When AI Knows Too Much: Safeguarding Sensitive Health Data Becomes Critical
Published in Bioengineering & Biotechnology, Research Data, and Computational Sciences
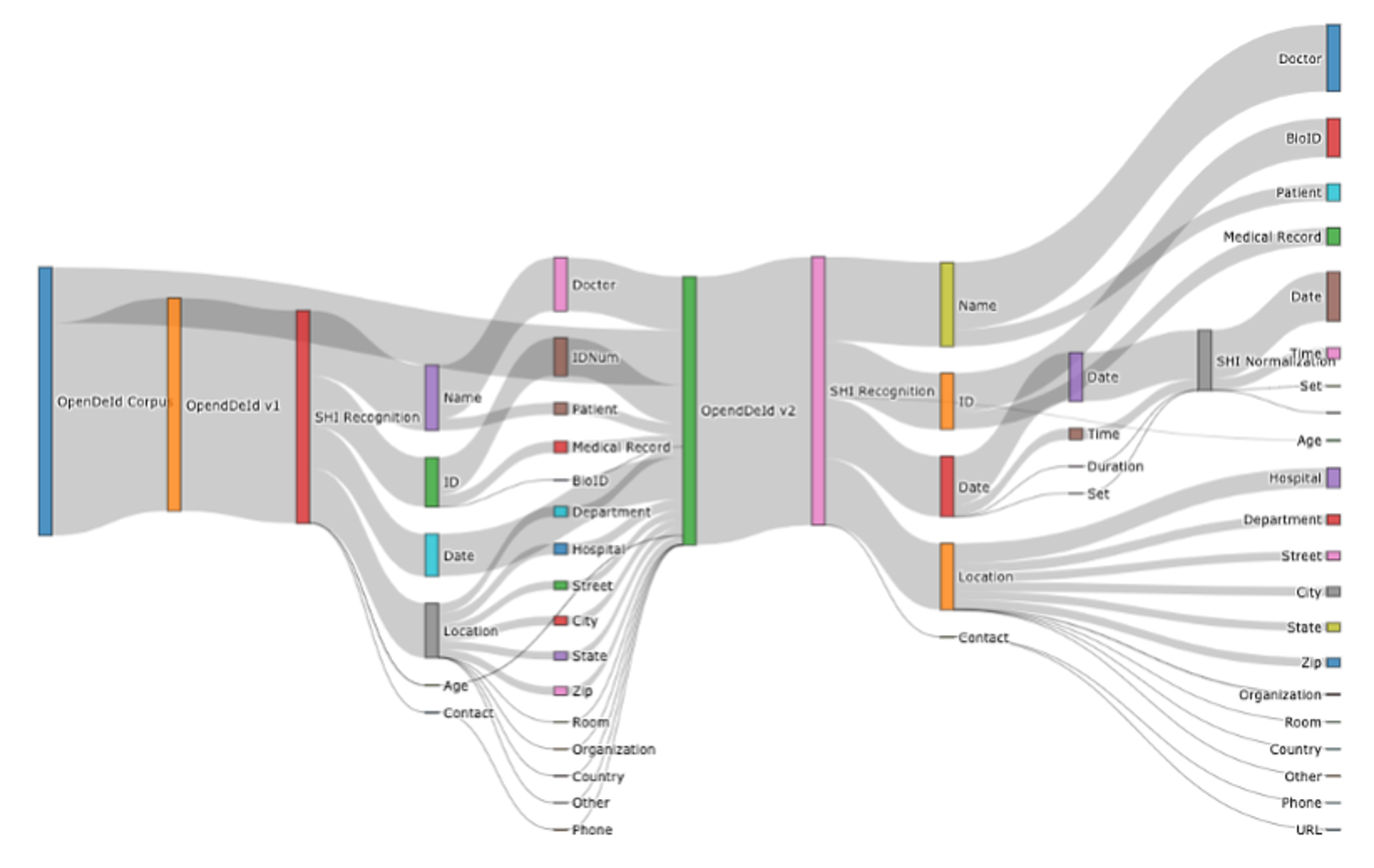
Recent advances in generative AI and NLP have positioned large language models (LLMs) as transformative tools in healthcare, enabling enhanced patient care and streamlined administrative workflow. However, significant concerns have emerged regarding user privacy, ethical considerations, and the potential leakage of sensitive confidential information. This leakage can occur either through direct interaction with the systems or through the training data used by these LLMs. On the other hand, health, medical and biomedical institutions at all levels around the world are using electronic health records (EHRs) for research. However, EHRs are often filled with private or confidential information related to patients. Fragments of information collected across various EHR systems can be used to deduce the real identity of a patient. Therefore, to properly utilize EHRs for secondary research and promote the development of innovative digital health applications, it is crucial to identify and remove patient private information.
