The COVID-19 XPRIZE and the need for scalable, fast, and widespread testing
Published in Bioengineering & Biotechnology
Posted on behalf of:
Matthew J. MacKay, Anna Hooker, Ebrahim Afshinnekoo, Marc Salit, Jason Kelly, Jonathan V. Feldstein, Nick Haft, Doug Schenkel, Subbu Nambi, Patrick Cai, Feng Zhang, George Church, Junbiao Dai, Cliff L. Wang, Jeff Huber, Hanlee Ji, Alison Kriegel, Anne L. Wyllie & Christopher E. Mason (affiliations provided below)
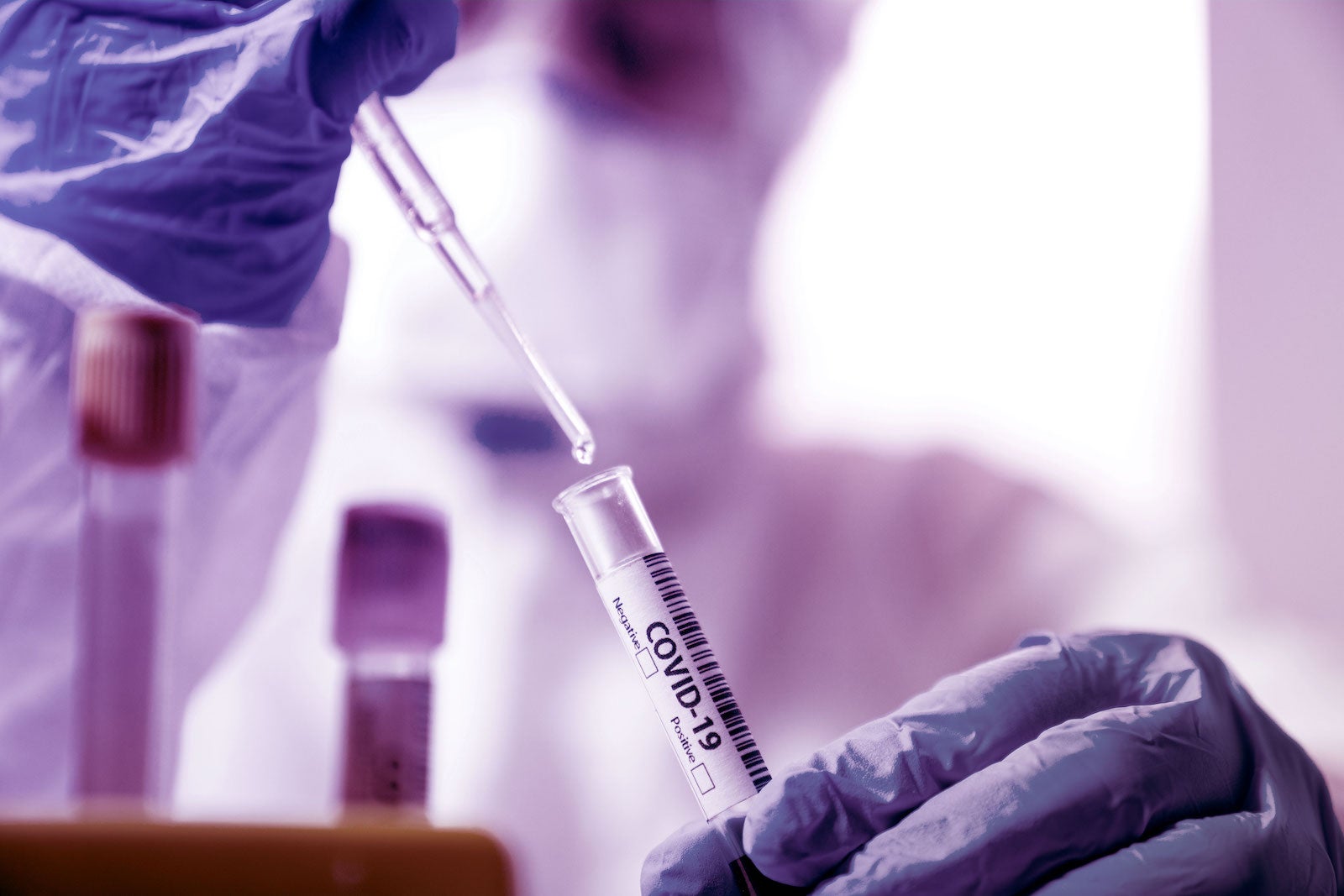
To the Editor — The US Food and Drug Administration (FDA) Emergency Use Authorization (EUA) and Instructions for Use (IFU) documents outlining the current approved virology tests for SARS-CoV-2 are largely unstandardized. As such, there remains an urgent need for a searchable interface allowing exploration of standardized information reported in these EUA and IFU documents. To gain an improved understanding of the current testing landscape and to galvanize future test development, we present here an online tool (http://www.resiliencehealth.com/tests) that profiles current and emerging virology tests for detecting SARS-CoV-2 (Fig. 1). We also call on the research community to respond to an open COVID-19 XPRIZE competition, OpenCovidScreen, seeking to identify cheap, high-quality, scalable testing solutions.
