Hi Neighbor - Have some of my glycerol
Published in Microbiology

Explore the Research

Glycerol metabolism supports oral commensal interactions - The ISME Journal
The ISME Journal - Glycerol metabolism supports oral commensal interactions
"Biofilm, city of microbes" (1) was one of the defining reviews for me as a new post-doc, quite a while ago. It made me realize that my path forward as a microbiologist has to be in the context of microbial ecology. Microbes do not dwell alone, quite the contrary, they thrive when others are around, like to chat, share, work together and sometimes fight with neighboring species co-inhabiting the biofilm. My group is interested in these interspecies interactions that are ultimately responsible for species distribution in the developing biofilm and determine overall biofilm architecture. The species distribution inside the biofilm is also intimately linked to the ability of a biofilm to cause disease or as even more important, but vastly understudied, support health. Our area of research is the oral biofilm.
We previously reported that two abundant and oral health associated species, Streptococcus sanguinis and Corynebacterium durum have a close relationship through the production of membrane vesicle and fatty acids leading to S. sanguinis chain elongation and overall increased fitness supporting their commensal state (2) and Fig 1.
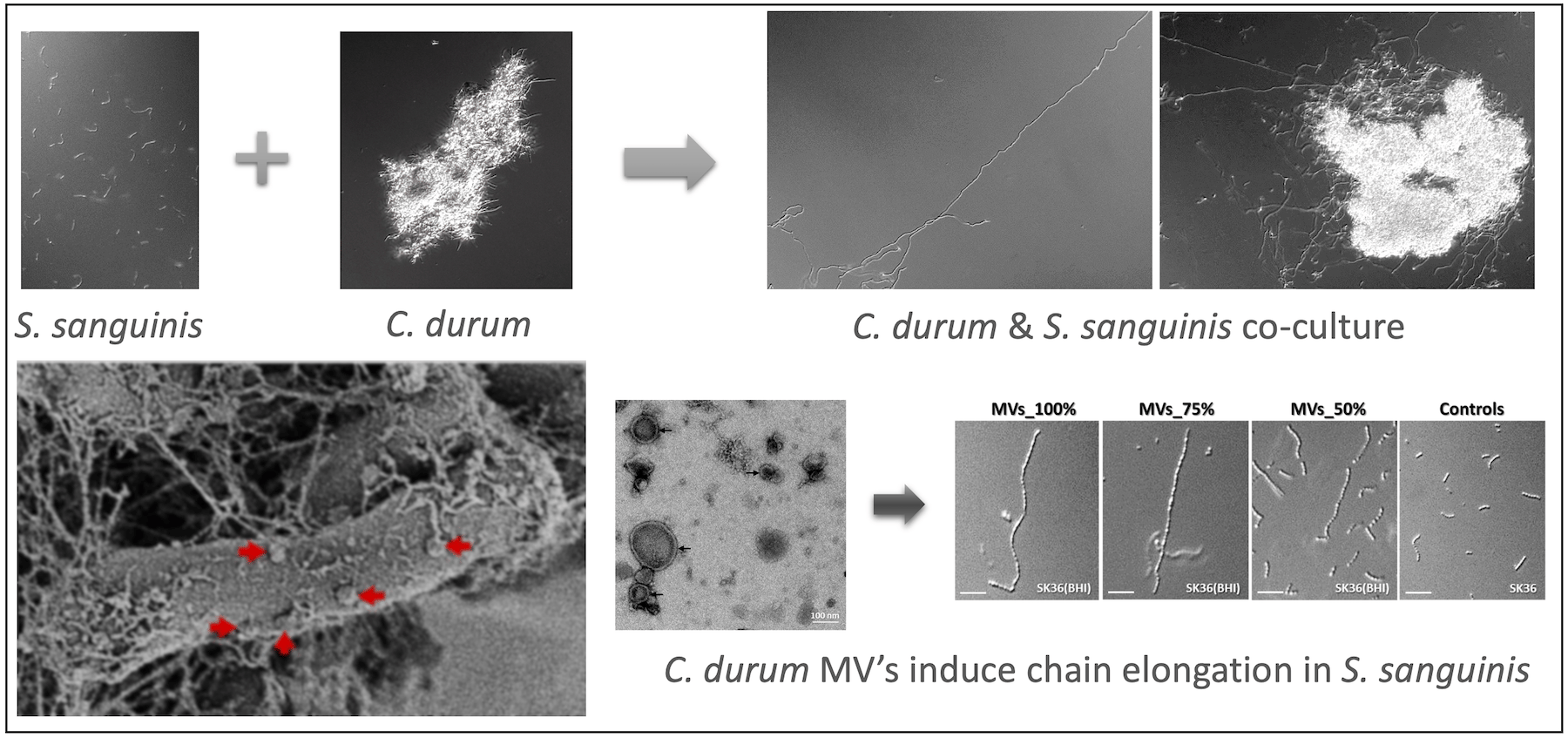

