A tunable orthogonal coiled-coil interaction toolbox for engineering mammalian cells
Published in Bioengineering & Biotechnology
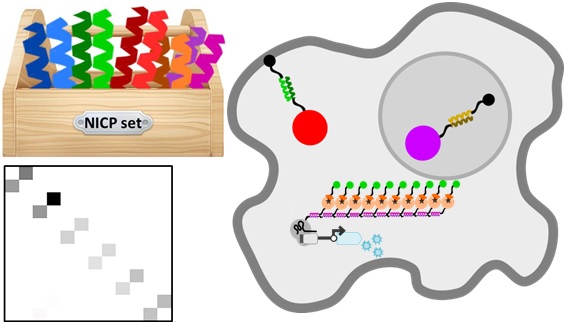
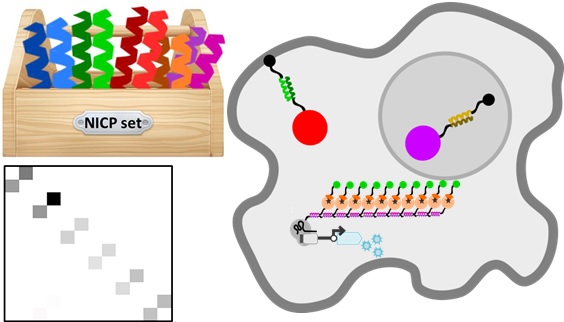
Protein-protein interactions are involved in almost all cellular processes, they define scaffolds, regulate cellular localization and participate in functioning in cellular machines, such as those performing transcription, replication, splicing and others. In order to manipulate cellular processes we need a reliable set of protein-protein interaction modules that is orthogonal, meaning that those modules minimally interact with the existing cellular components and other members of the set. The modules that we investigated here are de novo designed coiled-coil (CC) dimers whose specificity is defined by a combinatorial combination of electrostatic and hydrophobic interactions. Those are some of the smallest polypeptide dimers whose specificity rules are quite well understood and we have used them before to design fast responsive signaling pathway based on proteolysis (SPOC) and a new type of protein folds, coiled-coil protein origami, where the structure is defined by pairs of CC modules that self-assemble into polyhedra unknown in nature within a single polypeptide chain.
Coiled-coil dimers are also used in nature, particularly to encode leucine-zipper transcription factors in yeast, plants and animals. Using natural CC dimers introduces the danger of crosstalk with some natural components as indicated by the analysis of the interaction partners in human proteome. Proteome interaction screening of de novo designed modules (NICP set) that have been used previously for the in vitro assembly of new protein fold topologies and on the other hand indicated much lower number of predicted partners in the human proteome and suggested good orthogonality, therefore we decided to investigate if we could use those modules as tools to regulate mammalian cells.
Indeed the analysis of pairwise interactions of 12 CC peptides that formed six pairs in the complex environment of mammalian cells confirmed very high orthogonality with typically three orders of magnitude preference for the correct pairs over the background. Moreover the affinity of each CC pair can be tuned in an almost continuous manner by modifying residues at noncontact sites and by combining several peptides, to tune the stability within the 100-fold range. Employing several CC pairs in the same time (multiplexing) within the same cell was demonstrated by their use as cellular compartment localization tags, where different fluorescent proteins demonstrated localization defined by CC-pairing specificity.
Coiled coil modules (the same or different peptide) can be used to tune transcriptional regulation where the selected number of peptide repeats was concatenated into a single chain, which precisely defined the binding stoichiometry. We demonstrated that the concatenated CC peptide tag (CCCtag) could amplify the transcription of selected reporter or genomic gene targets based on CRISPR/dCas9/gRNA guiding and achieved superior potency in comparison to several previously designed strategies, such as SunTag and SamTag. Moreover, the similar approach was used to amplify the response of the chemically- or light-regulated transcriptional activation, which strongly increased the response in mammalian cells to a chemical stimulus as well as in cells implanted into the animals.
The NICP set and its implementations therefore represent a valuable toolbox of minimally disruptive modules for the recruitment of versatile functional domains and regulation of cellular processes for synthetic biology and other uses in molecular biology. A particular advantage of this platform is that it has quite small footprint, allows define the recruitment of one or several molecular partners in a selected stoichiometry and minimal interactions with other cellular components.
Lebar, T., Lainšček, D., Merljak, E. et al. A tunable orthogonal coiled-coil interaction toolbox for engineering mammalian cells. Nat Chem Biol (2020) doi:10.1038/s41589-019-0443-y
Read the paper here: https://www.nature.com/articles/s41589-019-0443-y
The Jerala group: https://www.ki.si/en/departments/d12-department-of-synthetic-biology-and-immunology/
Department of Synthetic Biology and Immunology, National Institute of Chemistry, Ljubljana, Slovenia
Follow the Topic
-
Nature Chemical Biology
An international monthly journal that provides a high-visibility forum for the chemical biology community, combining the scientific ideas and approaches of chemistry, biology and allied disciplines to understand and manipulate biological systems with molecular precision.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in