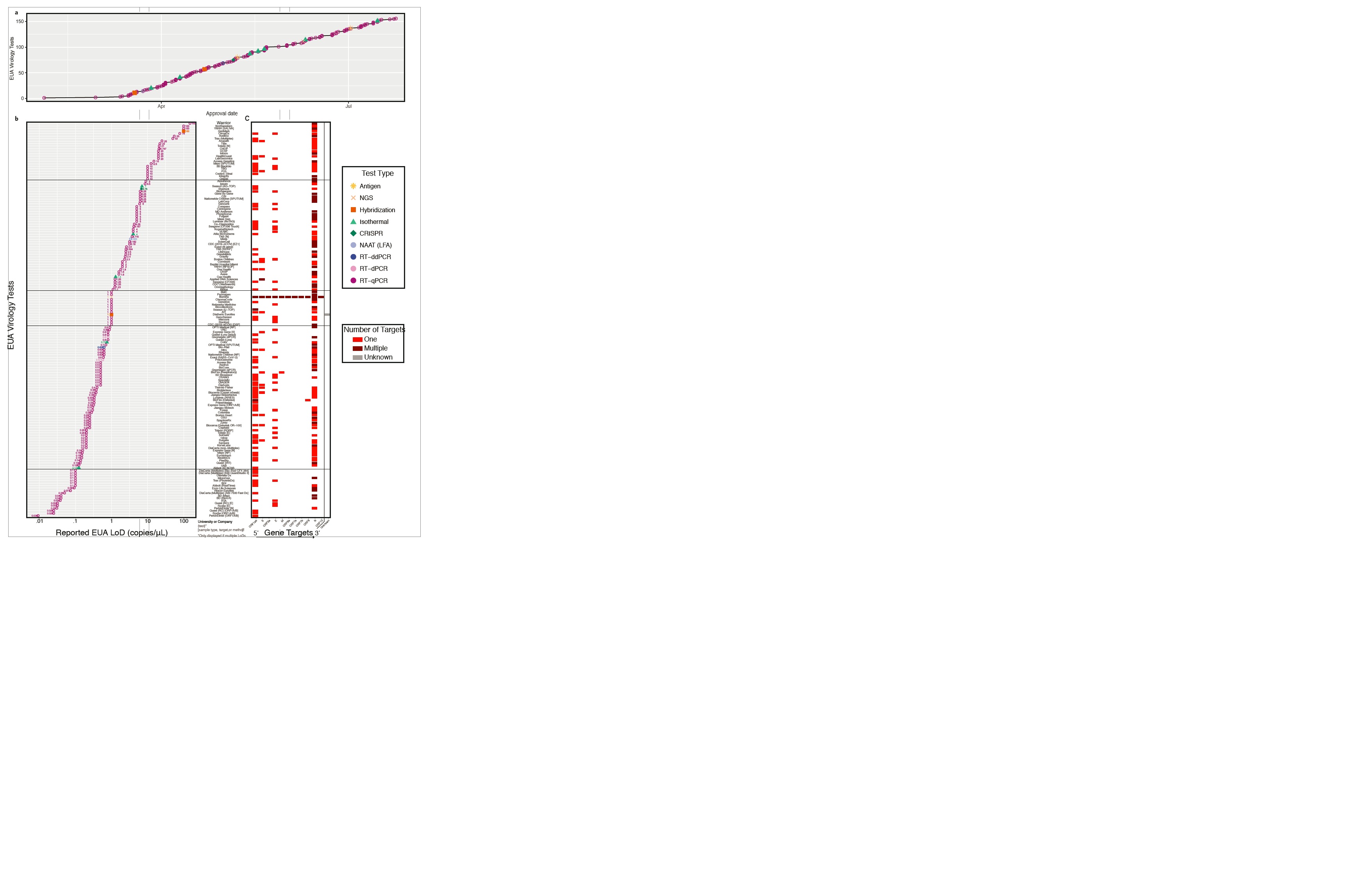

Fig. 1 | EUA Virology test types. a, Cumulative number of EUA virology tests approved from February to July 2020. b, Limit of detection (LoD) for tests that reported copies/mL or copies/mL (presented in copies/mL throughout). c, EUA virology test targets. SARS-CoV-2 5¢à3¢ genome4 on horizontal axis, with tests on the vertical axis and colors indicating whether the test has one target (red) or multiple targets (brown) in the specified region. Each line indicates a test source (company or university). If an EUA reported different LoDs for different targets, sample types or methods, each is displayed as a separate row. NGS, next-generation sequencing; NAAT, nucleic acid amplification test; LFA, lateral flow assay; RT, reverse transcription; ddPCR, Droplet Digital PCR; dPCR, digital PCR; qPCR, quantitative PCR.
As of 27 July 2020, an analysis of the FDA data on EUA SARS-CoV-2 virology tests reveal a wide range of limit of detection (LoD), spanning >5 orders of log10 differences. These metrics are of critical importance because each 10-fold increase in the LoD of a COVID-19 viral diagnostic test is expected to increase the false negative rate by 13%1.
Beyond this variable performance reported in IFUs for EUA tests, key attributes of many tests, such as primer sequences, protocol steps or viral gene targets, are either unclear or missing. Also, most approved EUAs use large multipliers (2- to 200-fold) on their own LoD for contrived or clinical samples to pass the minimum threshold for approval, based on a 95% positive/negative percentage agreement across at least 30 positive and 30 negative samples. As submissions stand, it is difficult to directly compare results and even understand how a test will actually translate into real-world or clinical settings.
Moreover, there has not been an independent assessment of these tests’ abilities or a comprehensive benchmarking of their strengths and weaknesses in different clinical settings, nor a consistent sample type (for example, nasopharyngeal, nasal, saliva) used across the EUAs. Thus, a comprehensive benchmarking effort on all methods on the market would be helpful, similar to ones conducting head-to-head studies of serological tests2 and other sites that annotate and analyze some of these tests, such as FindDx (https://www.finddx.org/), the US National Institute for Standards and Technology (NIST)’s Rapid Microbial Testing Methods Consortium (https://www.nist.gov/programs-projects/nist-rapid-microbial-testing-methods-consortium) and the COVID-19 Testing Project (https://covidtestingproject.org/index.html).
