So I have finally finished collecting my thoughts from the 3rd EMBO meeting in Vienna, which I was fortunate enough to attend from the 10th-13th Sept.
Hosted in the Austria Centre Vienna

the meeting was attended by around 1300 people, and with over 600 posters and 21 concurrent sessions there was lots to keep participants busy! As my first conference experience for a good few years, it was a little overwhelming, but in a good way!
Vienna
I found Vienna to be a lovely city, as was the weather for the weekend, which helps with the laid back atmosphere…
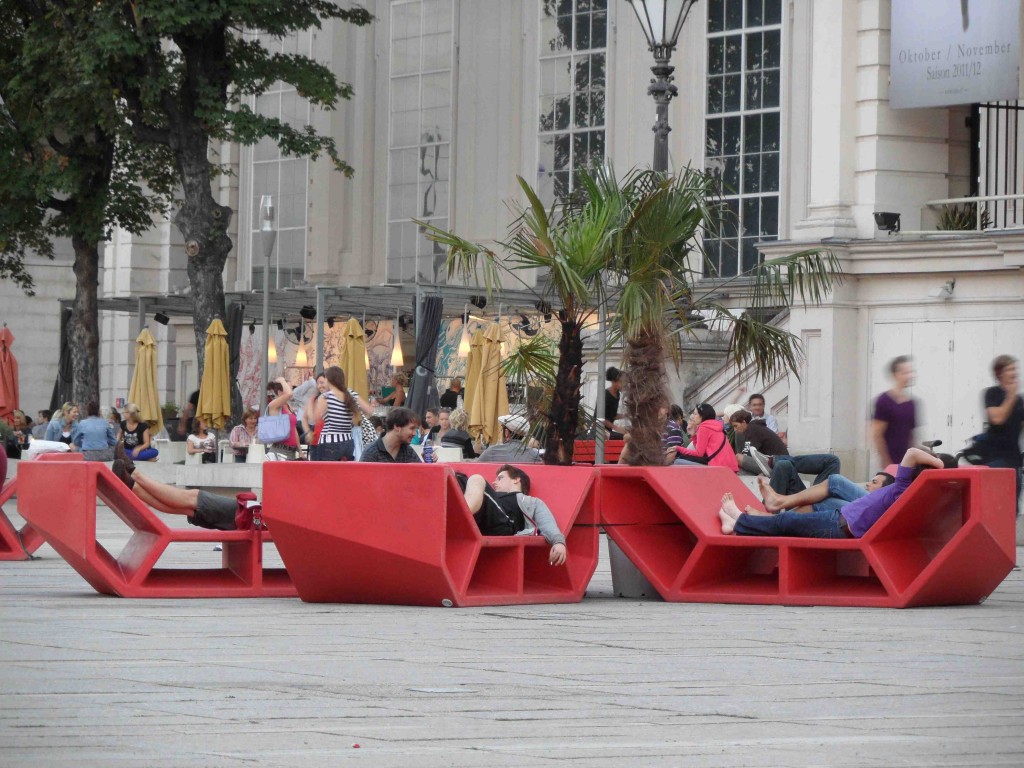

The architecture is amazing, with beautiful buildings on every corner
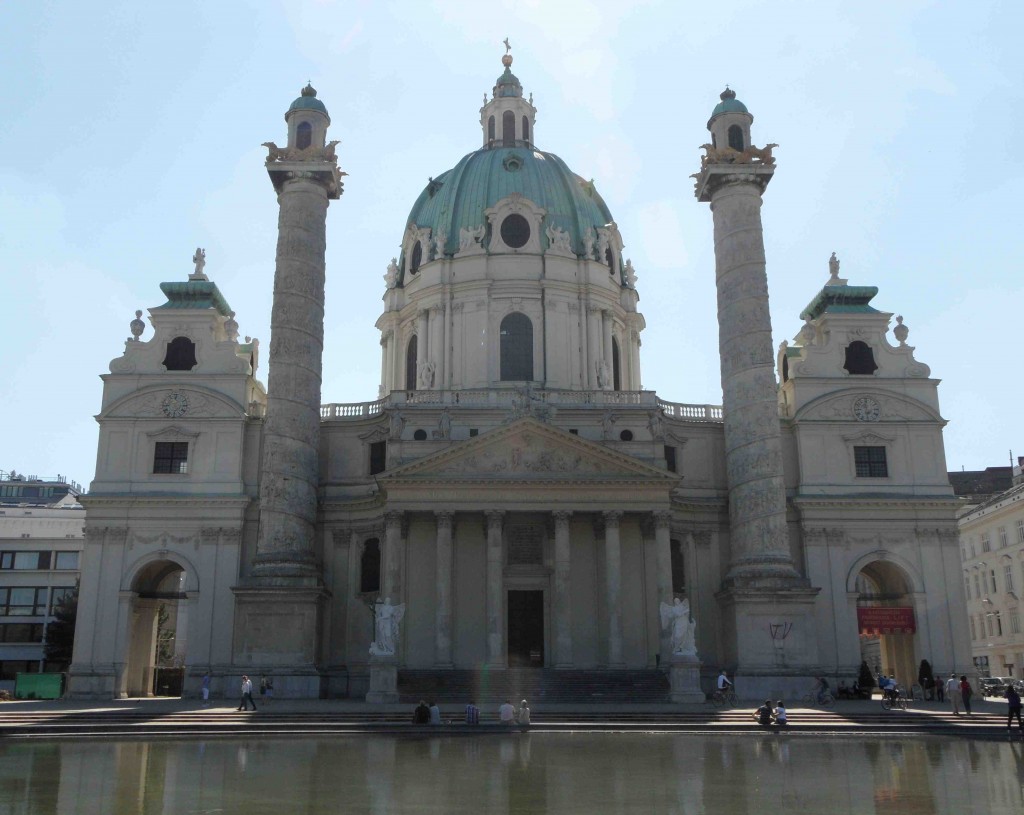

and although it was a little too cultural for my time-frame with many art galleries and museums that I didn’t manage to see, I did appreciate my visit to see the living art