Generative deep learning for the development of a type 1 diabetes simulator
Published in Social Sciences, Computational Sciences, and Immunology

Transforming Diabetes Simulation with Deep Generative Models
Biomedical simulators play a crucial role in healthcare by providing a cost-effective alternative to traditional patient and animal testing methods. These simulators, often based on mathematical physiological or pharmacokinetic models, are instrumental in developing and testing new treatments and therapeutic strategies. However, despite their utility, physiological models have inherent limitations in capturing the complexity of biological processes, particularly in conditions like Type 1 Diabetes (T1D). Factors such as unmeasured variables and external influences like lifestyle choices pose challenges for accurate modeling, leading to inherent errors that compromise the effectiveness of physiological models. T1D simulators, traditionally based on physiological models, suffer from the same fate.
Bridging the Gap: Introducing Deep Generative Models
Deep Generative Models (DGMs) have emerged as a promising alternative, offering the capability to learn complex distributions from data and generate novel samples with remarkable fidelity. Unlike discriminative models that focus on learning conditional boundaries, DGMs excel in generating realistic samples by capturing the joint probability distribution of data. DGMs, particularly Generative Adversarial Networks (GANs), have demonstrated remarkable efficacy in capturing the underlying distribution of complex biological phenomena, including the glucose-insulin dynamics of T1D patients.
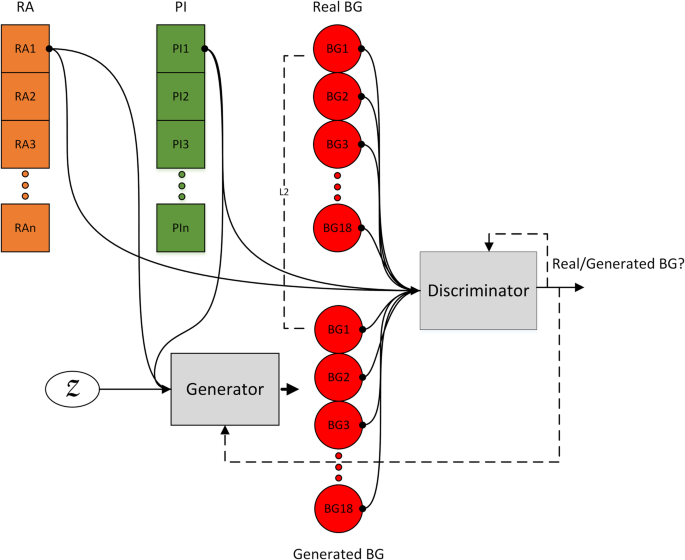
Unlike traditional simulators, which rely on predefined mathematical models, the proposed approach learns glycaemic trends directly from data, conditioning the generation process on key variables such as insulin and carbohydrate levels. By employing a sequence-to-sequence (S2S) GAN architecture, the model translates input insulin and carbohydrate data into corresponding blood glucose (BG) profiles, thereby capturing the relationships inherent to the T1D physiology. Moreover, the model learns causality by introducing shifted input and output pairs during the training process. Each pair of insulin and carbohydrate inputs is mapped to blood glucose values 90 minutes into the future. This enables the model to capture the effect of insulin and carbohydrates being translated into the blood glucose profile.
Assembling the Simulation Environment
The trained DGMs were then integrated into a simulation environment with a state-of-the-art controller. To ascertain the fidelity of the generated BG profiles, the study subjected them to validation under both open-loop (OL) and closed-loop (CL) therapy settings. The results revealed similarities between the simulated and real patient outcomes, affirming the model's capacity to emulate diverse clinical scenarios.
Of particular significance is the observation that the model's responses to CL therapy closely mirrored those observed in actual patients. This is a notable departure from conventional simulators, which often exhibit overly optimistic outcomes under CL therapy. The model's ability to replicate real-world complexities positions it as a potent tool for validating treatment strategies and exploring novel therapeutic avenues.
Shaping the Future of Diabetes Management
Looking ahead, the proposed methodology holds promise for shaping the future of T1D management. By leveraging latent space exploration and employing state-of-the-art DGM architectures, researchers can tailor synthetic patients with desired characteristics, paving the way for the creation of personalized models.
Moreover, the integration of advanced control strategies and variables like physical activity and time as conditional inputs opens avenues for refining the model's accuracy and versatility. This convergence of AI and healthcare promises a new era of precision medicine, where patient-specific simulations drive innovation and enhance clinical decision-making.
Conclusion
In conclusion, the convergence of DGMs and diabetes simulation suggests a new frontier in disease modeling and therapeutic development. By realistically replicating real-world dynamics and enabling personalized treatment simulations, these models empower clinicians, people living with diabetes, and researchers to navigate the complexities of T1D management with improved insight and precision. With continued advancements in DGM technologies and their integration into healthcare platforms, we believe that the landscape of diabetes management can be transformed, ushering in an era of personalized, data-driven care.
Follow the Topic
-
Communications Medicine
A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary across all clinical, translational, and public health research fields.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Public health and health governance in China
Publishing Model: Open Access
Deadline: Jul 31, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in