However, even if all current EUA tests for the SARS-CoV-2 virus performed with >95% sensitivity and >95% specificity, their combined capacities would still fall short of enabling large-scale, ubiquitous temporal monitoring (tens of millions per day), involving samples with varying viral load and substrates3. Also, tests with a higher LoD would not be readily applicable to pooling strategies, in which samples are by definition diluted before testing. Even the lower LoD tests, while promising for pooling, have not had independent LoD assessments. Moreover, truly city-scale or even national-level testing to decrease and control infection rates would require fast turnaround times (test result in less than one day), easy processing, and a low cost (per test and capital expense), such that a consumer, employer or government body could easily pay for multiple tests a week for each person.
To address the above issues, we have designed the COVID-19 XPRIZE competition, OpenCovidScreen (https://opencovidscreen.org/), to identify economically viable, high-quality, scalable testing options (Fig. 2). After competitors are selected on the basis of their results, methods, cost, scalability and speed, they will then be sent blinded samples to analyze. Results will be uploaded to the XPRIZE site and analyzed for overall performance, LoD and false positives. The finalists, based on their overall methods, results and innovation, will go into the clinical validation round, in which OpenCovidScreen will follow the methods laid out by the competitors to assess the reproducibility of their results. Top teams from this validation round will be awarded a prize and selected to set up and deploy testing sites, wherein OpenCovidScreen will help scale their tests and expand them into more locations, as well as to coordinate with government, industry and non-profit efforts (for example, the NIH RADx Program and Testing for America; https://www.nih.gov/research-training/medical-research-initiatives/radx).
