A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer
Published in Cancer

Over the last years, immunotherapy using immune-checkpoint blockage (ICB) revolutionized cancer treatment in terms of quality of life and life years saved. By now, hundreds of clinical studies investigate the potential of ICB therapy in multiple cancer types. In breast cancer (BC), it was recently shown that ICB therapy combined with neoadjuvant chemotherapy improves pathological complete response. Despite the ground-breaking clinical success, ICB therapy responses are variable and unpredictable, while resistance, relapse and side- effects often occur. As societal, financial and emotional burdens are increasing, the need for a conclusive and clinically applicable predictive biomarker becomes more and more clear. Unfortunately, many predictive biomarkers have been proposed yet proven inconclusive. In BC, the combined treatment with neoadjuvant chemotherapy and ICB therapy imposes additional difficulties in disentangling effects of ICB versus chemotherapy. A better understanding of the breast tumor and its tumor micro-environment (TME) during ICB with and without neoadjuvant chemotherapy is thus paramount to identify and implement new biomarkers in ICB-based BC treatment.
To highlight the importance of tumor biopsies in answering these key questions, we invite you to discover their journey.
It all started when we set up a window-of-opportunity study (BIOKEY) to characterize the immune environment of BC patients by immunohistochemistry. BIOKEY allowed early breast cancer patients to receive one dose of ICB therapy (pembrolizumab; anti-PD1; n=29) or neoadjuvant chemotherapy prior to ICB therapy (n=11) approximately 10 days before their surgery. Paired biopsies were collected immediately prior to anti-PD1 treatment (‘pre-treatment’) and during subsequent surgery (‘on-treatment’). Soon after initial testing, we were confronted with the challenge of picking up differences in immune composition via immunohistochemistry slides shortly after ICB therapy, and decided to implement single-cell RNA sequencing for the detailed evaluation of the tumor micro-environment. In 2017, we founded a unique collaboration between experienced clinicians of UZ Leuven and excellent researchers of KU Leuven. This ambitious project was funded by VIB and the Flemish government through the VIB Grand Challenges program. We realized effective communication and synchronized efforts from both teams would be key to make this unique project successful, and therefore clearly defined the journey the biopsy had to follow during the subsequent steps of our protocol. The biopsy, freshly collected in the surgery room, had to arrive in the research lab in less than 30 minutes to allow for sample preservation; a challenge overcome thanks to the close proximity of the research labs and surgery rooms in our institution (Gasthuisberg, UZ Leuven, Leuven, Belgium). Practically, one team member collected the BC tissue in the surgery room, racing through the corridors of the hospital to deliver the sample as soon as possible to the department of pathology. There, the sample was inked and prepared for diagnostic purposes, and a small piece of the tumor was cut off for the single-cell analyses.
After a second race against the clock and through the corridors, the tumor sample arrived at the research laboratory. The sample was immediately dissociated into single-cells, ready for the next adventure in the 10x Genomics machine. Inside, every cell is captured and tagged individually in its bubble for transcriptome, T-cell receptor (TCR) or proteome sequencing. The resulting sequencing data is mapped to the reference genome, producing a massive amount of gene expression data for each individual single cell. Bio-informatic pipelines are applied for cell clustering, the process of dividing cells in different clusters based on differences or similarities in their transcriptomic profile. Celltype-specific marker genes allow to annotate these clusters to their according celltype and give an identity to each cell. TCR analyses enabled us to define T-cells that expand in response to anti-PD1 therapy. As this T-cell clonotype expansion could serve as a surrogate for response to treatment, we divided patients in so-called expanders (‘responders’) and non-expanders (‘non-responders’) for downstream analyses.
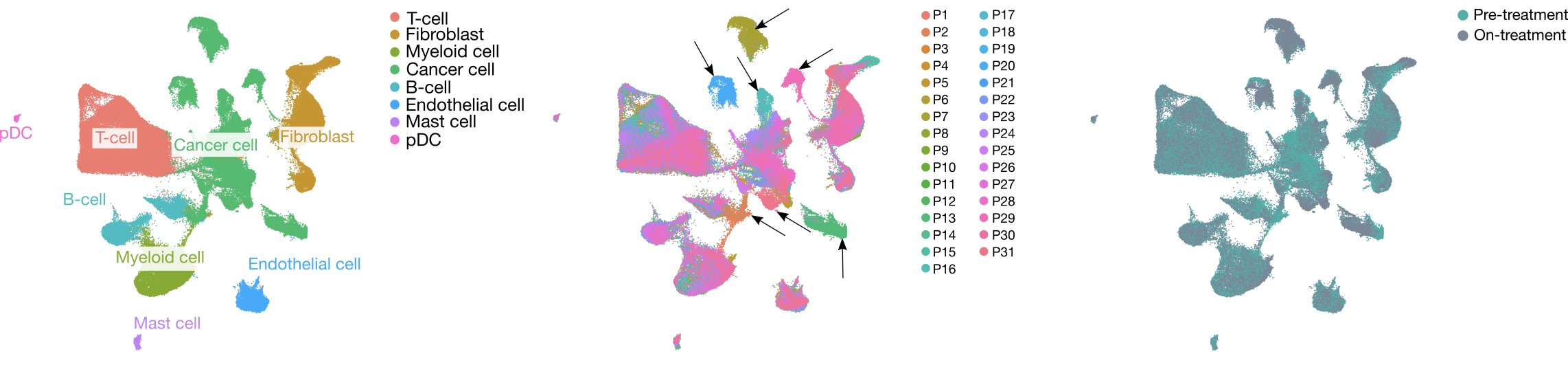
In our manuscript, we provide a detailed single-cell map of the cells in the TME before ICB therapy and report the dynamic changes implemented by this therapy. We highlight immunophenotypes and associated gene sets that are positively or inversely correlated with T-cell expansion after neoadjuvant anti-PD1 therapy in patients with early BC. Importantly, by shedding light on the heterogeneity of response to anti-PD1 therapy in BC, we stress the importance of biomarkers discriminating patients based on characteristics of the TME. As such, this study is not only important for the identification of novel biomarkers predictive for response to ICB therapy, but also to unravel the mechanisms driving resistance. Hence, the results of this paper provide an excellent resource of potential biomarkers and therapeutic targets for novel combinational therapies. While follow-up studies are needed to confirm our findings and validate potential therapeutic targets, we hope these insights lay the foundations to improve patient care and bring a biomarker for ICB response to the clinic.
The journey of the starring biopsies started with the real heroes; patients willing to participate in our window-of-opportunity study and contribute to research projects like ours. We sincerely hope that the results obtained through the combined efforts of our research teams and participating patients will benefit that same group of BC patients and many more in the near future. From bedside to bench and beyond!
Follow the Topic
-
Nature Medicine

This journal encompasses original research ranging from new concepts in human biology and disease pathogenesis to new therapeutic modalities and drug development, to all phases of clinical work, as well as innovative technologies aimed at improving human health.
Related Collections
With Collections, you can get published faster and increase your visibility.
Stem cell-derived therapies
Publishing Model: Hybrid
Deadline: Mar 26, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in