Bat influenza viruses transmit among bats but are poorly adapted to non-bat species.
Published in Microbiology

Unlike classical influenza A viruses that spread to a broad range of different hosts by utilizing host cell sialic acid residues, we recently showed that the bat-borne subtypes H17N10 and H18N11 gain entry into host cells via major histocompatibility complex class II (MHC-II) molecules. Moreover, the discovery that they efficiently use MHC-II proteins of several species, including humans, raised the question whether these bat flu viruses are a potential threat to the human population. To address these concerns we studied the virus’ biology in detail and investigated the growth and transmission abilities in mice, ferrets and in Jamaican fruit-eating bats (Artibeus jamaicensis).
This exciting bat flu journey started with the infection of susceptible cells in tissue culture. Although simple, this experiment had far-reaching consequences. To our great surprise, we observed the rapid appearance of better replicating virus mutants that harbored mutations in the H18 hemagglutinin head and expressed a truncated N11 neuraminidase protein either by the insertion of a premature stop codon or a partial deletion of the N11 open reading frame. Fascinated by these findings, we then dug deeper into the mechanistical role of N11. We found that its function is to facilitate viral release from the cell surface by presumably downregulating MHC-II surface expression. Importantly, functional N11 was only required in context of wild-type H18 but dispensable once H18 had acquired sufficient adaptive mutations.
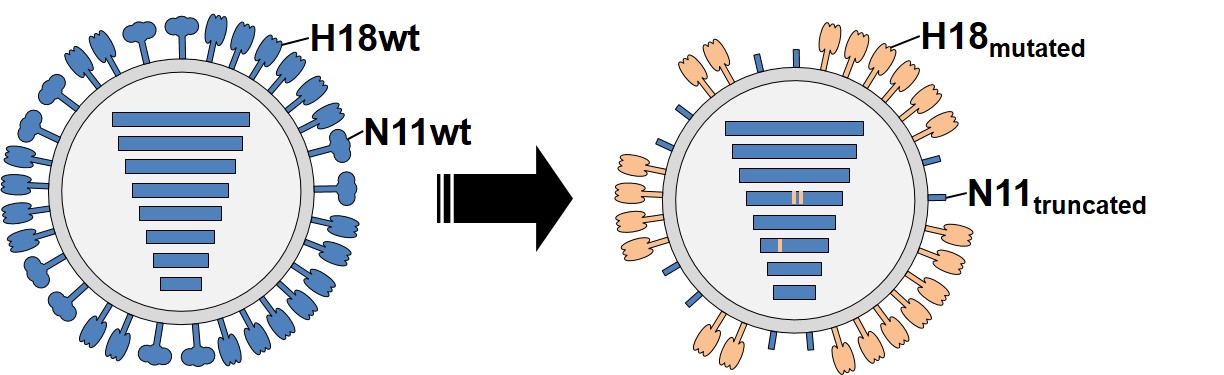
At this point we could not exclude the possibility that this unexpected flexibility to replicate to high titers without a functional N11 might also be possible in vivo. Therefore, we refined our study not only to test the wild-type H18N11 virus, but also the mutant variant harboring two mutations in H18 and a truncated N11 (due to a premature stop codon) in vivo. In mice both wild-type and mutant virus demonstrated the same constrained tropism to the upper respiratory tract, whereas in ferrets only the mutant was found to replicate in various tissues including immune competent compartments such as the tonsils. However, neither in mice nor in ferrets was virus transmitted to naïve cagemates. Now, there was only one urgent question to answer that had not been addressed since the virus’ discovery: is the bat-borne virus indeed infectious in bats? Coincidently, groups led by Tony Schountz at Colorado State University and Wenjun Ma at Kansas State University had found that Jamaican fruit-eating bats, close relatives of the initially identified host species and a likely reservoir host, are susceptible to H18N11. Following infection with the wild-type virus, the bats shed high viral loads via the fecal route and readily transmitted to naïve contact animals. Consequently, we began to collaborate on this project and determined that the follicle associated epithelium of the jejunal Peyer’s patches were found to be the infection hot spot. And what happened to the N11-deficient virus in bats? Although infection with the mutant variant resulted in a productive infection in index animals, viruses with a restored N11 rapidly emerged and were transmitted to contact bats.
In summary, the zoonotic potential of bat flu seems to be rather low, considering ferrets as the best available model to study these bat-derived flu viruses. This work is the result of a brilliant collaboration between several groups in Germany and the United States sharing complimentary expertise. Please see the paper and let us know what you think.
Link to the paper: https://www.nature.com/articles/s41564-019-0556-9.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in