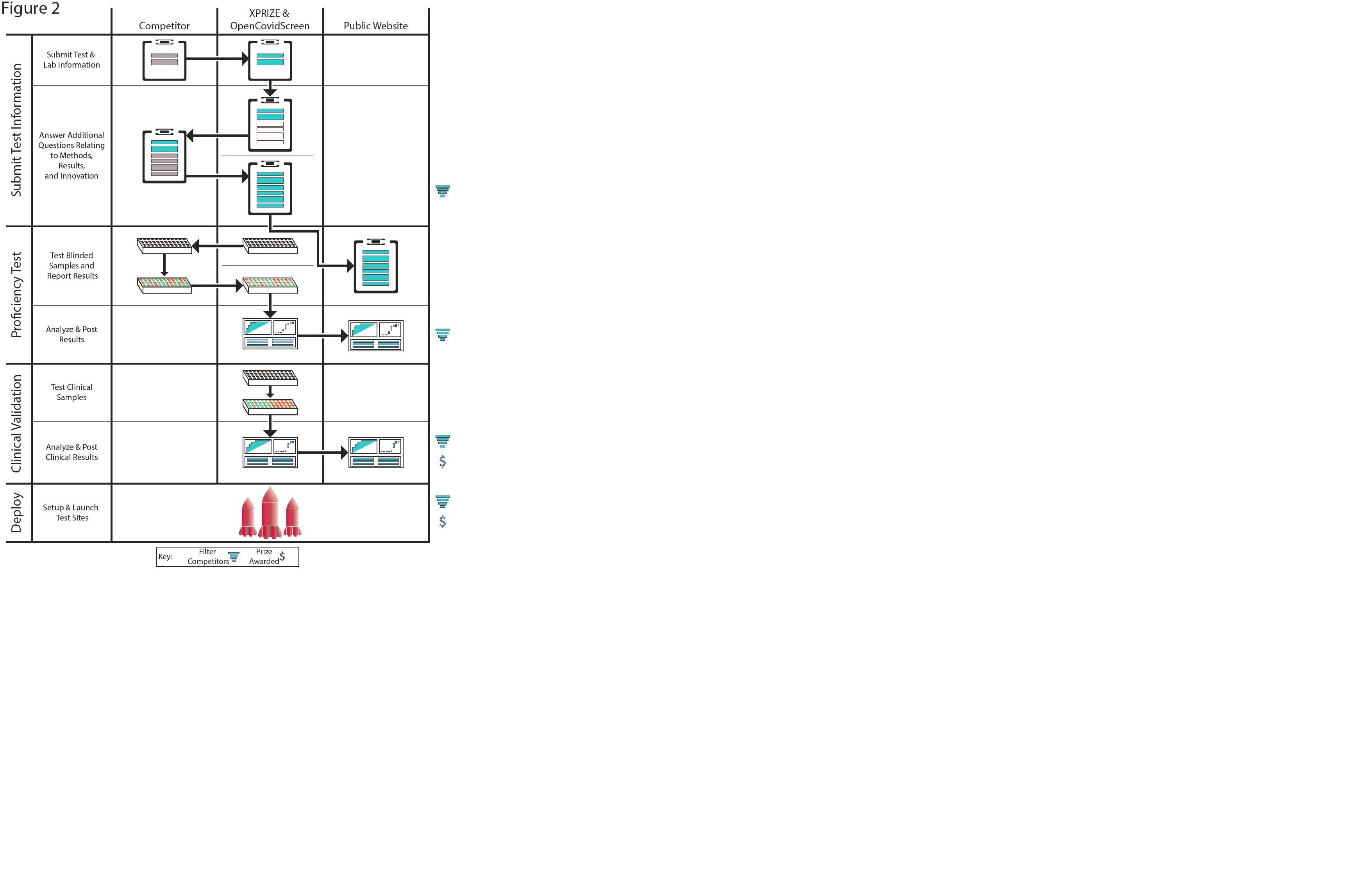
We invite readers to submit solutions for this XPRIZE, which is open to participants from all around the world (https://xprize.org/testing). This prize can serve as a springboard for both new and established technologies that can enable truly global viral testing and surveillance. Successful methods will depend on the availability of reagents, resources and automation, and as such, these metrics will also be used to identify the finalists. Winning methods will help regions increase testing capabilities by orders of magnitude and thus empower schools, businesses and cities to rapidly reopen safely, as well as pioneer technologies and platforms that can be used for future outbreaks. It is crucial to deploy rapid, scalable methods capable of tracking viruses to mitigate their detrimental impacts on society. All are encouraged to help in this fight.
