Novel Agents for Chronic Lymphocytic Leukemia to Address Resistance
Published in Cancer and Genetics & Genomics
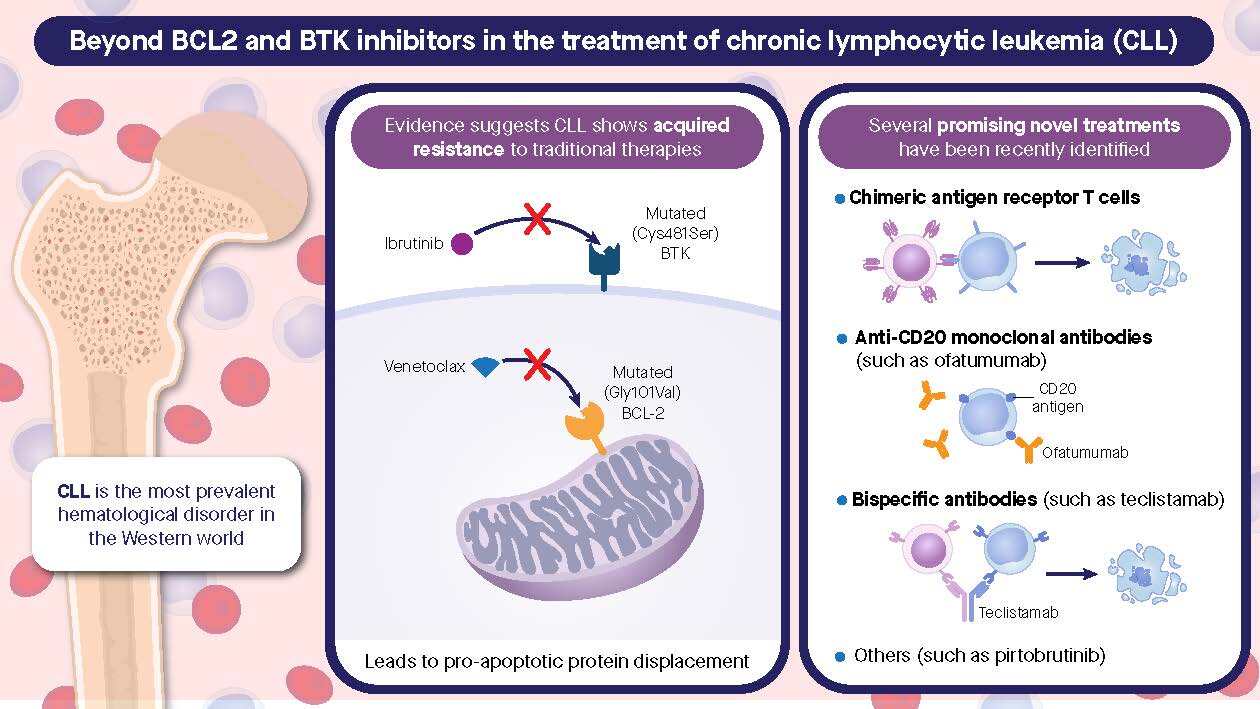
BTK Degraders
- Mechanism: These agents degrade Bruton tyrosine kinase (BTK), a key protein in B-cell receptor signaling, which is essential for CLL cell survival.
- Examples: NX-2127, BGB-16673, and NX-5948 have shown efficacy in relapsed/refractory (R/R) CLL.
- Non-Covalent BTK Inhibitors
- Mechanism: These reversible inhibitors target BTK without relying on covalent binding, overcoming resistance caused by mutations like C481S.
- Examples:
- Pirtobrutinib: Demonstrates high selectivity and efficacy, even in heavily pretreated patients.
- Nemtabrutinib: Similar efficacy but less well-tolerated compared to pirtobrutinib.
- BH3 Mimetics
- Mechanism: Mimic pro-apoptotic proteins to inhibit anti-apoptotic BCL2 family proteins, promoting cell death.
- Examples: Venetoclax targets BCL2, while newer agents like sonrotoclax aim to overcome resistance caused by mutations like Gly101Val.
- Monoclonal Antibodies
- Mechanism: Target antigens on CLL cells (e.g., CD20, CD19, CD37) to induce apoptosis or recruit immune cells for cytotoxicity.
- Examples:
- Obinutuzumab: Anti-CD20 antibody with improved efficacy over rituximab.
- Otlertuzumab: Anti-CD37 antibody triggering apoptosis and antibody-dependent cell-mediated cytotoxicity.
- Bispecific Antibodies
- Mechanism: Bind both CLL cell antigens (e.g., CD19, CD20, BCMA) and CD3 on T cells, redirecting T cells to kill leukemia cells.
- Examples:
- Blinatumomab: Targets CD19 and CD3.
- Teclistamab: Targets BCMA and CD3.
- Chimeric Antigen Receptor (CAR) T Cell Therapies
- Mechanism: Genetically modified T cells express CARs targeting antigens like CD19, enabling direct killing of CLL cells.
- Examples: Lisocabtagene maraleucel (CD19 CAR T therapy).
