From the BugBitten Archives: Clocking out -knocking out circadian clock gene disrupts key functions in Aedes mosquitoes
Published in Microbiology and Zoology & Veterinary Science

This blog was originally posted on the old BugBitten WordPress site . We are resharing it now on the Research Communities site.
Aedes aegypti is a species of mosquito that is a disease vector for several arboviral diseases such as dengue, Zika and chikungunya that affect millions of people globally. Many of these diseases do not have vaccines readily available, and so a lot of emphasis is put on controlling the spread of the vector. Molecular approaches to vector control have been emerging over the past ten years, with genetically modified mosquitoes being released into the wild to control breeding in mosquito populations.
Vinaya Shetty and colleagues have been looking at another way to disrupt mosquito populations, by investigating whether removing key genes related to the circadian clock of the mosquitoes could impact their clock-dependent biological processes.

Mosquitoes display time-related behaviours such as feeding at certain times of the day. For A. aegypti feeding is a day time activity – mainly in the early morning and late afternoon – whilst the anopheline mosquito is a nocturnal feeder.
In their 2022 study, Vinaya Shetty and colleagues knocked out the core clock cycle gene in A. aegypti using CRISPR/Cas9 and found the mRNA expression of 7 circadian related genes were changed. This knock out affected the mosquito’s longevity, feeding pattern and reproductive fitness. Their latest study explored this further by using transcriptome profiling and differential gene expression to determine the network of genes and biological pathways that were affected by disrupting the clock gene.
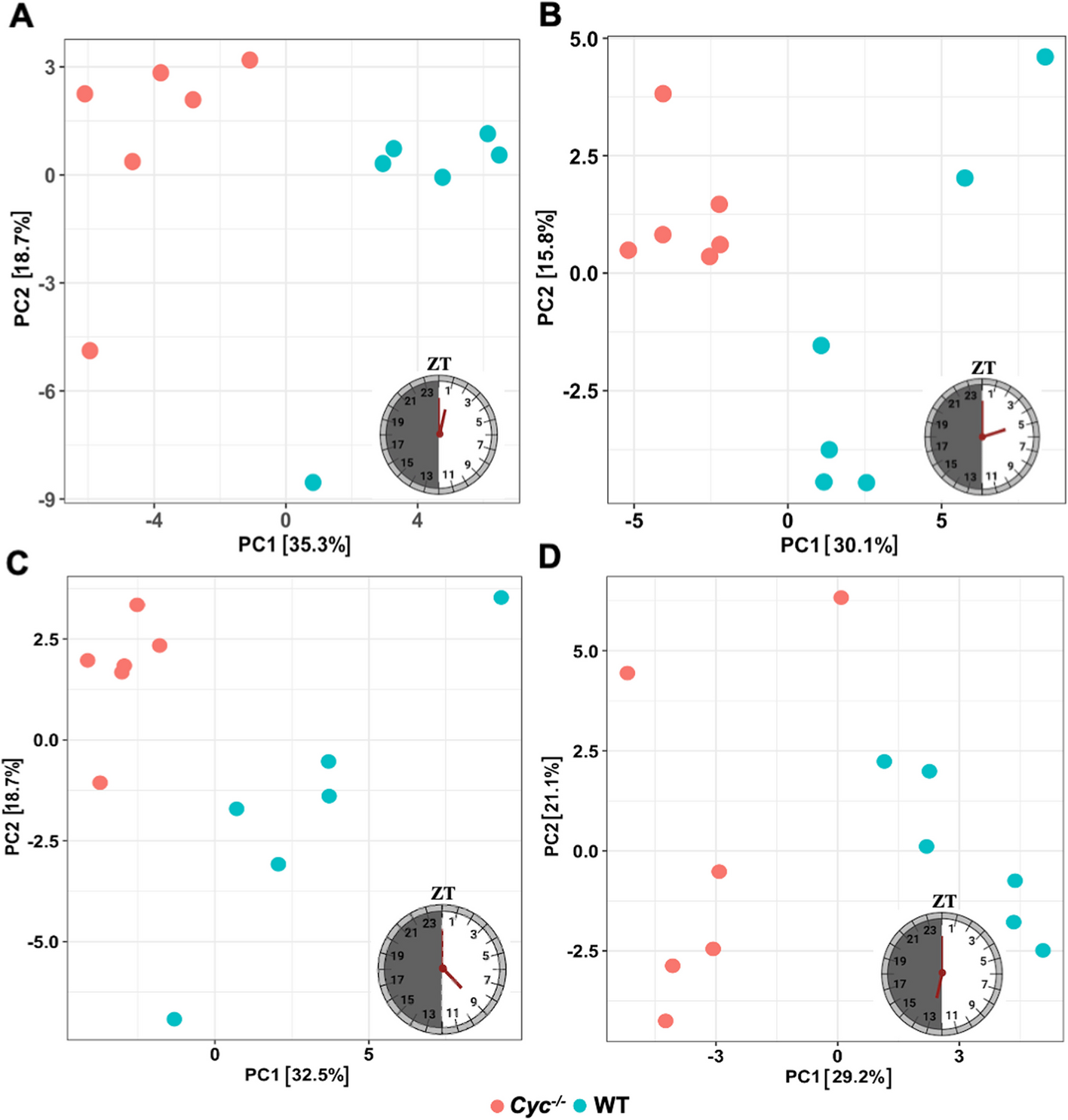
Differential gene expression enabled Shetty and colleagues to identify thousands of genes that were expressed differently in day-night cycles, and determine that removal of the cycle gene caused disruption in metabolic processes, signaling pathways and immune responses. This showed the vital role that the circadian clock plays in A. aegypti’s life, and targeting it could see massive changes to the mosquito’s behaviour and consequently neutralise its impact as a disease vector.
Cover image credit: Gerd Altmann from Pixabay
Follow the Topic
-
BMC Genomics
This is an open access, peer-reviewed journal that considers articles on all aspects of genetics, genomics and proteomics.
-
BugBitten
A blog for the parasitology and vector biology community.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics of human pathogens
For many years now, the study of pathogens’ genomes has enabled more accurate and timely diagnostics while allowing the tracking of specific strains or isolates. This can help discover the source of a specific epidemic and how it spreads, which in turn informs the best way to contain it. However, recent advances in third-generation sequencing techniques and the bioinformatics tools linked with them have transformed practice. It is now possible to sequence individual genomes much faster and much cheaper, which is opening new avenues for pathogen tracking, even in areas with limited funding such as the global South.
Advances in genomic technologies will also facilitate the emergence of personalized medicine approaches tailored to specific infections—such as genome-guided antiretroviral therapy for HIV or drug-resistance profiling in tuberculosis. Real-time genomic surveillance, exemplified by platforms like GISAID or other repositories during the COVID-19 pandemic, enables rapid detection and monitoring of emerging variants, thus improving public health responses.