How It All Began
To address this issue, the Ministry of Education in Taiwan sponsored a large nationwide competition, the Artificial Intelligence CUP 2023- Privacy Protection and Medical Data Standardization Challenge, through the nationwide project titled Ministry of Education Artificial Intelligence Competition and Annotation Data Collection Project. The project was organized jointly by the National Kaohsiung University of Science and Technology (NKUST), the SREDH Consortium, and Asia University. Leadership and guidance were provided by Dr Jitendra Jonnagaddallla (SREDH / UNSW), Dr Hong-Jie Dai (ISLab) and Dr Ching-Tai Chen (MLB Lab).
The Purpose Behind the Study
The aim was to seek de-identification and standardization solutions from researchers worldwide. The project brought together 721 participants, who formed 291 teams to employ various approaches in natural language processing to find a solution to the challenging problem of privacy in healthcare. The entire process was undertaken within the framework of the SREDH/AI CUP 2023 competition, hosted on CodaLab. The 291 teams explored how large language models (LLMs), both fine-tuned and prompt-based, can help detect and obfuscate sensitive health information (SHI) in pathology reports, as well as normalize temporal information to maintain usable timelines in clinical data across demographics. One of the immediate outcomes was the 2024 proceedings titled 2024 International Workshop on De-identification of Electronic Medical Record Notes (IW-DMRN), where the best-performing teams showcased their work. The proceedings of this workshop were published in 2024, by the Springer in Communications in Computer and Information Science.
Our Approach and Process
In a broader context, we adopted a three-pronged approach, focusing on corpus creation, baseline model development, and competition analysis.
-
Corpus creation
Built upon the existing OpenDeID corpus with added pathology reports to form OpenDeID-v2. SHI spans and date values were annotated using surrogate generation, ensuring consistent date shifting within a patient’s record and maintaining longitudinal integrity. -
Baseline modelling
Fine-tuned Pythia models across sizes using LoRA and full tuning; also tested prompt-based ICL setups with examples to prompt the models without further tuning.
-
Competition analysis
These baselines were compared to real entries from the participating teams. Methodological insights, such as reliance on decoder-only transformers and hybrid rule-based/LLM pipelines, were distilled from top performers.
What We Accomplished
First, we created and released a novel deidentification corpus, named OpenDeID v2, containing 3,244 pathology reports with annotations marking SHI and normalized date surrogates that preserve temporal consistency. We then used the Pythia model suite (ranging from 70 million to 12 billion parameters) as both a baseline fine-tuned system and for in-context learning (ICL) experiments. Finally, we analyzed model scaling effects by comparing LoRA (parameter-efficient tuning) and full-parameter tuning methods, and their impact on performance, particularly noting how models with more than 6 billion parameters sometimes overfit on this moderate-sized dataset.
Key Insights and Discoveries
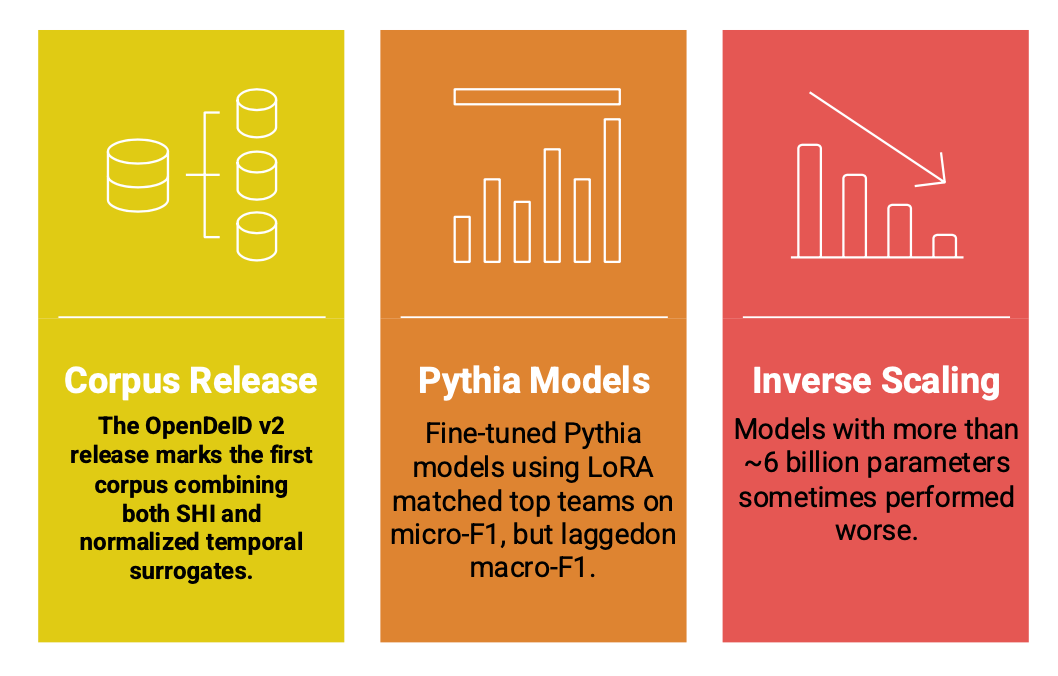
- The OpenDeID v2 release marks the first corpus combining both SHI and normalized temporal surrogates, which is significant for researchers building compliant clinical NLP systems.
