DeMyStifying the role of DMS during predator-prey interactions
Published in Microbiology

Written by Adva Shemi and Assaf Vardi
Elucidating the chemical signals that facilitate microbial predator-prey interactions is fundamental for our understanding of how marine food webs function. In the ocean, minute photosynthetic cells called phytoplankton serve as primary producers (as plants are on land). These unicellular algae produce about half of the oxygen we breathe. Their sworn enemies are a group of diverse microorganisms (mostly protists) collectively called microzooplankton, which consume about half of the phytoplankton biomass in the oceans every day. In their microscopic world, the main mode of communication with potential phytoplankton prey is chemical signaling, which is the exchange of molecules known as infochemicals. Although phytoplankton-microzooplankton interactions are of great ecological importance, the chemical nature of those infochemicals is often unknown. How do tiny predators locate prey in a highly diluted environment? Which prey-derived chemical cues mediate defense against microzooplankton? and which chemical signals attract microzooplankton to their prey? As a young PhD student in Prof. Assaf Vardi’s lab, I was highly motivated to discover such mechanisms, aiming to characterize the role of specific infochemicals during grazing interaction.
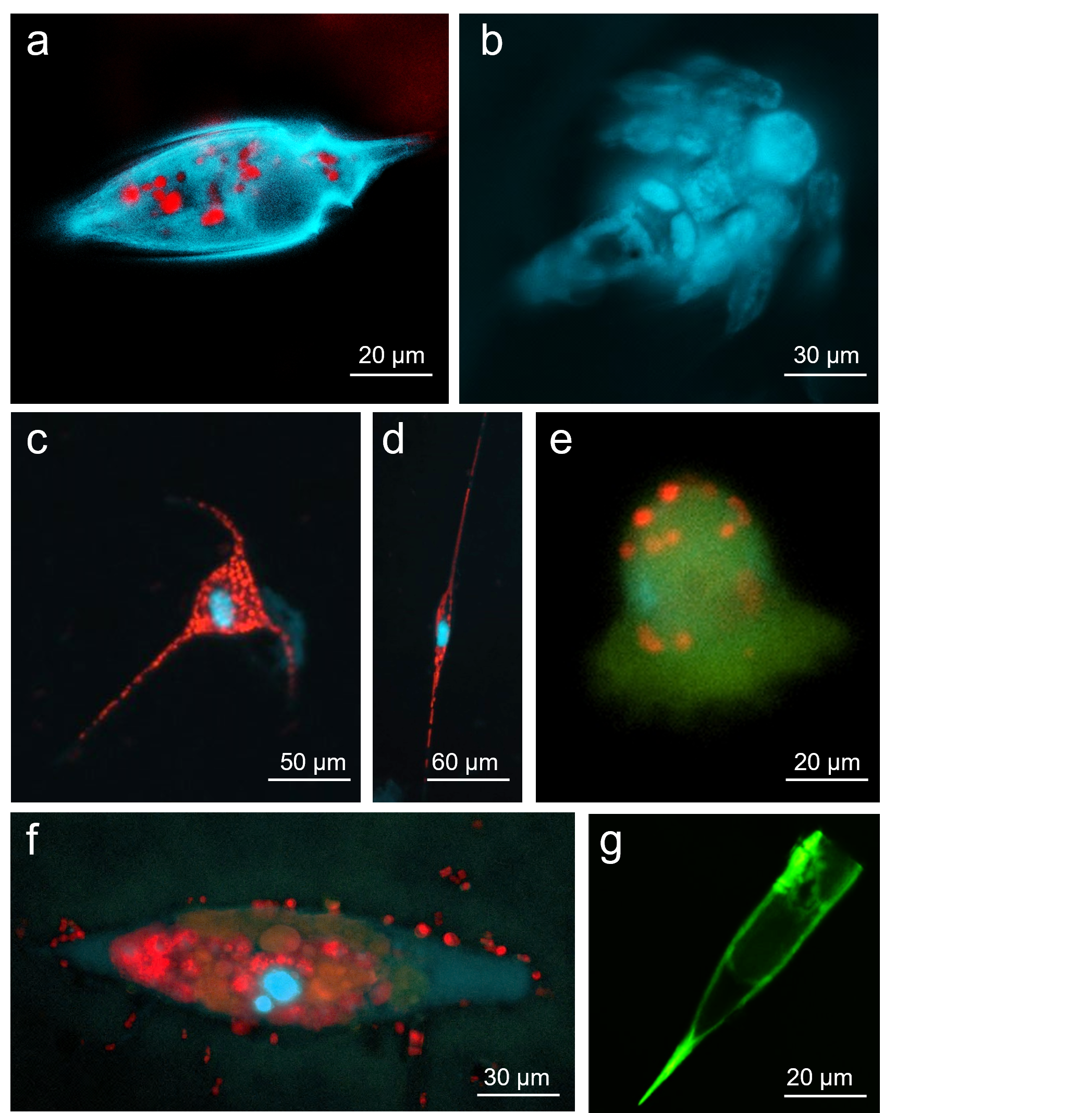
Microzooplankton, the major consumers of phytoplankton biomass in the ocean. Fluorescent micrographs of natural microzooplankton collected from the Red Sea. a, A dinoflagellate b, A copepod nauplius. c-d, Two Ceratium dinoflagellates. e, A tintinnid ciliate. f, The ciliate Tiarina. g, A tintinnid ciliate. Red represents the chlorophyll of ingested phytoplankton cells, but may also represent endogenous chloroplasts, as in mixotrophic species (c,d), or symbiotic microalgae, as in (f). DAPI nuclear stain (blue) and DiOC6, a general membrane stain (green) were used to visualize the cells.
While I was finalizing my research proposal, a fellow PhD student in the lab, Uria Alcolombri, made an important discovery. Alcolombri, a biochemist, identified the algal DMSP lyase enzyme1 (herein, the DL) that releases the volatile dimethyl sulfide, or DMS from algal cells. This sulfuric gas gives seawater its unique sweet-rotten smell, which was originally described 86 years ago by the British biochemist Paul Hass2. DMS is a significant source of sulfate aerosols in our atmosphere and therefore has an influence on the global climate3. Although DMS and its precursor (dimethylsulfoniopropionate, DMSP) were known to scientists for almost a century, the actual enzymatic conversion of DMSP to DMS was not fully understood. Alcolombri and his co-authors isolated the DL Alma1 from Emiliania huxleyi, a bloom-forming coccolithophore, and revealed that the DL gene is shared by globally distributed phytoplankton taxa and in other marine organisms. Their findings added “a missing link in the marine sulfur cycle”1.