This Collection invites submissions looking at human pathogens genomes and data sets to build on the knowledge already accumulated. We will also consider rapid diagnostics datasets and new techniques in genomic studies of human pathogens.
If you have a data note to submit, please submit to our sister collection Genomics of human pathogens data notes in BMC Genomic Data.
Topics of interest include, but are not limited to:
- Genomic characterization of bacterial pathogens
- New genomes of pathogens or strains
- Viral genomics and infection dynamics
- Comparative genomics of human parasites
- Microbial resistance mechanisms in pathogens
- Genomic epidemiology of infectious diseases
- Host-pathogen genomic interactions
- Metagenomic approaches to pathogen discovery
- Evolutionary genomics of emerging infections
- Functional genomics and regulatory networks in pathogens
This Collection supports and amplifies research related to SDG 3, Good Health & Well-Being.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Jul 20, 2026
Cattle genomics
BMC Genomics invites researchers to contribute to our Collection on Cattle genomics focusing on understanding the genetic makeup of bovine species, which is essential for improving livestock breeding and health. Advances in genomic technologies, such as next-generation sequencing and RNA sequencing, have enabled researchers to reveal insights into traits such as growth, meat quality, milk production, disease resistance, reproductive fitness, and overall adaptability in bovine genomes. This Collection aims to highlight the latest research developments in cattle genomics, encompassing both genomic and transcriptomic studies that contribute to the understanding of bovine biology.
Recent breakthroughs in genomic selection and precision breeding techniques have already shown promise in increasing efficiency in cattle production. The use of CRISPR-Cas genome editing, for example, has allowed for precise modifications to the cattle genome, introducing beneficial genetic variations without the linkage drag associated with traditional breeding methods. Additionally, the integration of omics technologies is paving the way for a holistic understanding of cattle biology, allowing for more effective management and breeding strategies. Studying the rumen microbiome using genomics, transcriptomics, proteomics, and metabolomics has revealed how microbial communities contribute to feed efficiency and nutrient absorption. This comprehensive approach enables targeted nutritional strategies that improve cattle health and productivity while reducing environmental impact. Such integrative studies facilitate the selection of cattle with optimal microbiome compositions, leading to more sustainable and efficient cattle production systems.
As research in cattle genomics progresses, we can anticipate the development of more sophisticated genomic tools that will enable precise manipulation of genetic traits in bovine populations. This may lead to enhanced resilience against diseases, improved reproductive performance, and better adaptation to changing environmental conditions. Ultimately, continued innovation in this field holds the potential to reform cattle production systems, ensuring sustainable livestock farming for future generations.
- Genomic selection in cattle breeding
- Transcriptomic analysis of bovine traits
- Pathogenicity and disease resistance genomics
- Advances in RNA-Seq applications for cattle
- Omics approaches to cattle health and productivity
- Genetic mapping of economically important traits
- Gene editing
- Metagenomics of the bovine gut microbiome
- Epigenetic regulation of growth and reproduction
- Comparative genomics of cattle and other livestock species
This Collection supports and amplifies research related to SDG 2, Zero Hunger.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: May 26, 2026







Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in