- Fine-tuned Pythia models using LoRA matched top teams on micro-F1, but lagged on macro-F1, especially in Subtask 2 (temporal normalization task), highlighting uneven performance across diverse SHI types.
- Interestingly, the inverse scaling phenomenon emerged; models with more than ~6 billion parameters sometimes performed worse, likely because the dataset, although well-annotated, was not large enough to fully leverage the massive model capacity.

This study highlights the importance of data quality and methodological nuances in clinical NLP. The OpenDeID v2 corpus fills a critical gap, enabling more accurate benchmarking and research on consistent temporal de-identification. The scaling analysis cautions against the common assumption “bigger is always better”. Additionally, the performance gap between micro- and macro-F1 suggests that while overall accuracy is important, handling rare or variable SHI categories requires more sophisticated strategies, possibly ensembles, data augmentation, or hybrid rule-based methods.
Study Progress Summary
Our study offers meaningful value to both researchers and clinicians by contributing to the expansion of the corpus and diversification of data sources to better support the fine-tuning of large models. Building upon the success of the SREDH/AI-CUP 2023 competition and the subsequent 2024 workshop proceedings, we expanded our work to incorporate multimodality by integrating doctor-patient audio consultations, further enhancing privacy in healthcare data. This second phase of the study followed a similar approach, involving a competition (hosted on CodaBench), a 2025 workshop, and forthcoming proceedings that will document the outcomes. The international seminar, serving as the closing event for the competition, was held during MedInfo 2025, featuring presentations from the top-performing teams. Building on the outcomes of the first International Workshop on Deidentification of Electronic Medical Records Notes (IW-DMRN, January 2024), the second workshop aimed to advance AI algorithms capable of effectively identifying and removing SHI from medical speech datasets, promoting the development of robust privacy-preserving AI systems in healthcare.
Future Directions and Opportunities
Possible avenues for future studies include expanding deidentification and temporal-normalization methods to more diverse clinical text sources, such as multi-institutional, multilingual, and varied EHR note types, to enhance generalizability. Another important direction is improving the processing of complex temporal expressions, particularly relative or ambiguous time references, through more advanced hybrid or temporally aware models. Future research may also examine the privacy robustness and interpretability of LLM-based deidentification systems to ensure regulatory compliance and reliable surrogate generation. Additionally, studies could explore the use of synthetic clinical text to address data scarcity and strengthen model performance while preserving patient privacy. The data and baseline models used in this work can also be requested through the provided link (OpenDeID), enabling further research and replication.
Closing Thoughts
This study provides both a valuable corpus and deep insights into how model scale, data, and methodology interact in the sensitive yet essential task of clinical deidentification, offering a roadmap for safer, more privacy-conscious medical AI.
Follow the Topic
-
npj Digital Medicine
An online open-access journal dedicated to publishing research in all aspects of digital medicine, including the clinical application and implementation of digital and mobile technologies, virtual healthcare, and novel applications of artificial intelligence and informatics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Artificial Intelligence in Sports Medicine
Publishing Model: Open Access
Deadline: Jul 21, 2026
Impact of Agentic AI on Care Delivery
Publishing Model: Open Access
Deadline: Jul 12, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in