly linked to the absence of the phosphatase, which is not targeted anymore by Rif1 at origins. Therefore, the model predictions proved to be correct. How exactly Rif1 could block these factors from chromatin binding and thus from activating origins will be very interesting to see in future studies.
Conclusions
Altogether our study shows that the early embryonic S phase is not running at its fastest biochemical rate and can be accelerated by Rif1 depletion in vitro and in vivo, contrary to the prevailing opinion in the field. But most important, our results point to the role of Rif1 in regulating a replication timing program at the level of larger chromatin domains in early embryos. We propose a model for Rif1 function in early embryos (Fig. 1) where Rif1 binding to domains restricts the accessibility of initiation factors to chromatin by keeping close domain contacts and by establishing locally high phosphatase activity, which slows down S phase progression, similar as it has been suggested for somatic cells4. This could be by directly inhibiting initiation factors like Treslin/MTBP and/or indirectly by modulating DDK access to chromatin and the ability of MCM2-7 to compete for limiting factors. In the absence of Rif1, structurally more relaxed domains and the lack of PP1 would allow the binding of DDK kinase and firing factors, such as Treslin/MTBP and RecQL4, to origins. This would boost initiations in whole clusters, as we observed by DNA combing analysis. By regulating chromatin access to replication factors, when specific subnuclear compartments and eu/heterochromatin differentiation have not yet been established in early embryos, Rif1's role could be to maintain a global epigenetic state, preparing the ground for changes occurring later during differentiation and development.
What made the difference in this study in the end? Using a numerical model to back up the experiments was valuable here. We directly looked at Treslin/MTBP or DDK because the literature described these factors as rate-limiting for DNA replication for several years5, but no direct link to Rif1 had been established. A general take-home message is to use numerical models in biological workflows more often and as early as possible to gain insights into complex biological processes.
- Alver, R. C., Chadha, G. S., Gillespie, P. J. & Blow, J. J. Reversal of DDK-Mediated MCM Phosphorylation by Rif1-PP1 Regulates Replication Initiation and Replisome Stability Independently of ATR/Chk1. Cell Rep. 18, 2508–2520 (2017).
- Meléndez García, R. et al. A non-transcriptional function of Yap regulates the DNA replication program in Xenopus laevis. eLife 11, e75741 (2022).
- Ciardo, D. et al. Organization of DNA Replication Origin Firing in Xenopus Egg Extracts: The Role of Intra-S Checkpoint. Genes 12, 1224 (2021).
- Foti, R. et al. Nuclear Architecture Organized by Rif1 Underpins the Replication-Timing Program. Mol. Cell 61, 260–273 (2016).
- Collart, C., Allen, G. E., Bradshaw, C. R., Smith, J. C. & Zegerman, P. Titration of four replication factors is essential for the Xenopus laevis midblastula transition. Science 341, 893–896 (2013).
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in