Cell-Based Glycan Arrays for Probing Glycan–Glycan Binding Protein Interactions
Published in Chemistry

The paper in Nature Communications is here: http://go.nature.com/2F4Y5n7
(Peng Wu) In 2002 when I was still a graduate student, the development of glycan microarray technologies was reported by Denong Wang, Milan Mrksich and Ten Feizi. In the past 16 years, glycan microarrays have been used to analyze substrate scope and specificity of glycosyltransferases, assess host receptor specificity of human viruses, and compare the antibody repertoire in normal serum with serum of patients with various diseases to identify glycan-specific biomarkers, just to name a few specific examples.
Today, most glycan microarrays, including the ones used by the Consortium for Functional Glycomics, are fabricated by coupling individual sugars synthesized chemically to N-hydroxysuccinimide ester functionalized glass slides. The construction of such microarrays is time-consuming and requires highly specialized experts. It is difficult for scientists who are interested in studying the biological functions of glycans to fabricate such microarrays in their own labs. In addition, glycans attached to a solid support do not fully recapitulate their natural arrangements on the cell surface.
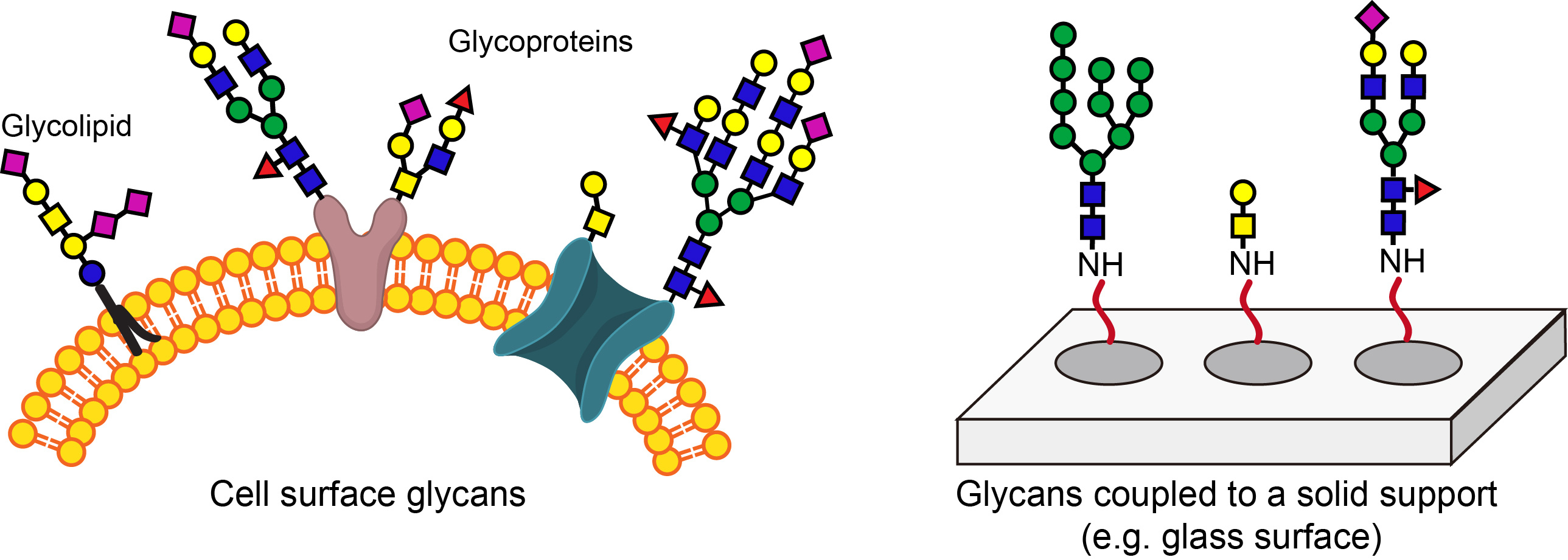
To address these problems, I came up with an idea to create glycan arrays directly on the cell surface. Because my lab has extensive experiences with CHO glycosylation mutant Lec2 cells, I asked Jennie Briard and Hao Jiang (now an Associate Professor at Ocean University of China) to find out if we could use these cells as the starting platform to construct such arrays. Lec2 cells have a comparatively narrow repertoire of cell-surface glycan epitopes and do not express periphery sialic acid and fucose. Therefore, sialyltransferase and fucosyltransferase, respectively, can be used to add sialic acid and fucose or their analogs in situ to the cell surface of these cells to create cell arrays displaying a diverse array of natural and unnatural sialosides and fucosides. By choosing a sialyltransferase or a fucosyltransferase with different specificity, sialic acid or fucose can be introduced to the cell surface with different linkages.
Jennie and Hao demonstrated that such arrays could be constructed in a few hours using tools and instruments that are found in most biology labs. Importantly, they confirmed that these arrays could be used to assess glycan-binding specificity of glycan-binding proteins and the readout produced as fluorescent signals could be easily processed using a flow cytometer compatible with 96-well microplates.
After we confirmed that we could using in situ glycosylation to add sugars directly to the cell surface to form cell-based glycan arrays, we were asking ourselves what kind of applications this new technology could be used for. Matthew Macauley, at the time working on the same floor, told me that probably we could try to use it to search for high-affinity and selective ligands for Siglec 15—a sialic acid-binding lectin involved in osteoclast differentiation. Jennie then made a cell-based array displaying unnatural sialic acid analogs via in situ sialylation and a biocompatible click reaction—the ligand assisted CuAAC. Screening this array against a panel of recombinant human Siglecs made by Matt, Jennie found a few high-affinity ligands for Siglec 15, among which one was also selective. Exposure of human osteoprogenitor cells to Lec2 cells displaying a high-affinity Siglec-15 ligand during differentiation suppressed osteoclast formation. We were very excited about our result, which suggested that this ligand can be further optimized for clinical evaluation.
(Matthew Macauley) While a member of the Paulson laboratory, at The Scripps Research Institute, I became well acquainted with the utility of the glycan microarray and its ability to advance our understanding of the specificity of glycan-binding proteins. However, I noticed that for some glycan-binding proteins, the glycan microarray just didn't seem to product any binding. Was it that the affinity of the glycan-binding proteins for their glycan ligands was too low, such that they were lost during washing steps? Or was it that even the impressive 600 glycan CFG library of glycans didn’t have the biologically-relevant glycans for the particular glycan-binding protein under investigation? While I am still uncertain what the answer(s) are, it is clear that taking advantage of the natural glycan scaffolds on the cell surface does ensure that the diversity cellular glycans is covered. For this reason, I am very excited about the cell-based glycan array that we have developed and see the broad potential in its use to study glycan-binding proteins.
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
I remember our lab using a glycan array to test influenza virus - sialic acid binding back in 2005 - great to see how the technology has moved on!