Nuclear magnetic resonance spectroscopy in medical research: Workflow and possibilities
Published in Biomedical Research

NMR is a measurement method that can detect a variety of metabolites, such as amino acids, ketone bodies, lipoproteins, as well as some inflammatory parameters. The areas of application of NMR are broad and include metabolic diseases such as diabetes, infectious diseases, neuropsychiatric diseases such as Parkinson's disease and Alzheimer's dementia, as well as the large field of cancer. NMR metabolomics is a growing field of research with the goals of enabling earlier diagnosis, more specific therapies, and monitoring of therapy response, paving the way towards personalized medicine. An example of the wealth of information that can be obtained using NMR is the analysis of lipoproteins in serum. The lipoproteins HDL, LDL, IDL and VLDL can each be broken down and measured according to particle size and density. For HDL, there are four sizes; for LDL, for example, there are six. Furthermore, the proportions of their individual components can be measured. These include apolipoproteins, cholesterol, free cholesterol, phospholipids, and triglycerides. Since the lipoprotein profile is altered in many diseases and plays an essential role in metabolic processes, more precise insights into disease processes and diagnostics can be gained.
Advantages of NMR
In our study, we worked with serum samples, although other sample materials can also be examined using NMR, for example urine, cerebrospinal fluid, organ, and tumor tissue. NMR offers several advantages over other measurement methods in the field of metabolomics research, such as mass spectrometry. Sample preparation for NMR measurements requires only a few steps, little sample material is needed (in the case of serum mostly 400 µl) and the sample material is not destroyed, so it can be measured as often as needed. Furthermore, many samples can be measured in a short time, which makes the work very efficient. NMR can therefore be used to measure and compare samples from large cohorts, for example, to determine disease-related differences. However, very specific questions can also be investigated, e.g., metabolic processes in tumor tissue.
Background of the patient samples
The serum samples of the COVID-19 cohort came from Heidelberg. Our research group, consisting of Christoph Trautwein, Georgy Berezhnoy and me, collaborated here with Uta Merle from Heidelberg. The samples were collected as part of a study initiated and led by Professor Merle, which ran from September 7, 2020 to May 17, 2021. This study was intended to monitor and care for COVID-19 patients through a "coronataxi".
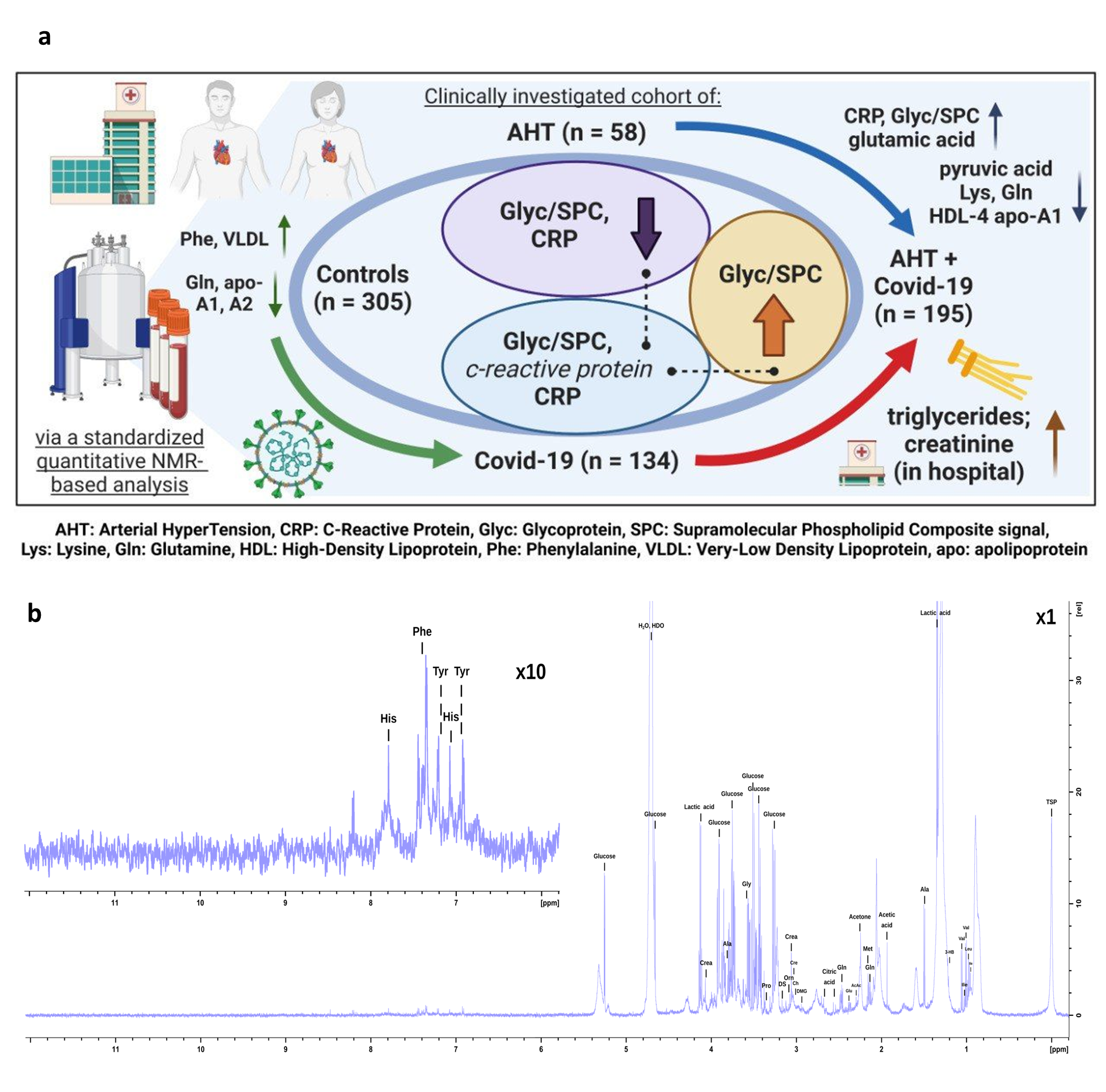
a This figure is intended to summarize the work process and results of this study. The three cohorts are shown with numbers of participants, as well as the subgroup AHT + COVID-19 with 195 patients. As typical striking metabolic parameters in COVID-19 disease, an increase of CRP, Glyc/SPC and gluatmic acid, as well as a decrease of pyruvic acid, lysine, glutamine, and HDL-4 fractions like HDL-4 apolipoprotein A1 are noted.
b illustrates a CPMG spectrum of quantified metabolites. The aromatic region (8.0 to 6.0 ppm) is zoomed by factor 10. HDO = semi heavy water, Crea = creatinine, DS = dimethylsulfone, Cre = creatine, Ch = choline, DMG = N, N-dimethylglycine, AcAc = acetoacetic acid, 3-HB = 3-hydroxybutyric acid, Val = valine, Ile = isoleucine, Leu = leucine, TSP = 3-(trimethylsilyl) propionic-2,2,3,3-d4 acid.