To further understand the relationship and define the molecular mechanisms that lead to the observed chain elongation phenotype, we performed global transcriptome analysis of the co-cultures to identify differentially expressed genes in S. sanguinis. Two operons caught our interest, the glp-operon encoding important genes for glycerol metabolism and the pil-operon responsible for type IV pilus formation on the surface of S. sanguinis.
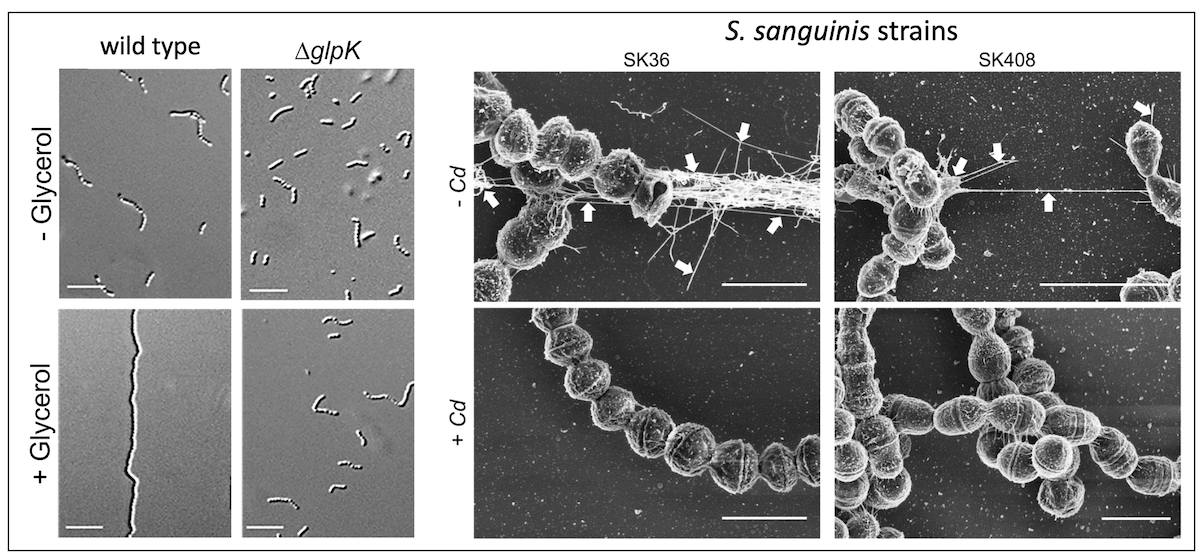
Bacterial type IV pili are known to be essential for various functions influencing microbial ecology, including adhesion, bacterial-bacterial and bacterial-host interactions. Surprisingly, in S. sanguinis (at least in some strains) type IV pili are also responsible for twitching/gliding motility. Thus, a C. durum dependent repression of pilus gene expression seems to favor the commensal state of S. sanguinis since it would abolish its ability to move away. It might also decrease its ability to adhere to other species thus aiding in the close interaction of both providing the ideal commensal relationship.
Why is research into commensals and "molecular commensalism" important?
Polymicrobial diseases affecting mucosal surfaces do not strictly adhere to Koch’s postulates, as no single invading species can be identified as a causative agent. Consequently, eliminating a single organism from the body is a largely ineffective strategy to resolve such complex diseases these types of diseases. A holistic approach may be useful for identifying mechanisms to support or strengthen molecular commensalism (3) of our microbiomes in order to avert sickness or at least understand these organisms’ roles in supporting health and homeostasis.
References:
(1) Watnick P, Kolter R. Biofilm, city of microbes. J Bacteriol. 2000 May;182(10):2675-9. doi: 10.1128/JB.182.10.2675-2679.2000. PMID: 10781532; PMCID: PMC101960.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in