Writing Microbial Memory into Intestinal Stem Cells
Published in Microbiology, Cell & Molecular Biology, and Immunology
Our interest in this work began years ago with a discovery that reshaped how we think about intestinal development. During the suckling period, intestinal stem cells undergo striking changes in DNA methylation - a chemical modification of DNA that is one of the best-understood epigenetic mechanisms and can be copied as cells divide. Development, we realized, was not just morphological; it was epigenetic. And when we removed the microbiota, part of that program stalled.
At the time, we focused on a handful of candidate genes. We saw that bacteria mattered, but we did not know how broadly they shaped the epigenome, or whether those changes represented something more lasting than a temporary response.
What we were really searching for was memory.
By epigenetic memory, we mean durable changes in gene regulation that persist after the original signal fades - changes that are maintained as stem cells divide and passed on to the cells they produce. To demonstrate that kind of memory, we needed to study purified, long-lived stem cells and remove every confounding variable we could.
That realization shaped the study.
We used Lgr5 reporter mice to isolate pure intestinal stem cells. To truly understand the role of microbes, we rederived the strain under germ-free conditions. Pups were delivered by timed cesarean section and raised by germ-free foster mothers. Months of careful validation ensured completely no bacteria or other microorganisms before experiments began. Establishing this system required patience, but it gave us clarity.
Diet was another often-overlooked variable. Germ-free animals are fed sterilized diets to eliminate bacterial contamination, but sterilization breaks down nutrients such as folate and certain vitamins, which must be supplemented. To avoid nutritional confounding, we fed all mice - regardless of microbial status - the same sterilized diet. Only microbial exposure differed.
What we found shifted our thinking.
The most pronounced microbiome-dependent epigenetic remodeling did not unfold gradually across early life. Instead, it was concentrated at weaning - a brief developmental window. As solid food was introduced, microbes expanded, triggering a transient immune surge known as the weaning reaction, and at the same time, intestinal stem cells underwent a focused wave of epigenetic change.
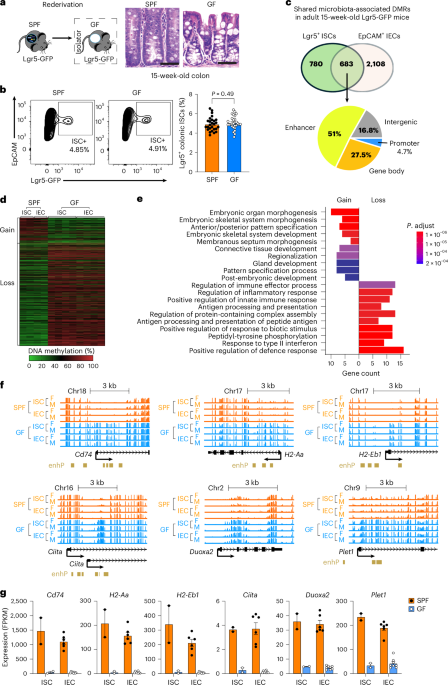
The key discovery was unexpected: after weaning, enhancers controlling MHC class II genes - crucial for how cells communicate with the immune system - lost DNA methylation, but only in the presence of microbes. It was a precise remodeling event centered on immune regulatory elements and confined to this narrow window of development. The trigger was brief. During the weaning reaction, gram-positive bacteria induced a short pulse of IFN-γ, and that signal was enough to initiate demethylation at these enhancers. The signal did not need to be sustained - a transient encounter was sufficient to set the process in motion.
Yet the consequences were lasting.
Because these changes occurred in long-lived stem cells, the remodeled enhancer landscape persisted into adulthood. The immune signal faded, but the regulatory configuration remained. Stem cells retained that information and passed it to the epithelial cells they generated.
To better understand how this worked, we turned to organoid models - miniature epithelial tissues grown from stem cells. In this simplified setting, we could focus purely on epithelial behavior. What we observed was remarkable: epithelial cells, like immune cells, were capable of “remembering” prior stimulation.
After an initial exposure, cells derived from previously stimulated stem cells mounted a stronger response to a second challenge. This heightened reaction was not due to lingering microbes or cytokines; it reflected the enhancer remodeling established earlier during weaning.
Timing proved critical. Altering microbial or immune signals during weaning changed the epigenetic outcome, whereas similar interventions in adulthood had little effect. The window opened – and then it closed.
This became the conceptual turning point.
Weaning is not simply a dietary transition. It is a developmental pivot point. As gram-positive microbes expand, they trigger a brief wave of immune activation. That transient IFN-γ signal drives stable demethylation of MHC II enhancers in intestinal stem cells. In doing so, microbial exposure becomes embedded in the regulatory architecture of the tissue.
What began as an extension of our earlier findings ultimately reframed how we think about host-microbe interactions. The microbiome does more than influence physiology in real time - during a defined developmental window, it helps establish the epigenetic foundation of mucosal immunity. Signals encountered during this period do not simply fade; they become part of the stem cell program, shaping gut function long after the original exposure.
In this way, early microbial cues leave a lasting imprint on epithelial cells, positioning them as innate immune sentinels with a form of trained transcriptional memory. Decoding this early-life dialogue between microbes and stem cells may open new ways to treat intestinal disease.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in