Workflow in a NMR laboratory and statistical analysis
After measuring the samples, the spectra (exemplary in Fig. 1 b) must first be translated into absolute concentrations for the individual metabolites. For this purpose, the data of the COVID- 19 cohort were transferred to Bruker BioSpin GmbH in Ettlingen. With the data the statistical questions could now be investigated. Very common for this is Metaboanalyst 5.0, a software package with univariate and multivariate statistical operations.
COVID-19 and hypertension: The main part of the Paper
In addition to the COVID-19 cohort and the healthy control cohorts, we also had the opportunity to measure samples from a cohort of patients suffering from hypertension and include them in our analyses. These samples came from a cohort at the University Hospital Tübingen and were stored in the Interdisciplinary Biomaterial and Database of the University Hospital Würzburg in Würzburg.
I prepared the samples for this cohort (AHT control cohort) with our technical assistant and measured them with the help of Georgy Berezhnoy. Since sample preparation for blood sera is so straightforward and the workflow is always the same, this was easy to do even for a lab novice like me and was done within a week. Once we received the serum concentration data from Bruker, I was able to compare the NMR data from this cohort with that of the hypertensive COVID-19 patients. We also wanted to investigate what differences were present in the metabolome of hypertensive COVID-19 patients, since hypertension was often mentioned in the literature as a risk factor of severe COVID-19 progression.
The international COVID- 19 Network
After some time of statistical analysis, I gained some interesting results and had the opportunity to present them at our international NMR COVID network meeting. The meeting was part of the NMR international COVID- 19 research Network, which was launched by Bruker in April 2020. In this network, research groups from Australia, Germany, Italy, Spain, Austria, Norway, and the USA are conducting research and exchanging information on NMR and COVID-19. My results regarding the metabolic profile of COVID-19 patients were generally consistent with the previous results of the working groups and the already published literature. However, the approach of comparing COVID-19 patients with nonCOVID-19 subjects within a group with a specific salient feature (in this case, hypertension) to remove this factor from the analyses was still novel. This provided a more accurate picture of the metabolic profile of COVID-19 disease.
Final words
One challenge in writing the paper was to present the results in a clear and understandable way. After all, NMR measurements provide a mass of data. In addition, the NMR methodology and its evaluation in applicable clinical medicine is still very new and little established. Papers like this one should change this and show what additional information can be generated by NMR, which opens up further possibilities for a better understanding of disease processes and diagnostics.
Follow the Topic
-
Communications Medicine

A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary across all clinical, translational, and public health research fields.
Related Collections
With Collections, you can get published faster and increase your visibility.
Reproductive Health
Publishing Model: Hybrid
Deadline: Mar 30, 2026
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in