HALD, a human aging and longevity knowledge graph for precision gerontology and geroscience analyses
Published in Research Data
The published literature is one of the most accessible data sources of molecular and disease information related to aging and longevity. However, due to the huge amount of biomedical literature, it is time-consuming and inefficient for researchers to conduct information retrieval from the major databases of medical journals like PubMed. Integrated datasets with comprehensive knowledge are crucial for researchers to leverage existing resources. In the field of life sciences, a biomedical KG can not only link biomedical entities through certain relations, but also predict the potential relationships between existing entities and discover new relational facts.
In this paper, we presented HALD, a human aging and longevity dataset of the biomedical KG from human aging and longevity-related literature in PubMed. Figure 1 illustrates the workflow of biomedical literature mining using multiple NLP techniques.
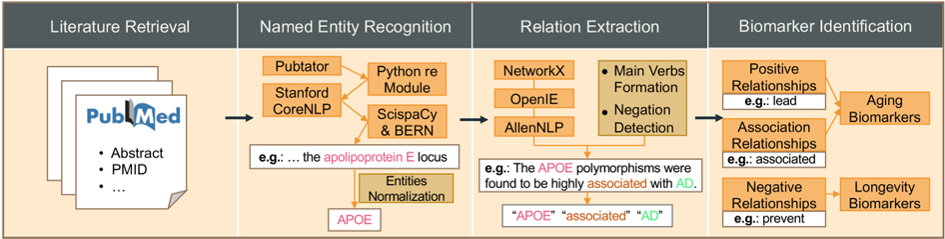
Up to September 2023, we had annotated 339,918 abstracts from PubMed and curated 12,227 entities in 10 types (gene, RNA, carbohydrate, peptide, lipid, protein, pharmaceutical preparations, toxin, mutation, and disease entities), 115,522 relations, 1,855 aging biomarkers, and 525 longevity biomarkers in HALD. The distributions of entities and relations are shown in Fig. 2a,b.
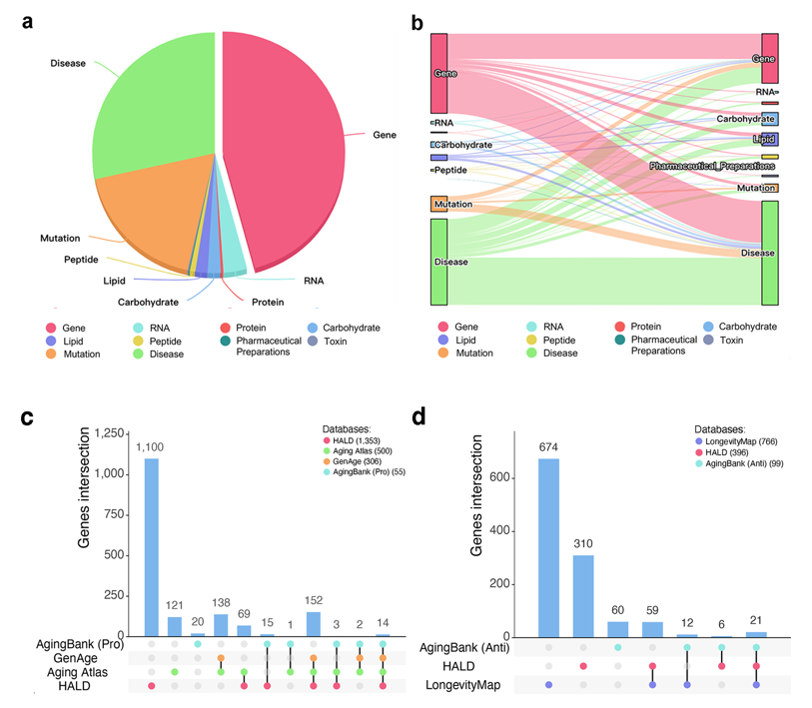
The contributions of HALD are listed as followings:
-
HALD is the first human aging and longevity knowledge dataset of the biomedical knowledge graph mined from published literature using NLP technologies.
