Peeling back the genetics of groundnut blanchability
Published in Agricultural & Food Science

Understanding the trait: a small step with big consequences
Groundnut (Arachis hypogaea) is a familiar crop on our plates, yet its journey from field to food product involves several intricate steps. Whether it is groundnut butter, roasted snacks, or confectionery, one critical process stands between harvest and consumption-blanching, the removal of the thin seed coat surrounding the kernel.
At first glance, blanching may seem like a routine mechanical step. But anyone working in the groundnut industry quickly realizes that not all groundnuts behave the same way. Some release their testa effortlessly after roasting, while others stubbornly retain fragments. This variation, known as blanchability, has significant implications for processing efficiency, product quality, and economic returns.
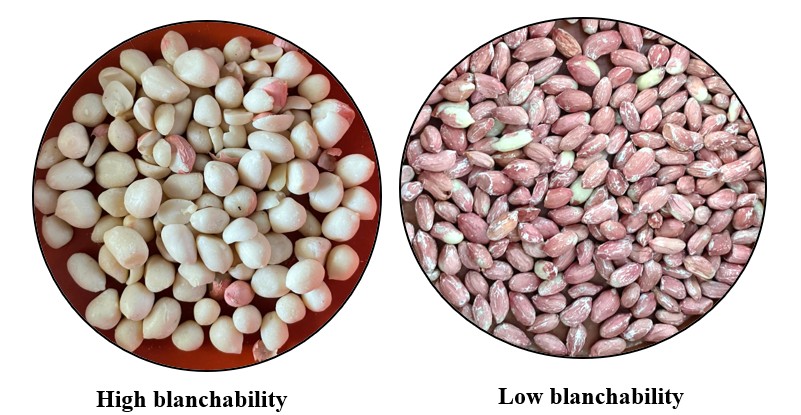
Image 1: Variation in blanchability among groundnut genotypes
Interestingly, blanchability is not always about “better is higher.” While high blanchability is preferred for clean kernel production, low blanchability also has value in specific products such as coated nuts and confectionery, where testa retention is desirable. This dual importance makes blanchability a uniquely complex trait from a breeding perspective.
Despite its importance, the genetic basis of blanchability has remained largely unexplored. This gap became the starting point of our journey.
Blanchability is not governed by a single factor. It reflects a combination of biological processes, including seed coat thickness, adhesion between seed coat and cotyledon, and underlying biochemical composition. These interconnected mechanisms make it a quantitatively inherited and environmentally sensitive trait.
One of our earliest challenges was phenotyping. Groundnut grown during the rainy season typically produces lower pod yield and smaller, less uniform kernels due to high humidity, and foliar diseases, thus posing a challenge for blanchability phenotyping using the standard protocol, where 200-g seeds are required. In contrast, post-rainy (rabi) crops have higher pod filling, better seed uniformity, and greater kernel availability. In essence, our findings revealed that when dealing with large-scale blanchability screening in low yielding rainy season, this modified approach based on 50g provides scope trait mapping.
Genebank exploration: where diversity tells the story
To truly understand a complex trait, one must first explore diversity. This led us to the rich germplasm resources conserved in the ICRISAT genebank.
Cultivated groundnut consists of two major subspecies:
- Arachis hypogaea subsp. hypogaea
- Arachis hypogaea subsp. fastigiata
These subspecies differ in plant architecture, growth habit, and maturity duration. To capture this diversity, we worked with a wide collection of genotypes sourced from global germplasm. These collections capture groundnut evolution, domestication, and breeding, making them invaluable for uncovering hidden genetic variation.
Working with such diversity was both exciting and challenging-it required balancing large datasets with consistent phenotypic evaluation. This highlights an important message: the solutions to modern agricultural challenges often lie hidden in existing diversity.
From diversity to discovery: GWAS and genomic insights
With robust phenotypic data and diverse germplasm in hand, the next step was to connect phenotype with genotype.
We employed whole-genome resequencing (WGRS) followed by genome-wide association studies (GWAS) to identify genomic regions associated with blanchability. This integrative approach allowed us to systematically scan the genome and detect significant marker-trait associations, single nucleotide polymorphisms (SNPs)-minor DNA changes commonly used in genetic studies.
However, as the analysis progressed, it became evident that individual SNPs could only provide a partial picture. For a complex trait like blanchability, controlled by multiple interacting loci, focusing solely on single markers risked oversimplifying the underlying biology.
This realization prompted a shift in our approach: haplotypes.
A haplotype represents a group of genetic variants inherited together as a block. Rather than examining isolated mutations, haplotypes allow us to capture the combined effect of multiple genetic changes, providing a more realistic picture of how traits are controlled in nature.
As, we began to visualize haplotype patterns across the genome, something unexpected emerged. The haplotypes associated with blanchability were not uniform across all genotypes. Instead, they showed distinct patterns specific to each subspecies.
This finding was both surprising and insightful. It suggested that the genetic architecture of blanchability is not universal-what works in one subspecies may not work in another. In other words, the “best” genetic combination for blanchability depends on the genetic background.
Moments like these-when patterns begin to emerge from complex data-are among the most rewarding in research. They transform scattered information into meaningful biological insight.
