Quantifying polysaccharide uptake by marine bacteria, one cell at a time
Published in Microbiology
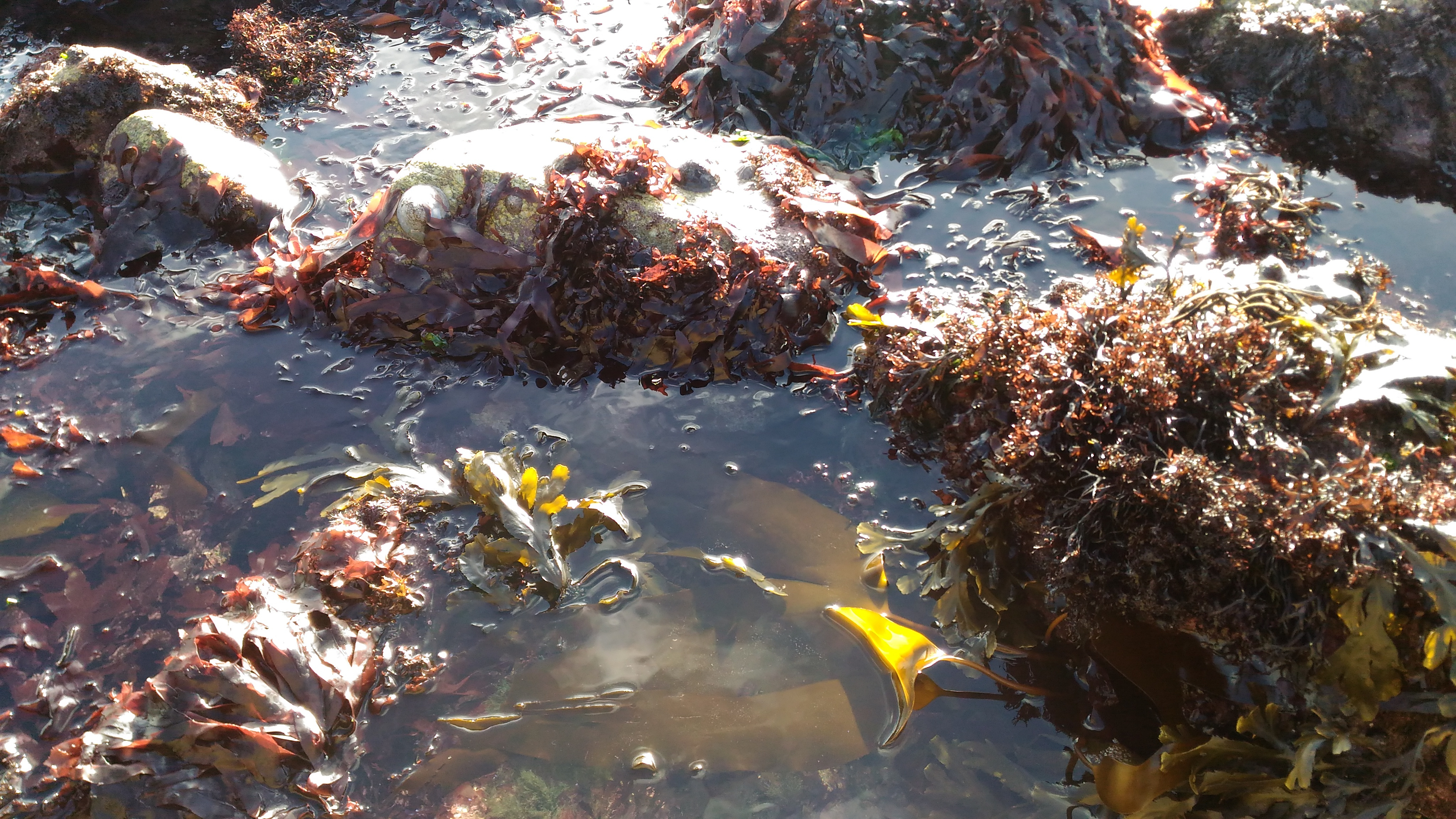
People lucky enough to live along the coast or to spend vacations at the seashore know that seaweeds (a.k.a macroalgae) can be very abundant and create genuine submarine forests. Indeed, macroalgae are considered dominant primary producers in coastal environments. Through photosynthesis, they fix a large amount of carbon as polysaccharides that can further be released as dissolved or particulate organic matter in the seawater during their life cycle or when they eventually die. Algal polysaccharides therefore constitute a huge reservoir of substrates for the marine biosphere and especially the heterotrophic bacteria, which are central to the oceanic carbon cycle. Yet, every marine bacterium cannot use these compounds as particular enzymatic pathways are required. Identifying which bacterial taxa actively assimilate algal polysaccharides in seawater communities remains a challenge, let alone quantifying their individual contributions to polysaccharide recycling.

