Establishing semantic interoperability across cohorts at European level
Published in Healthcare & Nursing
Our work aimed at enhancing interoperability within the international research community in the field of COVID-19. The ORCHESTRA project gave us the opportunity to continue the ‘FAIRification’ process of COVID-19 –related data by use of international IT standards started with the definition of the German Corona Consensus Data Set (GECCO).
ORCHESTRA is a multinational initiative funded by the European Commission to advance the knowledge of the SARS-CoV2 infection and its long-term effects. Within the project, it was decided to use the REDCap® web application for patient data collection of the prospective clinical studies that were being developed. Based on our experience with GECCO, we used international interoperability standards to define the study data elements of the prospective and retrospective ORCHESTRA clinical studies. The purpose of this effort was to enable interoperable processing of the collected data across countries and across cohorts. International standard terminologies were used to unambiguously identify the study variables. A data variable is composed of a question or statement and an answer that can allow for a textual reply or would contain a list of permissible answer choices.
To facilitate data harmonization within the project, we first considered the option of working with the clinical partners across several work packages to define a common data set comprised of the most relevant variables that ought to be collected by all ORCHESTRA cohorts. Soon enough we realized that this was not feasible for several reasons. There was significant time pressure to get the planned studies to launch quickly in order to gather valuable data during the case number peaks of the waves. In addition, the cohort structures and legal as well as ethics board requirements were so diverse that our first option did not seem practical and efficient. That is because ORCHESTRA includes many partners and clinical teams; some of them had ongoing COVID-19 trials and others were about to launch new clinical studies. Furthermore, we also faced some skepticism to the data standardization and harmonization process because its inherent value was not apparent to all partners, fearing that it might create extra work for them, some took longer to reach out to us and share their data definitions. The sum of these challenges led us to take a different approach: we explored the metadata definitions that already existed and compared them to the new ones that were being defined for use in prospective studies’ case report forms (CRFs).
We started this standardization process off by focusing on the first new prospective ORCHESTRA study, referred to as the “Long-term Sequelae” (LTS) study, for which we received the CRF from the clinical partners. This turned out to be a good starting point for our activities as the study was quite comprehensive and diverse in terms of categories of variables. Not only did it contain baseline information, but the team also intended to conduct several exams and gather laboratory parameters at follow up visits. Additionally, it included all the genomics variables that a dedicated ORCHESTRA team was suggesting.
While we were mapping the LTS study variables to standard terminologies, we also began to work on standardizing study variables for a second clinical study. The “Fragile Population” (FP) study had at that point already started collecting patient data, but they intended to add new variables to include long-time monitoring aspects. This second study was also large and comprehensive in terms of information categories covered. Working on the two studies in parallel led to frequent comparisons between the studies’ variables, many of which were similar in their phrasing and intended meaning. Whenever we noticed two similar variables and deemed it to be appropriate, we suggested adjustments so that both clinical groups could adjust the variables to be identical and therefore facilitate analysis of data across studies. Hence, all identical variables were mapped to the same standard code which was incorporated in the variable identifier (ID). The most comprehensive terminology for clinical information, SNOMED CT, was the main source of codes assigned to variables in ORCHESTRA studies. SNOMED CT allows for the license-free use of a subset of its terminology codes which are contained in the Global Patient Set (GPS). In response to the pandemic, the GPS was significantly enriched with many COVID-19-related concepts to support interoperable research in the field. However, about a third of all SNOMED codes used to represent variables in ORCHESTRA were not part of the GPS. Thus, we applied for and obtained a fee waiver for the usage license.
When it came to variables describing laboratory examinations, many of the assays used were new and COVID-19-specific so that they had not been included in the standard terminologies yet. This was true also for some genomics related tests involving specific characteristics of the virus. To address this issue, we prepared numerous submissions of genomics, serology, questionnaire and other terms to the standard developing organizations (SDOs). With regards to our submissions to the standards developing organizations LOINC (for laboratory parameters, questionnaires and documents) and NCIt (for omics-related variables), our contribution was recognized by being invited to propose improvements to the submission process (for LOINC) and being listed as COVID-19 terminology contributors (with NCIt). In the case of submitting copyrighted questionnaires to SDOs for coding, we reached out to the respective authors to obtain permission before we could submit their works for coding. Fortunately, permission was granted in most cases.