of the Spanish Riding School! Sadly we weren’t allowed to take pictures of the action, but here is one of the stars dozing in the sun
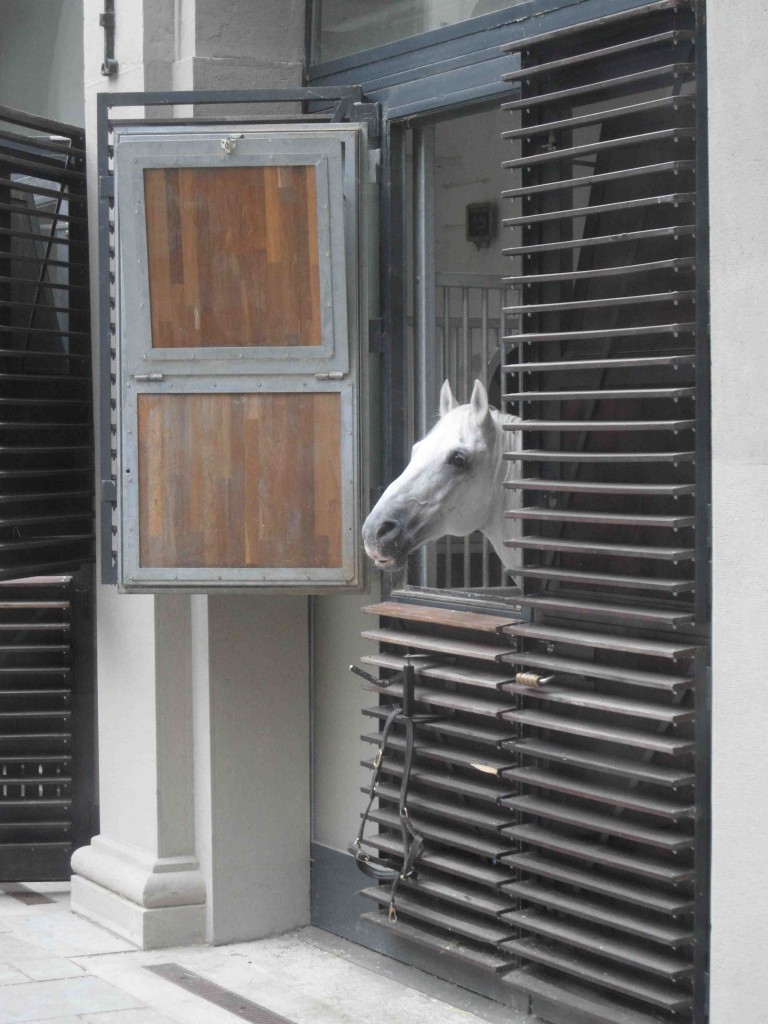

Anyway, on to the conference, which began with a series of plenary lectures on hosts & microbes, including a really interesting talk by Bonnie Bassler who explained their work investigating quorum sensing inhibitors as potential therapeutics to prevent pathogenesis. Although there is still some way to go before inhibitor compounds are made into drugs, this sounds like an exciting avenue of future research, especially given the multidrug resistance problems nowadays. I also really enjoyed the final talk in this session from Brett Finlay (despite the high content of toilet humour!) on the importance of our gut microbiota in our health and their role in resistance to allergic diseases such as asthma.
