Lions & sea lions & bears, oh my: the value of natural history collections for studying the developing skeleton
Published in Ecology & Evolution, Zoology & Veterinary Science, and Anatomy & Physiology
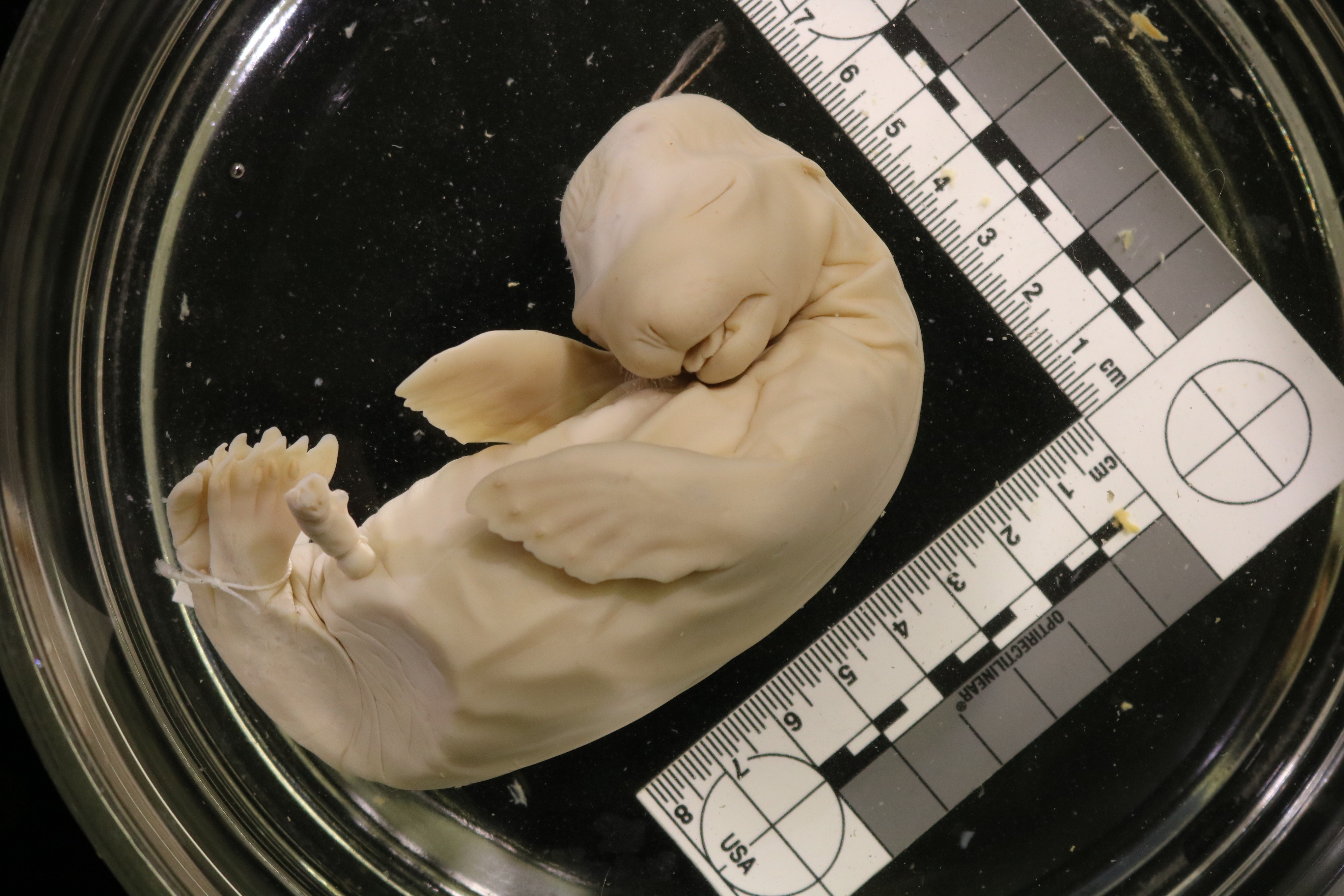
Most mammalian embryos develop for weeks to months inside the mother’s body. This developmental strategy of giving birth to live young –known as viviparity– both protects the developing offspring and allows the mother to continuously supply nutrients over an extended period. Apart from monotremes, or egg-laying mammals, all other mammals give birth to live young, making studies of their early development difficult. Collection of prenatal materials for species other than established models is extremely challenging, both practically and ethically, due to factors such as the body size, habitat, reproductive biology, and conservation status.
