Go with the flow: high throughput 3D localization in cells by depth-encoding imaging flow cytometry
Published in Bioengineering & Biotechnology
In localization microscopy, the distribution of signal photons in each image is analysed to estimate the underlying positions of emitters. Point-spread-function engineering refers to modifying the way these photons are patterned in the image to create distinctive shapes that encode additional information. For 3D imaging, i.e. depth encoding, we normally calibrate the shape-to-depth response of the microscope by scanning a stationary object at various levels of defocus to build an image dictionary. In the fast-moving microfluidic channel of an imaging flow cytometer, however, scanning is not feasible. Further complicating the problem is that the positions of objects within the flow are distributed stochastically.
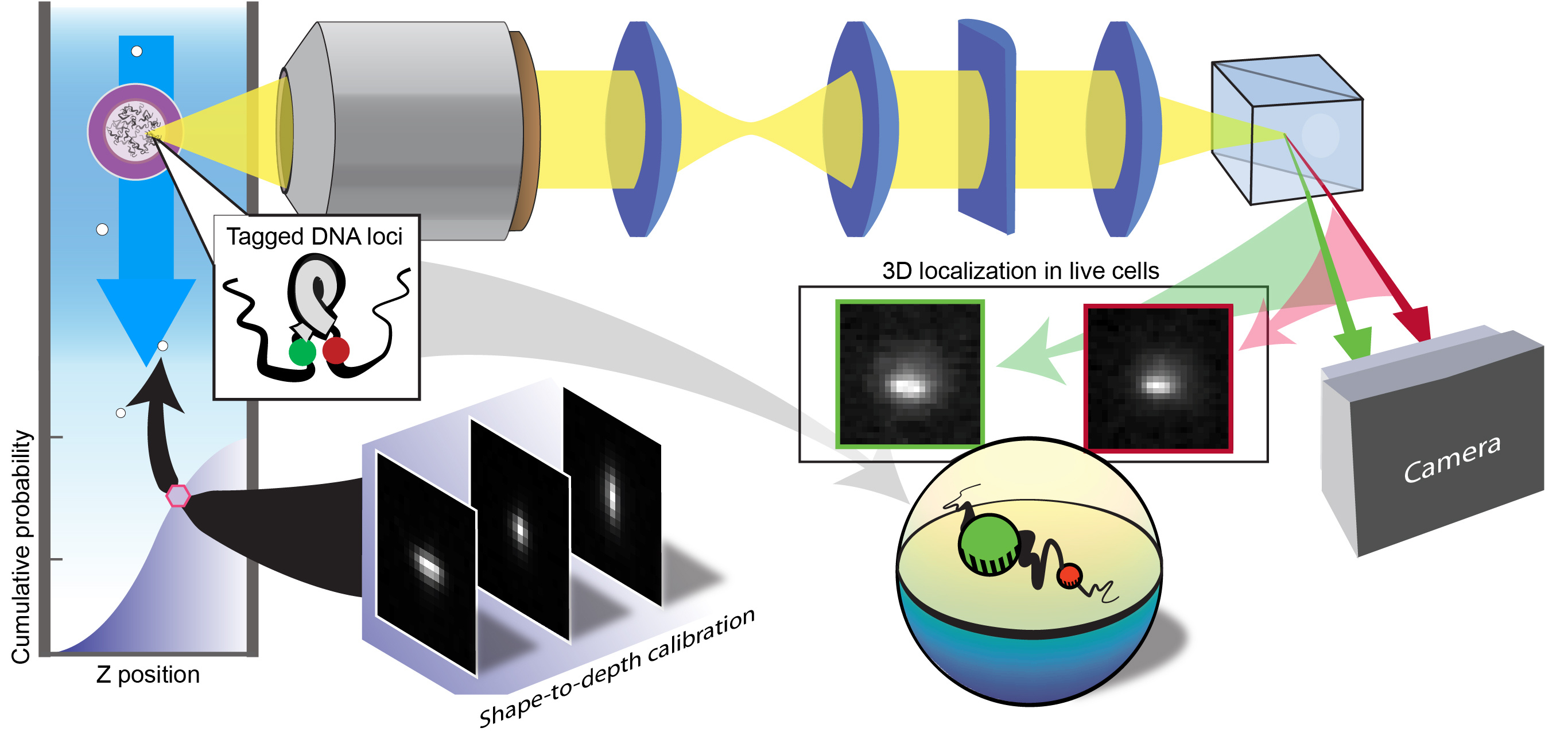
A key breakthrough in our approach came when we recognized that we could readily detect the signal from objects even at the very edge of these stochastic distributions using bright, nanoscale fluorescent objects, i.e. tiny beads loaded with fluorophores. In other words, we could sample the entire distribution so long as we recorded enough measurements. Furthermore, while the z positions of individual particles were unknown, the overall order in any set of measurements, in terms of depth, could be deduced by comparing the relative shapes in the images. Combining these two pieces of information, namely (1) that we can sample the full distribution and (2) a set of measurements can be ordered by depth, the calibration falls into place! Most importantly, this calibration can then be applied to new samples irrespective of how they are spatially distributed such as fluorescent objects within flowing cells.
This method enables certain experiments that were challenging to perform previously, particularly tracking the dynamic changes in cell populations with robust statistics. The throughput is essential here due to the inherent heterogeneity in these samples that often have subtle, yet important effects which can only be quantified from large populations. Harnessing the throughput of our approach to characterize one such system, we performed 3D measurements of DNA compaction in live yeast cells - recording over 1000 cells each minute! Interestingly, we found that when our cells were loaded at higher concentrations, they tended to aggregate. In other words, we were no longer limited by the microscopy hardware, but had reached a biological detection limit for single-cell imaging!
To further demonstrate the capabilities of high-throughput 3D localization, we quantified the uptake of nanoparticles in cancerous white blood cells. In the context of nanomedicine, the combination of throughput and precision enabled here can be applied to study how various formulations of medicinal nanoparticles could influence cellular uptake, intracellular distributions, therapeutic effects, and more - all of which can help in the design of targeted nanomedicines.
Follow the Topic
-
Nature Nanotechnology
An interdisciplinary journal that publishes papers of the highest quality and significance in all areas of nanoscience and nanotechnology.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in