-
HALD provides 10 types of credible human aging and longevity biomedical entities.
-
HALD links biomedical entities through certain relations and predicts the potential relationships.
-
HALD identifies aging and longevity biomarkers from curated entities and elucidates their associations with aging-related diseases.
Literature retrieval
A search for (“aging” [Title/Abstract] OR “ageing” [Title/Abstract] OR “longevity” [Title/Abstract] OR “centenarian” [Title/Abstract] OR “the elderly” [Title/Abstract] OR “the aged” [Title/Abstract] OR “old people” [Title/Abstract] OR “older people” [Title/Abstract] OR “old age” [Title/Abstract] OR “gerontology” [Title/Abstract] OR “geroscience” [Title/Abstract] OR “lifespan” [Title/Abstract] OR “healthspan” [Title/Abstract] OR “life expectancy” [Title/Abstract] AND “Journal Article” [ptyp] AND “humans” [MeSH Terms] AND “English” [lang]) was used to retrieve PubMed biomedical journal articles related to human aging and longevity directly with the Bio.Entrez python package.
Named entity recognition
We combined web-based, dictionary-based, rule-based, and DL-based methods to conduct NER, and recognized 10 types of entities including gene, RNA, protein, carbohydrate, lipid, peptide, pharmaceutical preparations, toxin, mutation, and disease.
Relation extraction
Once two entities co-exist in one sentence, and a main verb lies between the 2 entities at the meanwhile, there is likely to be some relationship between these two entities. We selected sentences with no less than two entities to conduct RE through the NetworkX, OpenIE, and AllenNLP methods.
Human aging and longevity biomarkers identification
We further identified human aging and longevity biomarkers by investigating the characteristics of the relationships between gene, RNA, protein, carbohydrate, lipid, peptide, pharmaceutical preparations, toxin, mutation entities and disease entities. The relationships between the potential human aging and longevity biomarkers and disease entities were divided into three classes as follows:
-
Positive relationship. Positive relationships like “lead” and “cause” were considered aging-promoting relationships.
-
Association relationship. Relationships that can indicate an association like “associated” and “related” were considered aging-promoting relationships.
-
Negative relationship. Negative relationships like “prevent” and “ameliorate” were considered longevity-promoting relationships.
Framework of KG
Generally, resource description framework (RDF) and graph database are two main storage forms of KG. RDF is convenient for designers to publish and share data, while graph dababase provides a user-friendly interface to browse data. Thus, we developed the graph database-based HALD to explore the human aging and longevity-related KG. The front end was built with React (https://react.dev/) and Elasticsearch (https://www.elastic.co/) was used to realize a real-time search and management. We employed Neo4j (https://neo4j.com/) to offer an intuitive network demonstration of the entities and relations knowledge. All analyses in this study were done inside JupyterLab (https://jupyter.org/) notebooks with the Python kernel. Automatic updates would be executed monthly to keep the KG up-to-date.
The presentation of the HALD dataset aimed to provide significant convenience to researchers in the field of human aging and longevity and reduce the workload of further sifting through vast amounts of data. Additionally, HALD predicted biomarkers of aging and longevity from published literature, making it a valuable reference for precision gerontology and geroscience analyses. HALD is publicly available at Figshare, an open scientific data repository. For scientific researchers who want to explore the dataset intuitively, please visit https://bis.zju.edu.cn/hald for user-interactive browsing.
Users are welcome to contribute data and give suggestions in the Feedback module on the website at any time, by directly filling the form and click the “FEEDBACK” button to submit it. We will promptly check all the feedback, respond via email, and make necessary adjustments as soon as possible.
Code availability
All code used in this paper can be downloaded on GitHub at https://github.com/zexuwu/hald.
Follow the Topic
-
Scientific Data
A peer-reviewed, open-access journal for descriptions of datasets, and research that advances the sharing and reuse of scientific data.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genomics in freshwater and marine science
Publishing Model: Open Access
Deadline: Jul 23, 2026
Genomes of endangered species
Publishing Model: Open Access
Deadline: Jul 01, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in