Catestatin antimicrobial activity shapes the colonic microbiota composition
Published in Microbiology
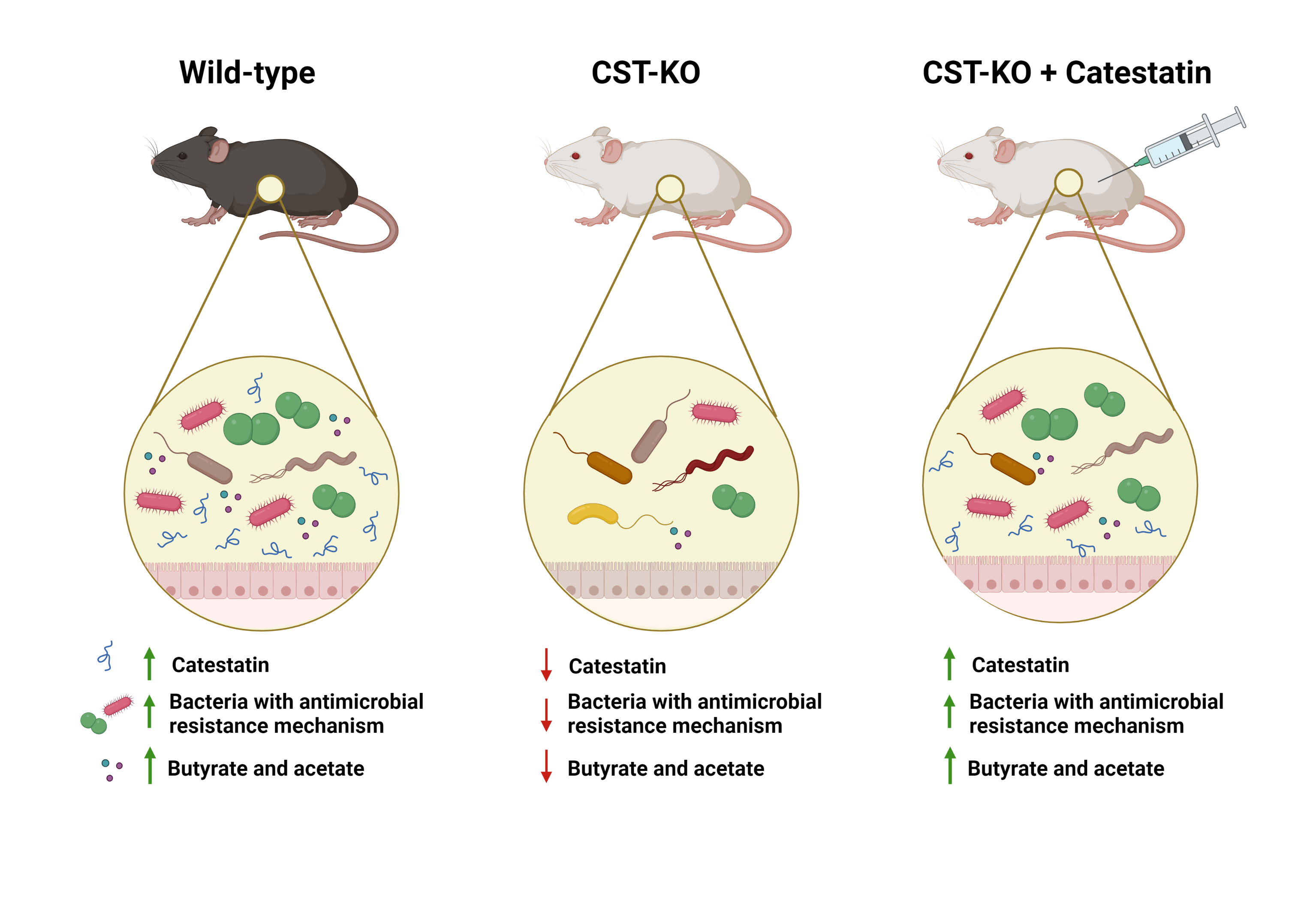
Our group, Host-microbe metabolic Interactions, strives to understand the chemical dialogue between the gut microbes and their host, and how this cross-talk maintains a mutual environment. For example, the gut epithelium, secretes a variety of antimicrobial peptides. Antimicrobial peptides play an important role in protecting against infection while maintaining intestinal homeostasis by supporting mutualistic interactions with the gut microbiota. In turn, the gut microbiota produces metabolites that directly regulate Antimicrobial peptides production to orchestrate immune responses. Among these antimicrobial peptides, is catestatin, which is a degradation product of chromogranin A, a neuroendocrine secretory protein present in a subtype of the epithelial cells, known as enteroendocrine cells. Our publication unraveled the crucial role of catestatin in altering the microbiota composition. The colonic microbiota profiling indicated that the absence of catestatin is associated with a lower abundance of short-chain fatty acid (SCFA-producing bacteria such as Roseburia and Intestinimonas and, in turn, lower levels of SCFAs, in particular, butyrate and acetate. However, the supplementation of catestatin to a catestatin -depleted environment did not promote exclusively health beneficial microbiota, but rather led to a more equilibrated composition, including pathobiont taxa. Although changes in the microbiota composition have been observed in healthy wild-type mice treated with catestatin, the changes were much less prominent and were dominated by the colonization of gut bacteria with a capacity to resist the antimicrobial effect of catestatin that became more abundant upon catestatin treatment.
A significant increase in the abundance of taxa that harbour the catestatin peptide-resistance enzyme, phosphoethanolamine transferase (eptA) was observed in catestatin-knock out and wild-type mice treated with catestatin. The latter finding implies that catestatin may regulate the microbiota composition via its antimicrobial properties and that during its depletion, the microbiota was not primed with catestatin, thus do not evolve mechanisms to tolerate its antimicrobial effect.
As a defence mechanism to the production of antimicrobial peptides, the gut microbes have evolved antimicrobial resistance mechanisms to counteract the killing effect of the secreted peptides. Here, we found members of the gut microbiota to harbour proteases with the capacity to degrade catestatin. For example, the gut bacteria, E. coli, was able to degrade catestatin via the action of its outer membrane protease omptin, which is known to cleave other cationic antimicrobial peptides. Although the resulting peptides were not further tested, we speculate that omptin may cleave at arginine–glycine residues in catestatin, yielding cateslytin and a hexapeptide. However, other different peptides can emerge due to the proteolytic process of catestatin in the intestinal lumen from different members of the gut microbiota in specific homeostatic or pathogenic conditions.
Findings from our publication are of significant importance to diseases associated with altered levels of intestinal antimicrobial peptides, such as inflammatory bowel disease, where altered levels of endogenous antimicrobial peptides have been reported, and the administration of these peptides has ameliorated intestinal inflammation in murine models of the disease.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in