For the next 3 days, there was a great diversity of topics presented so I was too often spoilt for choice about which session to attend! Day two started with an excellent keynote lecture from Susan Lindquist discussing the role of protein folding in the inheritance of environmentally acquired traits via both Hsp90- and prion-mediated mechanisms. She explained how Hsp90 buffers the effects of genetic variation and acts as a potentiator of evolution, enabling previously hidden genetic variation to be revealed in times of stress. In the plenary session on genome evolution that followed, Paul Rainey presented evidence of the involvement of an epigenetic switch in bacterial genome evolution, reinforcing the theme that genome evolution is about so much more than just the underlying genetic variation.
Next-generation sequencing
Another theme that spanned many sessions was the use of next-generation sequencing and the advances made possible in a variety of fields as a result of the development of these techniques. This was illustrated by Mike Stratton’s plenary lecture on the exome sequencing and analysis of cancer genomes. Hand-in-hand with such technological advances of course comes the need for methods to analyse the vast quantities of data produced. These ideas were further explored in the session on human genome variation & disease with a talk from Elaine Mardis covering whole genome sequencing approaches used to analyse cancer genomes, including their application to personal diagnosis and treatment.
Personal medicine
Continuing on the topic of personal medicine, Russ Altman from Stanford next introduced a pharmacogenomics database designed to allow the effect of human genetic variation on drug responses to be assessed. This became quite entertaining when he reported results from the analysis of the personal genome of his colleague, Steve Quake; he had obviously agreed to everyone at EMBO and beyond knowing how he is likely to respond to a variety of drugs as well as his genetic predisposition to various diseases, but it is quite wierd to feel like I know far more about his genome than about the man himself?!
Computer vision
As well as the increase in the throughput of sequencing methods, there is also now a need for computational methods to keep up with advances in imaging methods. This was explored in a stimulating session on computer vision of cellular function in which Andre Brown and Sean Megason (amongst others) described systems to image C. elegans behaviour and zebrafish embryo development, respectively, as well as outlining the software they designed to analyse the image data obtained. The real take home message from this session was that it’s no use having the means to obtain data on this scale if you don’t also have algorithms and software to enable efficient and accurate analysis of this data to extract the biological meaning. So I think this will be a big area for methods development going forwards.
TNTs
I also found a new area of molecular biology to me and was really interested to learn about the recent discovery of tunneling nanotubes (or TNTs). In a talk by Hans-Hermann Gerdes we heard how these nanotubes form and that they are used for long distance electrical coupling between cells, and as a method of cell-cell communication for the transfer of cytoplasmic molecules and organelles. Fascinating stuff!
And finally, the prize for the best talk goes to…
Florian Engert, for his virtual reality world for zebrafish larvae. Or as he put it, ‘The Matrix’ for fish! He presented work on adaptive motorlearning in zebrafish larvae and detailed the system he developed in which larvae are paralyzed and surrounded by a virtual reality environment, which allows the whole brain to be imaged at a single cell level in an awake animal as they respond to experimentally induced changes in the environment. This clearly provides an exciting system to study the neural circuitry underlying a range of visually-induced behaviours in zebrafish.
Featured protocols
I will finish with a reminder of a couple of Nature Protocols that I was really pleased to spot being put to good use in the work presented at EMBO.
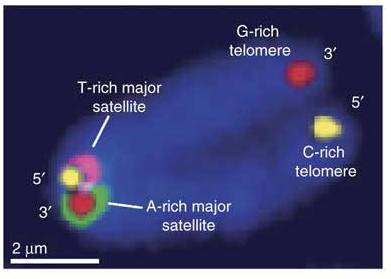
Firstly, “CO-FISH “:http://www.nature.com/nprot/journal/v5/n7/full/nprot.2010.102.html (chromosome orientation fluorescent in situ hybridization) was being utilised by both Shahragim Tajbakhsh and Peter Lansdorp in their talks on the non-random segregation of chromosomes in mitosis.
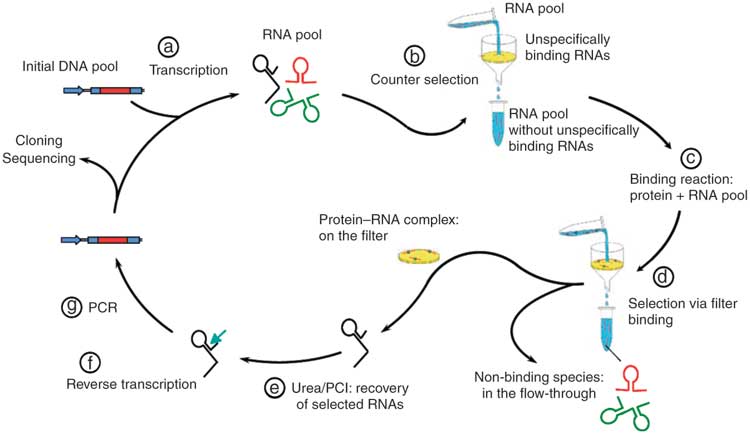
And genomic SELEX formed the basis of the work presented in Renee Schroeder’s talk in the RNA & RNA mediated regulation session.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in