The identification of haplotypes is not merely descriptive-it has direct translational value.
These haplotypes can be converted into molecular markers, enabling breeders to:
- Identify superior donor genotypes
- Track desirable genomic regions during selection
- Develop varieties tailored for specific processing needs
This forms the foundation of haplotype-based breeding, a strategy particularly suited for complex traits like blanchability, where multiple genes act in concert and also opens new avenues:
- Functional validation of key haplotypes
- Integration with other quality traits such as oil content and dormancy
- Deployment in breeding populations for product-specific improvement
As genomic tools continue to advance, the possibility of designing crop varieties with customized traits is becoming increasingly tangible.
A journey of discovery beyond data
Behind this work lies a journey of data, discussions, and collaboration.
From handling large genomic datasets to standardizing phenotyping protocols, each step required persistence and teamwork. One of the most exciting moments came when clear subspecies-specific haplotype patterns began to emerge-revealing that the groundnut genome holds answers to questions that breeders have faced for years.
Even in a well-studied crop like groundnut, there are still many hidden layers waiting to be uncovered. In essence, our journey revealed key insights for informed breeding decisions to customize groundnut for industrial applications.

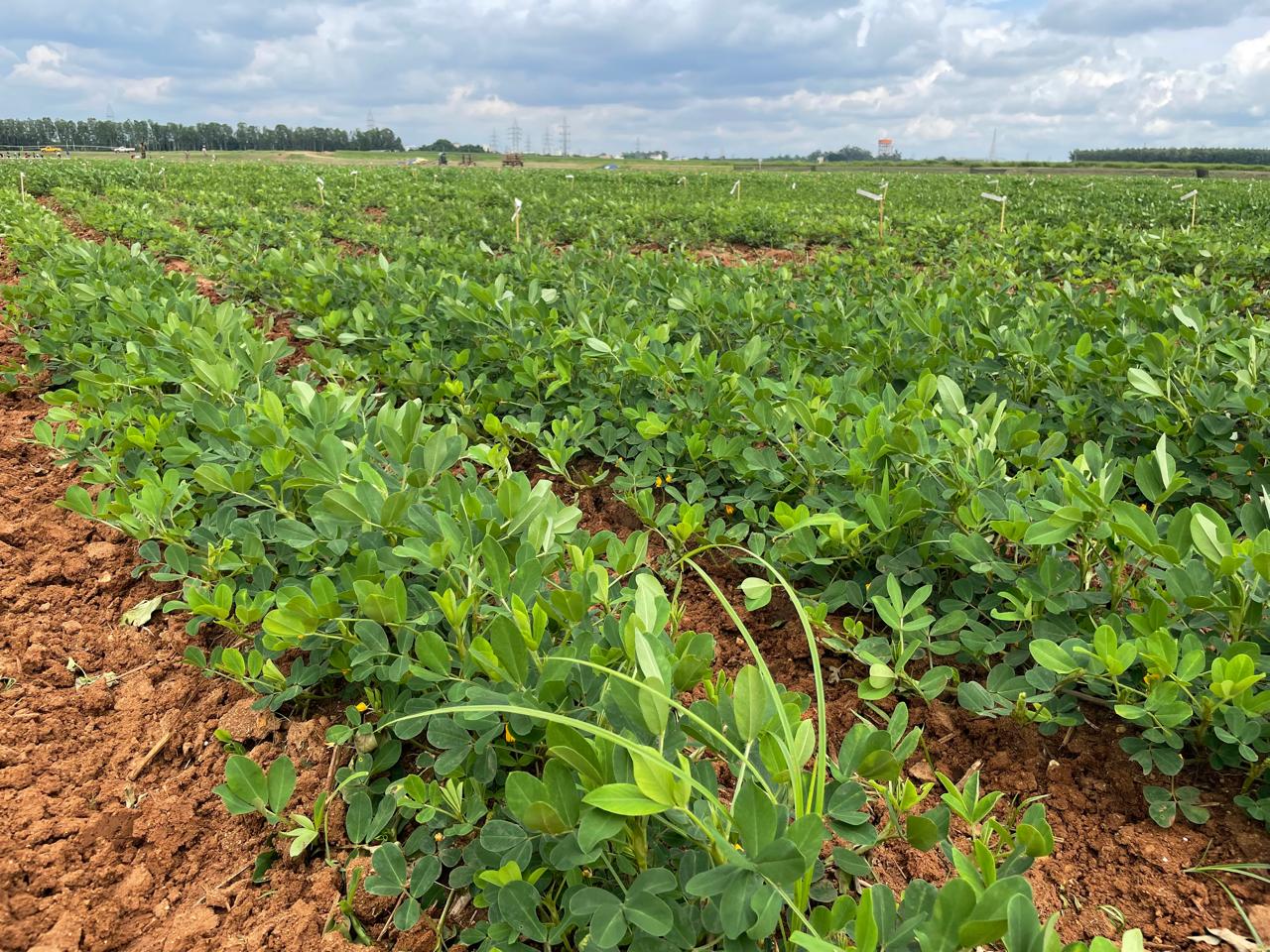
Image 2: A moment in the field-where collaboration, science, and innovation converge to explore blanchability genomics, informing industry-ready groundnut breeding.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in