Optimizing Lab Engines
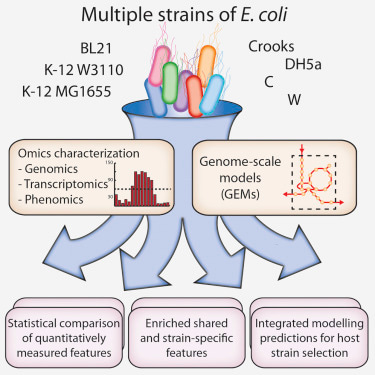
Quantification of strain-specific differences in seven widely used strains of E. coli (BL21, C, Crooks, DH5a, K-12 MG1655, K-12 W3110, and W) using genomics, phenomics, transcriptomics, and genome-scale modeling.
Published in Microbiology
Like
Be the first to like this
In the Paper a group studied various strains of E. coli using genomics, phenomics, transcriptomics, and genome-scale modeling.The highlight of this study is they checked the performance of E. coli strains under different conditions (Transcriptomic data GSE78756) which enhances understanding of optimization as well selection of strain suitable for use either for basic biology or for industrial use. The transcription factors, codon usage and global regulators rpoS also plays vital role in gene expression. Finally paper introduces a new star method format brought by Cell press.
Paper Link : http://www.sciencedirect.com/science/article/pii/S2405471216302903
Follow the Topic
Microbiology
Life Sciences > Biological Sciences > Microbiology



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in