Matthew J. MacKay1,2,3, Anna Hooker4, Ebrahim Afshinnekoo1,2,3, Marc Salit4, Jason Kelly5, Jonathan V. Feldstein6, Nick Haft6, Doug Schenkel7, Subbu Nambi7, Patrick Cai8, Feng Zhang9, George Church9, Junbiao Dai10, Cliff L. Wang11, Jeff Huber11, Hanlee Ji4,12, Alison Kriegel13, Anne L. Wyllie14 & Christopher E. Mason1,2,3,15†
1Department of Physiology and Biophysics, Weill Cornell Medical College, New York, NY, USA. 2The HRH Prince Alwaleed Bin Talal Bin Abdulaziz Alsaud Institute for Computational Biomedicine, Weill Cornell Medical College, New York, NY, USA. 3The WorldQuant Initiative for Quantitative Prediction, Weill Cornell Medicine, New York, NY, USA. 4Stanford University School of Medicine, Stanford CA, USA. 5Ginkgo Bioworks, Cambridge, MA, USA. 6Resilience Health, New York, NY, USA. 7Cowen and Company, Boston, MA, USA. 8Manchester Institute of Biotechnology, University of Manchester, Manchester, UK. 9Harvard Medical School, Cambridge, MA, USA. 10Shenzhen Institutes of Advanced Technology, Shenzhen, China. 11OpenCovidScreen, San Francisco, CA, USA. 12Stanford Genome Technology Center West, Palo Alto, CA, USA. 13Medical College of Wisconsin, Milwaukee, WI, USA. 14Department of Epidemiology of Microbial Diseases, Yale School of Public Health, New Haven, CT, USA. 15The Feil Family Brain and Mind Research Institute, New York, NY, USA.
†e-mail: chm2042@med.cornell.edu
References
- Arnaout R. et al. SARS-CoV2 testing: the limit of detection matters. Preprint at bioRxiv https://doi.org/10.1101/2020.06.02.131144 (2020).
- Whitman J. D. et al. Test performance evaluation of SARS-CoV-2 serological assays. Preprint at medRxiv https://doi.org/10.1101/2020.04.25.20074856 (2020).
- The COVID-19 testing debacle. Nat. Biotechnol.38, 653 (2020).
- Khailany, R. A., Safdar, M. & Ozaslan, M.. Gene Rep. 19, 100682 (2020).
Acknowledgements
We thank Scott A. Jackson, Peter M. Vallone, and Kenneth D. Cole from the National Institute of Standards and Technologies (NIST). We thank Testing for America (501c3) for their support, the OpenCovidScreen Foundation, Mayor Cory Mason, the Bert L and N Kuggie Vallee Foundation, Igor Tulchinsky and the WorldQuant Foundation, Bill Ackman and Olivia Flatto and the Pershing Square Sohn Cancer Research Alliance, Ken Griffin and Citadel, the US National Institutes of Health (R01AI125416, R21AI129851, R01AI151059), and the Alfred P. Sloan Foundation (G-2015-13964).
Competing interests
J.K. is cofounder and board member for Gingko Bioworks. C.E.M. is a cofounder and board member for Aevum, Biotia and Onegevity, as well as an advisor or compensated speaker for Abbvie, Acuamark Diagnostics, ArcBio, Bio-Rad, DNA Genotek, Genialis, Genpro, HomoDeus, Karius, Illumina, MilliporeSigma, Nanostring, New England Biolabs, Qiagen, Promega, Resilience Health, Tempus Labs, Whole Biome, and Zymo Research. A.H., H.J., C.E.M. and M.J.M. advise Resilience Health. J.H. is a board member or shareholder for Grail and Illumina. G.C. has conflicts: http://arep.med.harvard.edu/gmc/tech.html.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in