Natural history museums around the world offer a simple, ethical means of studying mammalian development because their collections consist of hundreds to thousands of historically (and opportunistically) collected prenatal specimens for a wide range of mammals. By using non-invasive imaging technologies, such as micro-computed tomography (micro-CT), researchers can image the developing skeleton without harming these precious specimens and digitally share these data with the scientific community through public repositories.
This study was inspired when we discovered that the wet collections of the Harvard Museum of Comparative Zoology (MCZ) Mammalogy Department contain hundreds of mammalian embryos preserved in jars of ethanol which had not been previously used in research. Many were collected over 100 years ago.

Harvard MCZ collection.
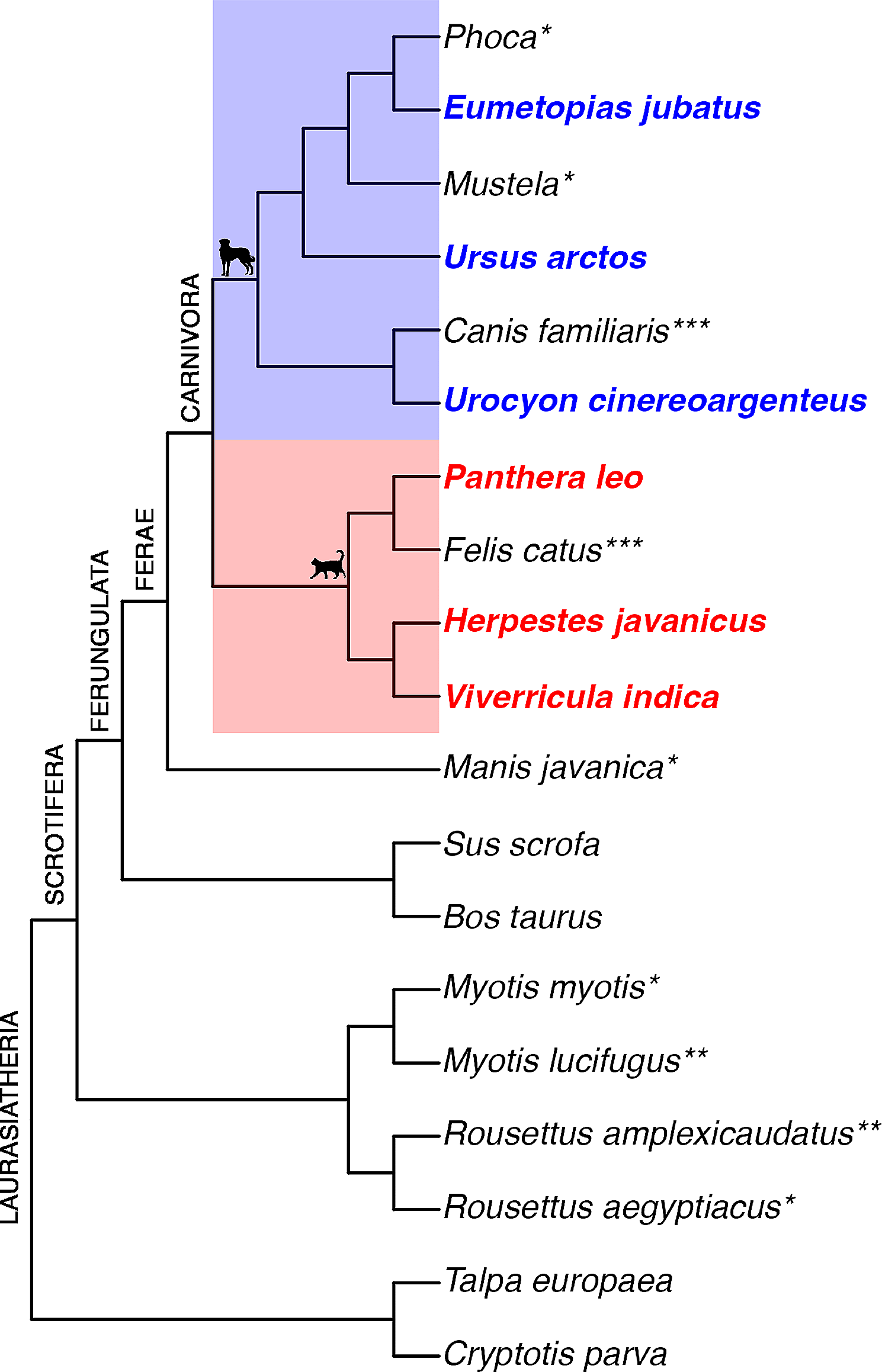
To conduct our research, we used micro-CT to study the developmental sequence of bone formation (ossification sequence) in the mammalian order Carnivora. Species included the Steller sea lion, small Indian mongoose, lion, gray fox, brown bear, and small Indian civet. We chose to focus on carnivorans because previous research on carnivoran ossification sequences was largely limited to two domestic species: dogs and cats. Furthermore, the MCZ collection included singleton embryos from a wide taxonomic range of species, as well as numerous Steller sea lion embryos of different developmental stages. This diversity allowed us to compare both within and between species, as well as among animals living in terrestrial versus aquatic environments.
After CT-scanning, we digitally reconstructed the developing skeleton of each specimen and identified which bones were present. Based on these data, we could determine the order of bone formation by comparing the bones observed with those expected based on the adult skeleton. For example, if the femur was present in one specimen while the tibia was absent, we infer that the femur ossifies before the tibia in this species. Using this logic, we constructed ossification sequences for the skull and postcranial bones of each species. We then used an analysis method called Parsimov-based genetic inference to identify heterochronic changes in ossification sequence between species and to reconstruct ancestral ossification sequences throughout the carnivoran tree. Briefly, this method treats ossification sequence as a single complex character and generates trees to identify the minimum number of heterochronic shifts necessary to explain the data.