Inspired by the discovery of the DL enzyme, I started to look into the ecological role of this important enzyme and its cleavage product, DMS, during prey-predator interactions. Interestingly, not only humans can identify the smell of DMS, but several marine animals are also sensitive to this molecule. When top-predators as whales4, penguins5 and seabirds6 smell DMS, they follow this signal and may use it to find ‘hot spots’ of plankton biomass7. But going down the food chain, the meaning of the DMS signal for primary consumers as microzooplankton is more debatable. Do tiny herbivores also attracted to DMS8 and use it to find algal prey? Or does DMS deter microzooplankton9,10 and protect the phytoplankton from their predators? While these contradicting roles for DMS were suggested during microbial predator-prey relationships so far, no direct link was demonstrated between prey cells with DL activity and reduced or enhanced grazing phenotype. Therefore, I was happy that with the isolation of the DL Alma 1, we are now able to apply genetic manipulations and perhaps clarify some of the mystery around its role during grazing. To accomplish this, we overexpressed the Alma1 gene from E. huxleyi (which is not genetically transformable) in the diatom Thalassiosira pseudonana (widespread phytoplankton and a great model for genetic transformation). The advantage of this approach was that by comparing grazing activity on the Alma1-overexpressing clones to that on the wild type T. pseudonana, we were able, for the first time, to directly characterize the effect of DL activity and DMS production on grazing. With the help of Dr. Daniella Schatz, we obtained a handful of clones with significant DMS production and started using them as prey in grazing assays with the model heterotrophic dinoflagellate Oxyrrhis marina. O. marina was often utilized to study the role of DMS/P as infochemicals11, and I was curious to know how this grazer will respond to our transgenic prey cells. The results were striking: the DMS-producing diatoms were not rejected by O. marina, but rather were consumed at least twice as fast as the wild-type cells that don’t produce DMS. These results were complemented with chemical inhibition experiments, where we treated E. huxleyi cells with a novel selective DL inhibitor12 before offering them as prey to O. marina. Following the treatment with the inhibitor, O. marina consumed significantly less prey compared to mock-treated cells. Altogether, our findings strengthen previous chemotaxis studies8 and showed that DMS in prey cells enhances uptake by microzooplankton predators.