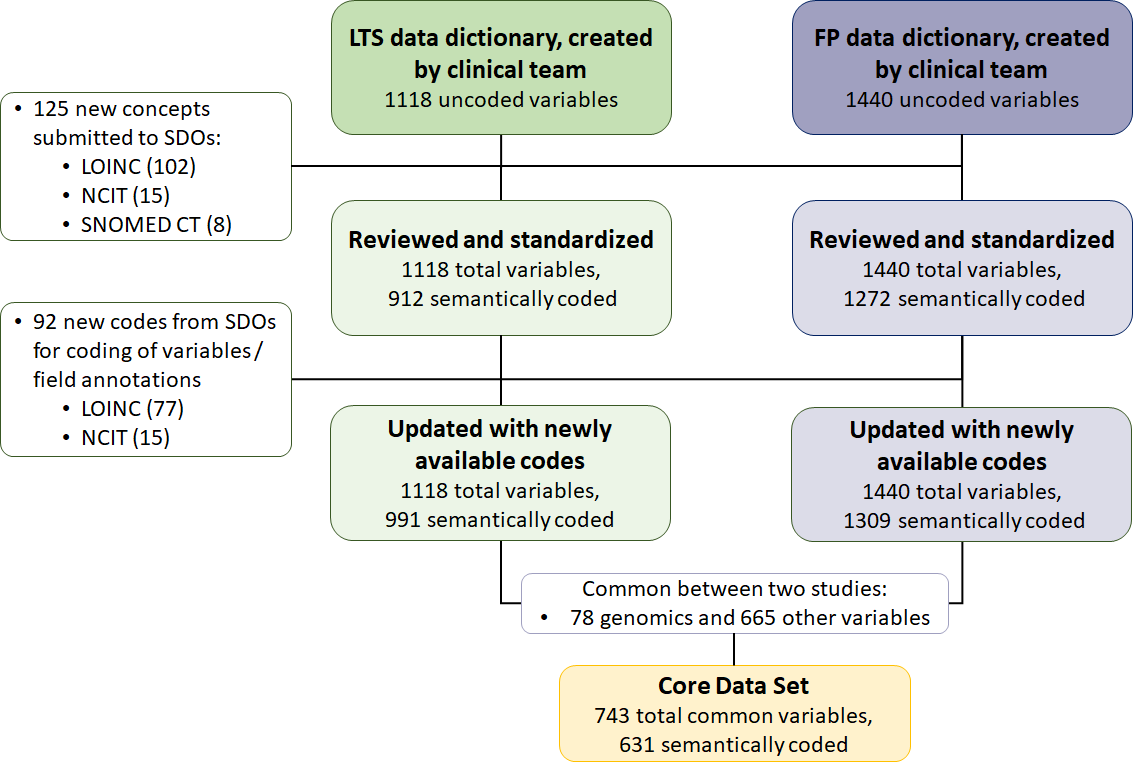
We used international standard terminologies and classification codes to represent an extremely wide range of clinical concepts and embedded them into the metadata definition of case report forms (CRFs). When a variable was used in at least two ORCHESTRA studies, it was identified as high priority and converged into the “ORCHESTRA Core Data elements for COVID-19" (OcDeC) pool of variables (data elements). By doing so, we created a Core Data Set of standardized variables (Figure 4) for future use with the goal to simplify the merging of data and increase metadata quality.
 |
|
Figure 4: Overview of harmonized data and submissions to standard developing organizations. |
Our pool of standardized variables (OcDec) can be used to develop new clinical studies within ORCHESTRA but also beyond the project’s borders by other research initiatives. OcDeC has in fact already been shared with two other European Projects working on COVID-19 who were interested in re-using a subset of the variables. In order to support the extended use of the variables, the variable definitions have been made freely available on the ART-DECOR® open-source platform that supports the creation and maintenance of Health Level Seven International (HL7) data sets. OcDeC is continuously being updated as our work in ORCHESTRA progresses.
Following the methodology of the American National Institute of Health, the OcDeC represents a list of common data elements (CDEs) for COVID-19. CDEs are standardized, precisely defined questions paired with a set of specific allowable responses and used systematically across different sites or clinical trials to ensure consistent data collection.
Using standards right from the inception of a study is of great advantage as it allows to map variables immediately to corresponding international terminology standard codes and where possible, also to match them with already coded elements from other studies.
Follow the Topic
-
npj Digital Medicine
An online open-access journal dedicated to publishing research in all aspects of digital medicine, including the clinical application and implementation of digital and mobile technologies, virtual healthcare, and novel applications of artificial intelligence and informatics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Evaluating the Real-World Clinical Performance of AI
Publishing Model: Open Access
Deadline: Jun 03, 2026
Impact of Agentic AI on Care Delivery
Publishing Model: Open Access
Deadline: Jul 12, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in