Carnivores are divided into two major lineages, the dog-like ‘caniforms’ and cat-like ‘feliforms.’ As luck would have it, each of these lineages includes a popular domestic species. We found that both the dog and cat are good models for the ossification sequence of their respective linages. This is a helpful finding since dogs and cats are much easier to study than other species, making them very useful models in both veterinary science as well as carnivoran biology in general. Pinniped species, however, were the exception; seals and sea lions showed differences in the timing and order of ossification of different bones – i.e. skeletal heterochronies – when compared both to each other as well as to the terrestrial carnivorans. This finding matches the high rates of heterochrony observed in the sutures of the crania of adult pinnipeds and may be related to adaptations to aquatic locomotion.
The micro-CT scans generated through this study are now publicly available via the Morphosource repository and we hope these data will facilitate the inclusion of embryonic data in future studies of carnivoran biology. We encourage other researchers to explore the collections available at their local natural history museum to gain critical insights into development and evolution.
We would like to thank the Genes, Organisms, and Ecosystems Research Experiences for Undergraduates (GEO REU) program for making this research possible, our reviewers for helpful comments on this manuscript, and the editor for inviting us to write this post.
Follow the Topic
-
BMC Zoology
This is an open access, peer-reviewed journal that considers articles on zoology, including comparative physiology, mechanistic and functional studies, morphology, life history, animal behavior, signaling and communication, cognition, parasitism, systematics, biogeography and conservation.
Related Collections
With Collections, you can get published faster and increase your visibility.
Advancing zoological research through museum collections
BMC Zoology is calling for submissions to our Collection, Advancing zoological research through museum collections.
Zoological collections in museums have long served as essential resources for scientific research, education, and conservation. These collections house valuable specimens and associated data that offer insights into the biology, behavior, physiology, evolution, and ecology of both extant and extinct species. Recent advances in technology and data management, including museomics, collectomics, imaging, and virtual collections, are transforming how researchers and the public access and use these collections, opening new avenues for discovery.