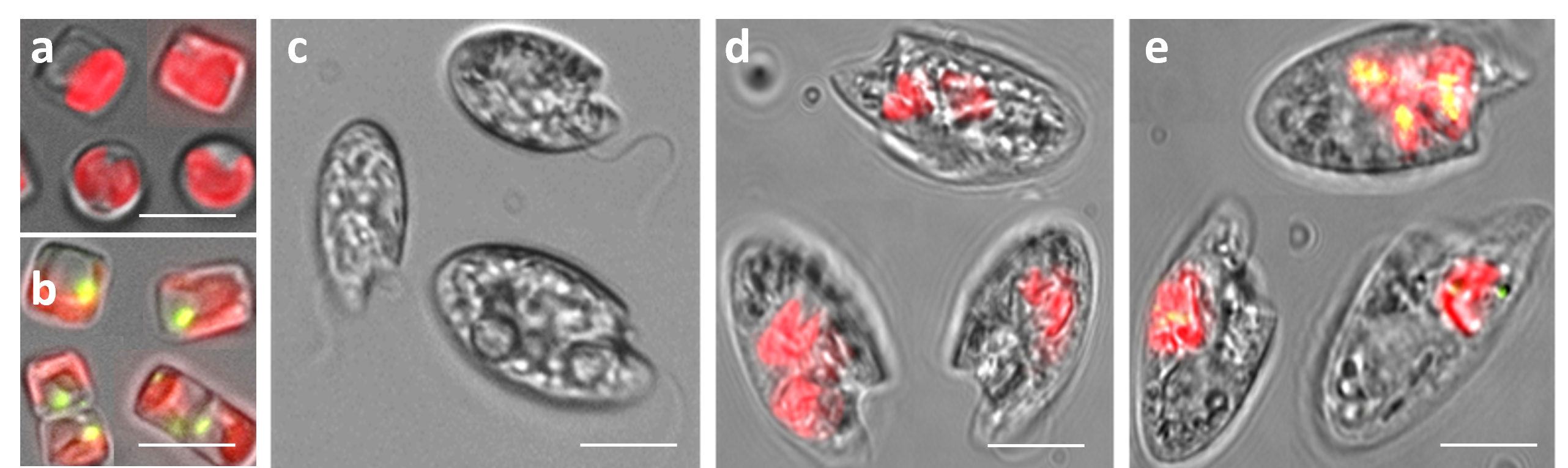
Model phytoplankton and microzooplankton used in the lab. a, Wild-type T. pseudonana. b, transgenic T. pseudonana. c, O. marina. d, O. marina with ingested wild-type T. pseudonana cells. e, O. marina with ingested transgenic T. pseudonana cells. Red represents T. pseudonana’s chlorophyll fluorescence. Yellow-green represents the expressed DMSP lyase enzyme in the transgenic cells (the Alma1 sequence was conjugated to GFP). Bar = 2 µm for a,b and 5 µm for c-e.