- Siglec-6 Monoclonal Antibodies
- Mechanism: Target Siglec-6, a novel antigen absent on healthy cells, to activate T cells and eliminate CLL cells.
- Examples: RC-1 and RC-2 antibodies engineered for high specificity and potency.
- ROR1-Directed Therapies
- Mechanism: Target ROR1, a receptor tyrosine kinase expressed selectively on CLL cells.
- Examples: Cirmtuzumab and zilovertamab vedotin.
- Vg9Vd2-T Cell Engagers
- Mechanism: Activate Vg9Vd2-T cells to lyse CLL cells by targeting CD1d, a molecule expressed on leukemic cells.
- Example: Bispecific single-domain antibodies designed to boost T cell responses.
- Anti-BAFF Antibodies
- Mechanism: Target BAFF, a factor promoting CLL cell survival, to induce apoptosis.
- Examples: Belimumab and ianalumab.
- Precision Medicine Approaches
- Mechanism: Use predictive biomarkers (e.g., TP53, IGHV mutations) and machine learning algorithms to tailor treatments based on tumor heterogeneity and resistance profiles.
These novel agents address unmet clinical needs by targeting specific pathways, overcoming resistance mechanisms, and enhancing immune responses, offering hope for improved outcomes in CLL treatment.
Follow the Topic
-
Discover Oncology
This is a fully open access general oncology journal that aims to provide a unified forum for researchers and clinicians. The journal spans from basic and translational science, to preclinical, clinical, and epidemiology, and welcomes content that interfaces at all levels of cancer research.
Related Collections
With Collections, you can get published faster and increase your visibility.
Single-Cell RNA Sequencing in Cancer Immunotherapy
Cancer immunotherapy is a hot area of current oncology research, with its core focus on activating or enhancing the body's immune system's ability to recognize and kill cancer cells. However, cancer cells possess complex heterogeneity and dynamics, which affect the efficacy of immunotherapy in many ways. Single-cell RNA sequencing (scRNA-seq) has emerged as a powerful tool in recent years, providing us with an unprecedented insight into the cellular heterogeneity and dynamics within tumors. This technology has revolutionized our understanding of cancer biology, especially in the context of cancer immunotherapy. By enabling researchers to analyze individual cells, scRNA-seq allows them to identify distinct cell populations, track cellular responses to treatments, and discover new therapeutic targets. This collection aims to compile cutting-edge research in this field and explore the various applications of single-cell RNA sequencing in cancer immunotherapy.
This collection will cover the following topics: 1. The latest advances in single-cell RNA sequencing technology in cancer immunotherapy, including research on technology optimization and data interpretation; 2. Using single-cell RNA sequencing to reveal the characteristics of immune cell subgroups in the tumor microenvironment and their interaction mechanisms with cancer cells; 3. Analyzing the molecular basis of immune therapy response and resistance through single-cell RNA sequencing, exploring new biomarkers and therapeutic targets; 4. Combining single-cell RNA sequencing with clinical studies of immunotherapy to assess treatment outcomes, predict patient prognosis, and optimize treatment plans.
Keywords: cancer immunotherapy, single-cell RNA sequencing, therapeutic targets, tumor microenvironment, treatment response
Publishing Model: Open Access
Deadline: Jun 30, 2026
Computational and Analytical Methods for Multi-omics Approaches in Cancer Research
Exploring the molecular mechanisms, clinical outcomes, and their interplay is crucial for cancer research. Several recent research works have highlighted the importance of understanding inter- and intra- relationship among molecular mechanisms assayed at several layers of biological process, e.g. genomic, transcriptomic, proteomic, metabolomic, and microbiomics. Advanced analytical methods are needed to better understand the crosstalk signals among the biological processes and to discover molecular markers for future therapeutic targeting. Understanding of such molecular processes and clinical outcomes will provide a more holistic approach to investigating cancer biology and therapeutics. Overall purpose of this collection is to provide a platform for discussion on emerging analytical methodologies for studying tumor heterogeneity, diversity, progression, and treatment response with a focus on advancing our knowledge of the complex disease better.
Considering the recent advances in statistical, bioinformatics, and machine learning approaches in cancer research, we propose a collection of the topics aiming to facilitate interdisciplinary dialogue and collaboration among researchers working in quantitative and clinical aspects of cancer biology and therapeutics. Authors are encouraged to submit articles that investigate the molecular mechanisms underlying the disease heterogeneity and progression, as well as studies that explore the potential therapeutic strategies targeting these interactions. We welcome submission of original research articles addressing the following themes but not limited to: (1) new analytic methods covering both traditional statistical, bioinformatics, and advanced machine learning methods, (2) review of the literature and perspectives, (3) tutorial on any recently developed analytical methods, (4) imaging and artificial intelligence analysis methods, etc. Submission of the articles from all types of cancer research using any type of quantitative analysis methods is welcome.
Keywords: tumor heterogeneity, statistical methods, bioinformatics, machine learning, cancer biology
Publishing Model: Open Access
Deadline: Jul 31, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Part of a collection:
The Convergence of Targeted Precision Medicine: Future of Oncology Therapeutics