The significance of zoological collections extends far beyond traditional taxonomy and species identification. New computational and molecular approaches, such as museomics and collectomics, now enable the study of genetic material, ancient biomolecules, morphological and ecological data, and other forms of “dark data” historically locked within specimens. These methods have the potential to reveal previously inaccessible zoological information, illuminate evolutionary histories, and inform conservation strategies. At the same time, it is becoming increasingly clear that certain preservation techniques may affect the integrity of such information, and that destructive sampling requests must be carefully evaluated with guidelines in place to ensure raw data is shared. Developing clear workflows and guidelines is therefore important to ensuring that valuable data locked in specimens are not inadvertently lost.
In parallel, new virtual collections and data-sharing platforms are broadening access to museum resources and facilitating collaborative research across institutions and disciplines. These developments underscore the importance of integrating zoological collections into interdisciplinary research frameworks to maximize their impact on science and society. Strengthened partnerships between museums, universities, conservation organizations, and the wider public are also important to fully unlocking the potential of these collections to support zoological research, conservation priorities, and education.
The Collection will consider studies on:
•Unlocking dark data in zoological collections, including museomics, collectomics, computational, molecular, or imaging approaches that reveal hidden or previously inaccessible information.
•Advances in ancient DNA and biomolecule recovery from zoological specimens. Evaluating the impact of preservation techniques on the integrity of genetic, morphological, biochemical, or other data contained in specimens
•Optimizing specimen preservation workflows, including best practices to maintain long-term data quality
•Development or assessment of guidelines for destructive sampling, including strategies to minimize data loss and improve decision-making frameworks
•Creation or enhancement of virtual collections, including digitization, 3D imaging, and online accessibility tools
•Open science practices and data-sharing workflows, guidance, or platforms that improve access to information sourced from zoological collections for research, education, and conservation
•Knowledge-exchange initiatives between museums, researchers, conservation practitioners, and the public
•Interdisciplinary approaches integrating museum collections with fields such as ecology, evolution, conservation science, informatics, and education
This Collection supports and amplifies research related to SDG 14: Life Below Water and SDG 15: Life on Land.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Oct 29, 2026
Animal movements, migration, and range shifts
BMC Zoology is calling for submissions to our Collection, Animal movements, migration, and range shifts.
The study of animal movements, migration, and range shifts is crucial for understanding ecological dynamics and the effects of environmental change on wildlife populations. As species navigate complex landscapes, their movement patterns provide essential insights into behavioral ecology, habitat use, and evolutionary processes. This Collection invites research that explores the mechanisms and influences behind these movements, including the effects of climate change, habitat fragmentation, and human activities. By examining the interplay between environmental factors and animal behavior from the Arctic tundra to tropical forests, open oceans, and freshwater systems, we can enhance our understanding of biodiversity and conservation.
Understanding animal movements and migrations has never been more important, particularly in the context of rapid environmental changes and biodiversity loss. Advances in tracking technologies, such as GPS and remote sensing, have significantly improved our ability to monitor wildlife movements and assess migratory patterns. This enhanced capacity allows researchers to investigate how animals adapt to shifting habitats, exploit new resources, and respond to anthropogenic pressures. By unraveling these complexities, we can inform conservation strategies and policies aimed at protecting vulnerable species and ecosystems.
Continued research in this field may lead to the development of innovative management practices, such as the design of effective wildlife corridors and the enhancement of migratory routes that facilitate species resilience. Future studies could also enhance our understanding of how species may adapt to climate-induced range shifts and the implications for ecosystem health and function. The integration of interdisciplinary approaches will be vital in addressing the challenges posed by human-wildlife interactions and fostering a sustainable coexistence with nature.
Areas of particular interest include, but are not limited to:
Navigational strategies in migrating animals, including sensory cues and the role of instinct, learning, and social interactions in route selection
Mechanistic modeling approaches for studying migration behavior and predicting movement patterns
Advances in wildlife tracking technologies and methodologies for monitoring movement and migration
Impacts of human activities such as habitat fragmentation, pollution, and climate change, on migratory patterns
Climate-driven range shifts and their ecological and conservation implications
The role of wildlife corridors, stopover sites, and habitat connectivity in supporting migratory species
This Collection supports and amplifies research related to SDG 14: Life Below Water and SDG 15: Life on Land.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Aug 07, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in