I found it fascinating that overexpressing just one gene in the prey led to a dramatic enhancement in the grazing rate by O. marina, and wondered how widespread this effect is, how will other predators respond to the elevated DMS production in prey cells? Indeed, we found that DMS-mediated herbivory may be ecologically relevant, since different grazers responded in the same way as O. marina, including natural grazers collected from the Red Sea prior to the experiments. However, not all grazers tested were feeding more on prey with enhanced DL activity, thus the response to DMS-producing prey may be species-specific. Overall, our research suggests that in the ocean, DMS-mediated herbivory may have a significant impact on bloom dynamics of DMS-producing species like E. huxleyi. During bloom events, minute plumes of algal DMS may initiate grazing interactions that release more DMS from grazed cells into the water. The DMS acts as an appetizing ‘eat-me’ signal, attracting microzooplankton and leading to enhanced grazing pressure on the phytoplankton population.
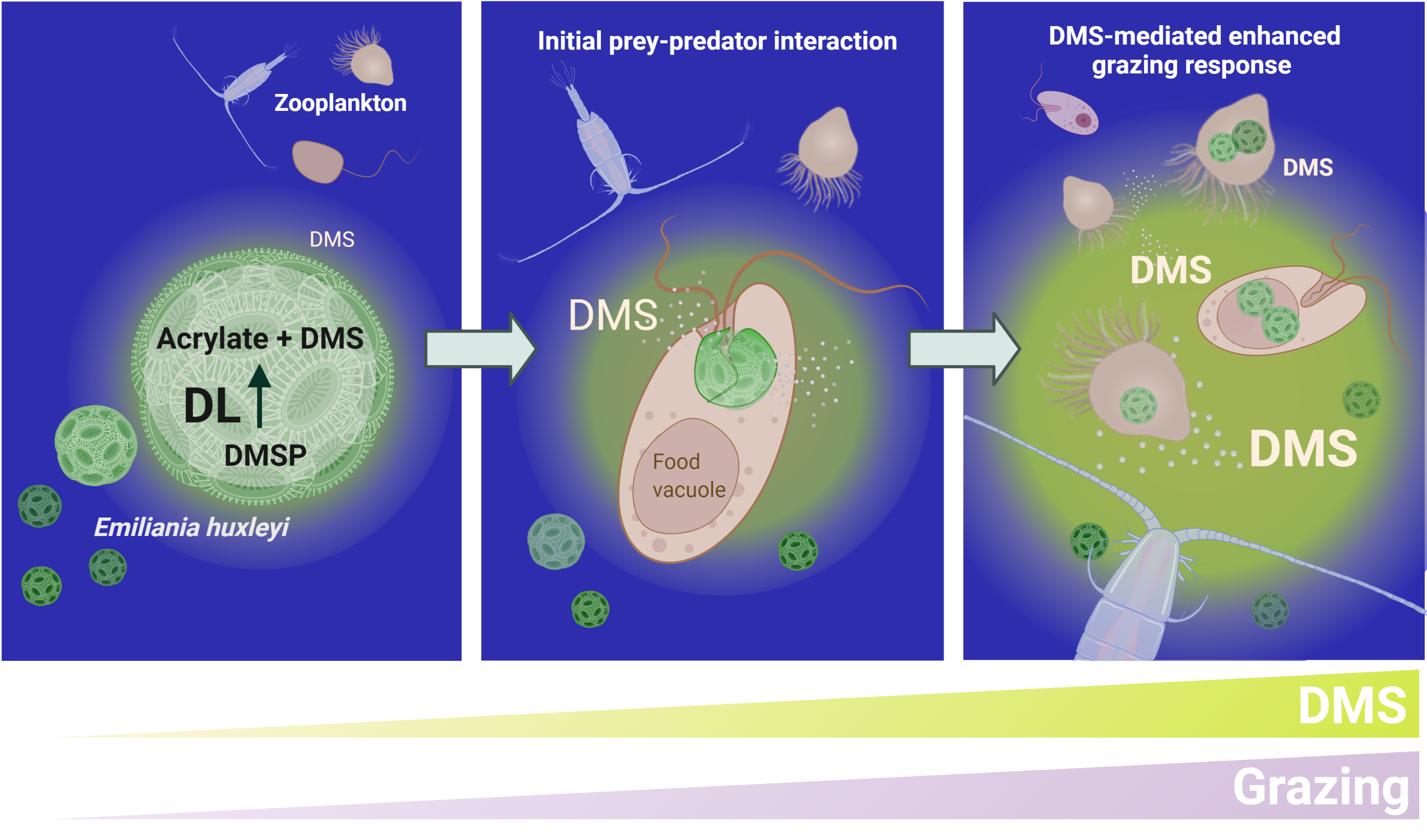
The ecological impact of algal DMS on planktonic prey-predator interactions. A conceptual model describing DMS-mediated herbivory. Chemoattraction of microzooplankton to leakage of DMS from phytoplankton such as E. huxleyi may facilitate initial grazing interaction (left panel). Upon dissociation of ingested cells during phagocytosis, the DL Alma1 degrades DMSP and releases more DMS into the water (middle panel). The microscale release of DMS may accumulate to high concertation, which induces an increase in the grazing pressure by diverse grazers on the entire phytoplankton population (right panel), and thus has a wide ecological implication in the marine environment.
The suggested pro-grazing role for the DL enzyme raises evolutionary questions regarding the retention of the alma1 gene in the genomes of ecologically successful phytoplankton as E. huxleyi, Phaeocystis and Symbiodinium1. Why do phytoplankton release DMS, an infochemical that enhances their removal by predators? What is the cellular role of the DL enzyme in phytoplankton? Can the DMS signal in the water induce trophic cascades, attracting primary consumers (microzooplankton) and secondary consumers (copepods) at the same time? These open questions remained to be answered by the marine chemical ecology community in the near future.