To tackle this question, we used stable isotope probing (SIP). This technique allows tracing the incorporation of substrates artificially enriched in stable isotopes into cellular material (here, 13C carbon). Yet, by contrast to simple compounds, no 13C-labeled algal polysaccharides are commercially available. We therefore had first to develop a new protocol to produce our own 13C-labeled algal biomass. We started from tiny juveniles of the kelp Laminaria digitata and grew them for 3 months in the laboratory with regular additions of 13C-bicarbonate that was incorporated in algal tissues. There definitely was a learning curve in pampering baby kelps, and it even got emotional when they all died due to an electric failure in the growth chamber! We eventually managed to obtain healthy algae in a subsequent attempt, from which we extracted 13C-enriched alginate, the major polysaccharide from kelps. With this precious alginate in hand, we could finally start the SIP experiments!

The alginate was added to coastal seawater samples collected along the shore of Roscoff (France) to trace its assimilation into specific bacteria within the microbial community. The results published in our recent paper show that most of the alginate is used by a few initially rare Flavobacteriaceae and Gammaproteobacteria taxa and provide the first single-cell estimates of alginate incorporation rates. We first combined SIP incubations with halogen in situ hybridization coupled to chemical imaging via nanoscale secondary ion mass spectrometry (HISH-SIMS). Aside from yielding colorful Andy-Warhol-style pictures (see below), this technique allowed us to specifically detect individual cells that had incorporated alginate, based on their isotopic content.
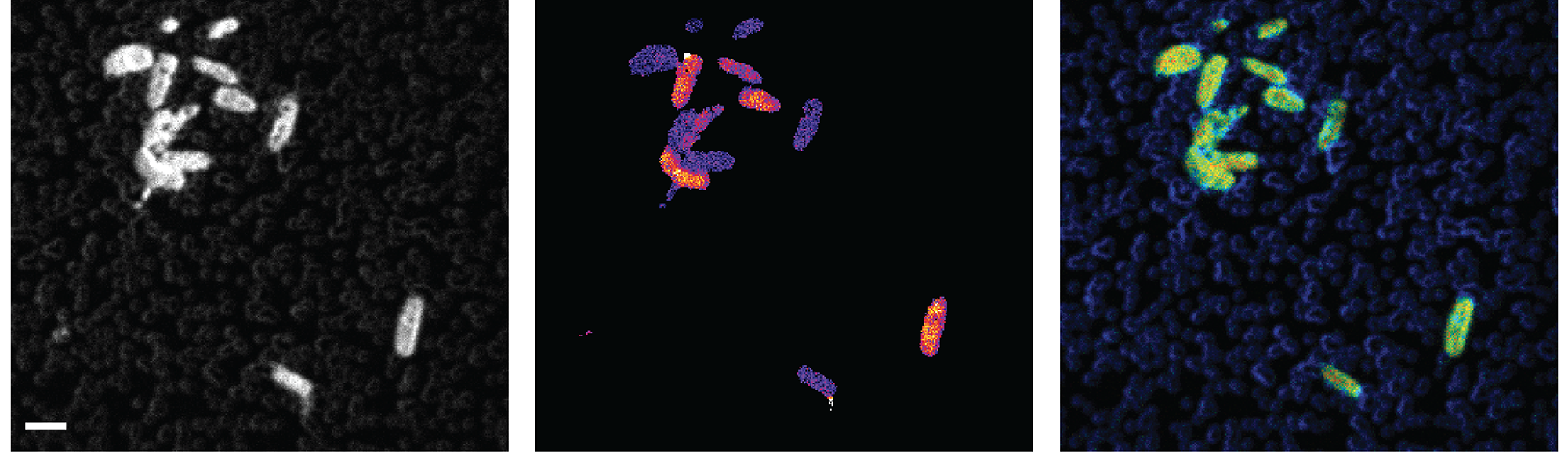
Rates of carbon assimilation from alginate were calculated for each individual cell using a mathematical treatment of HISH-SIMS data. They reached up to 27.5 fg C µm-3 h-1, as high as a quarter of the estimated partial density of carbon in bacterial cells. Total DNA from the same SIP incubations was further fractionated according to its buoyant density (the more the organisms have incorporated 13C from alginate, the "heavier" their DNA is) and analyzed using 16S rRNA gene metabarcoding to reveal the taxonomy of the alginate incorporators. Finally, functional screening of metagenomic libraries prepared from SIP incubations gave insights into the gene repertoire of some of these alginolytic bacteria.
Altogether, these novel results suggest that abundant algal polysaccharides might preferentially be consumed by a limited number of bacteria in coastal environments, highlighting the high degree of niche specialization in heterotrophic communities. Our estimates of uptake rates can help constrain the quantitative role these specialist polysaccharide-degrading bacteria play to unlock the carbon sequestered in algal biomass.
For full details, check our new paper at: https://rdcu.be/cj1R9.
Twitter: @FThomas_Roscoff



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in