
As microbial ecologists, we’re getting pretty good at collecting metagenomic data and recovering metagenome assembled genomes (MAGs) by now. We’re all at it.
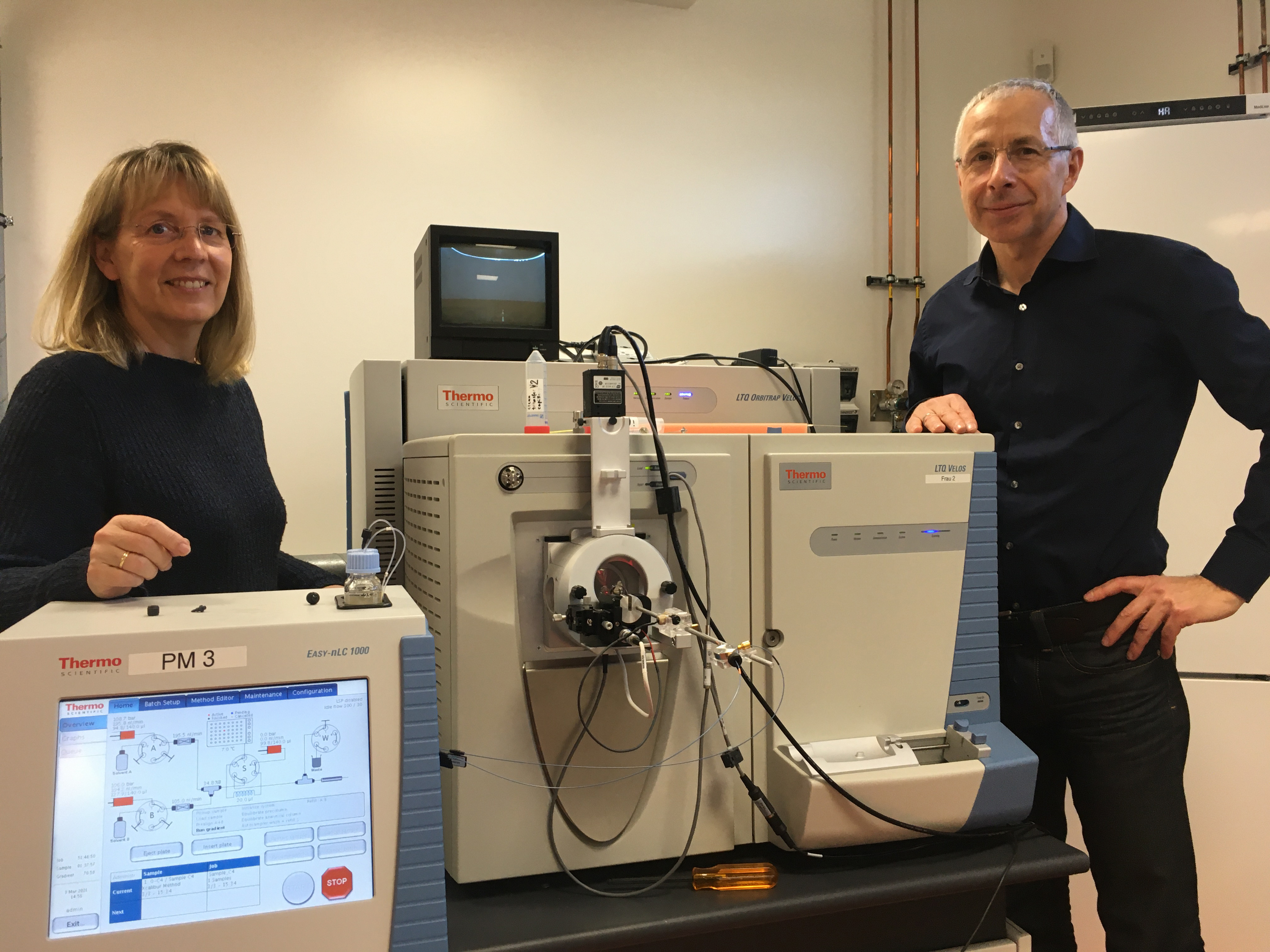
In our group we’ve been running sampling campaigns during spring phytoplankton blooms at the North Sea island of Helgoland for over a decade. In partnership with colleagues at the University of Greifswald we combine molecular microbial ecology and metagenomics with proteomics and metaproteomics to address not just the potential function of marine bacterial heterotrophs, but their activity as well.
In our latest collab, we did something we hadn’t managed before. In earlier years, we were able to get metaproteomes from a few key dates during spring blooms where we had corresponding metagenome data. But this time we sequenced metaproteomes in a time series across the entire spring bloom period, much like we’d previously done with metagenomic time series. This is where we made the most of our different skillsets, with the metaproteomics team generating very deeply sequenced metaproteomes that the metagenomics team could compare with our MAG database.

We’d learned a lot from earlier metagenomic series: what bacteria were present, how abundant they were, and a fair amount of detail about their implied polysaccharide substrate preferences. (Our major research theme is the fate of the large polysaccharide fraction of phytoplankton biomass released when algal populations die off.) We also knew that when bacterial abundance peaked, proteomes indicated large investments in TonB-dependent transporters specific to individual oligo/polysaccharide substrates.
But with this study we wanted to know more. Specifically, in what ways do the bacteria actually respond to the availability of polysaccharide? Do the responses fit with our assumptions about substrate quality? And finally, can expression of transporters be used for tracking uptake, and by inference, degradation, of polysaccharides that can be difficult to measure directly?
What the data showed was that, as usual, the bacteria in these environments commit a large amount of resources to their polysaccharide transporters. But they don’t appear to commit to individual substrates all at once. Instead it looks as though the community preferences for different substrates change from ‘easy’ substrates to ‘hard’ over time. By way of analogy, it’s as if the hot cakes are all selling like, well, y’know, and so anyone who got up late has to make do with the day-old bread. Or alternatively it could be more like when everyone’s done with their croissant, the pigeons come down and pick up the crumbs. To read that analogy translated into science, check out the paper!
Banner image: Coastline at Helgoland in the North Sea. © Max-Planck-Institute for Marine Microbiology, Naomi Esken

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in