Read the full story in our new Nature Microbiology paper.
References
1. Alcolombri, U. et al. Identification of the algal dimethyl sulfide-releasing enzyme: A missing link in the marine sulfur cycle. Science 348, 1466-1469 (2015).
2. Haas, P. The liberation of methyl sulphide by seaweed. Biochem. J. 29, 1297-1299 (1935).
3. Simó, R. Production of atmospheric sulfur by oceanic plankton: biogeochemical, ecological and evolutionary links. Trends. Ecol. Evol. 16, 287-294 (2001).
4. Bouchard, B. et al. Behavioural responses of humpback whales to food-related chemical stimuli. PLOS ONE 14 (2019).
5. Wright, K. L. B., Pichegru, L. & Ryan, P. G. Penguins are attracted to dimethyl sulphide at sea. J. Exp. Biol. 214 (2011).
6. Savoca, M. S. & Nevitt, G. A. Evidence that dimethyl sulfide facilitates a tritrophic mutualism between marine primary producers and top predators. Proc. Natl. Acad. Sci. U.S.A. 111, 4157-4161 (2014).
7. Owen, K. et al. Natural dimethyl sulfide gradients would lead marine predators to higher prey biomass. Commun. Biol. 4 (2021).
8. Seymour, J., Simó, R., Ahmed, T. & Stocker, R. Chemoattraction to dimethylsulfoniopropionate throughout the marine microbial food web. Science 329, 342-345 (2010).
9. Wolfe, G. V., Steinke, M. & Kirst, G. O. Grazing-activated chemical defence in a unicellular marine alga. Nature 387, 894-897 (1997).
10. Strom, S. et al. Chemical defense in the microplankton I: Feeding and growth rates of heterotrophic protists on the DMS-producing phytoplankter Emiliania huxleyi. Limnol. Oceanogr. 48, 217-229 (2003).
11. Breckels, M. N., et al. The role of dissolved infochemicals in mediating predator–prey interactions in the heterotrophic dinoflagellate Oxyrrhis marina. J. Plankton. Res. 33 (2011).
12 . Alcolombri, et al. Assigning the algal source of dimethylsulfide using a selective lyase inhibitor. ACS Chem. Biol. 12, 41-46 (2017).
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in