Robust scoring of selective drug responses for patient-tailored therapy selection
Published in Cancer

Two flavors of precision cancer medicine
The current implementation of precision cancer medicine is mainly driven by genomics, and the existing and emerging molecular-clinical tumor boards rely extensively on genomic characterization to identify personalized therapeutic interventions. However, genomic biomarkers can guide treatment decisions only for a fraction of cancer patients, and more often genomic data are neither actionable nor predictive1,2,3; therefore, there is an increasing interest in using functional assays to guide cancer treatment by capturing information from functional perturbations of patient-derived cells, such as those that directly test patient cells to hundreds of anticancer compounds in multiple doses3,4,5. Such ex vivo drug testing can extend both the biomarker space and patient populations suitable for precision oncology treatment selection.
Functional precision medicine has been spearheaded in hematological malignancies, where drug sensitivity profiles can be generated in three days, enabling rapid clinical translation1,2. Clinical value has been shown both in terms of improved patient outcomes and clinically actionable therapeutic insights for individual patients6-9. However, there remain experimental and analytical challenges, concerning the reproducibility and translatability of the ex vivo drug testing results that need to be considered in patient management3,4,5,10,11.These challenges are related to the broader ‘reproducibility crisis’ in preclinical research12-15. Furthermore, most functional precision medicine studies use simple drug response metrics, which cannot quantify the selective effects of drugs, i.e., differential responses between cancer and non-cancer cells.
Our computational solution
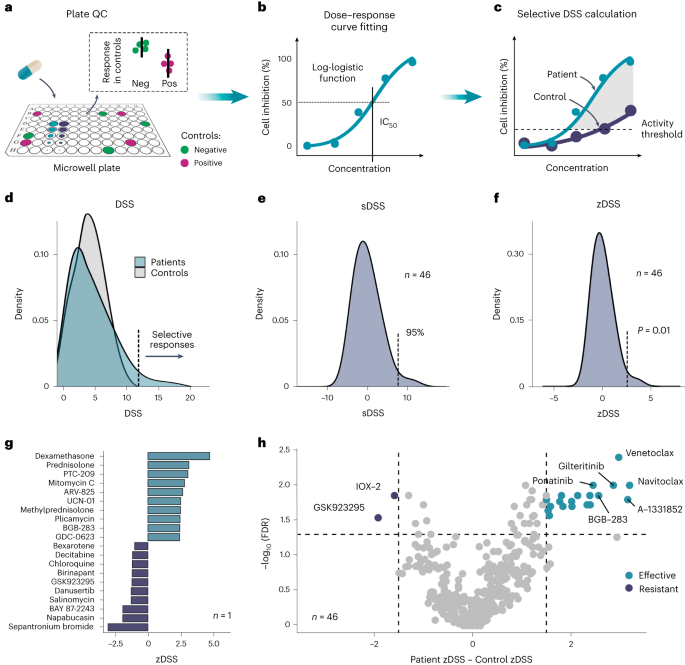
To address these challenges, we developed in 2014 a new computational approach, called drug sensitivity score (DSS)16, which integrates multiple dose-response parameters into a single response metric, and thereby enables accurate quantification of cancer cell sensitivity to both standard chemotherapies and molecularly-targeted drugs. Using pan-cancer screening data from two in vitro studies17,18, we showed that it is possible to achieve an improved consistency between different laboratories for drug response measurements by paying careful attention to harmonization of both laboratory assays and computational procedures19. Importantly, DSS enables identification of differential drug response patterns between cancer and healthy control cells. The selective response scoring uses the inhibition of non-cancerous cells as a proxy for potential drug toxicity, which can in turn be used to prioritize effective and safer treatment options.
Based on frequent requests from the functional precision medicine community, we have now implemented several new features, such as robust statistics and batch effect correction, which enable more reproducible quantification of cancer-selective effects of drugs. In this Protocol, we demonstrated its applicability in various precision medicine studies using real-life examples that showcase the best use of selective DSS calculation when optimizing treatment options for leukemia patients in three ongoing precision medicine studies in Europe and in the United States. The generic methods are widely applicable also to other malignancies that are amenable to ex vivo drug testing. The study was done in collaboration with researchers from the Institute for Molecular Medicine Finland (FIMM), Biotech Research & Innovation Centre (BRIC), Helsinki University Hospital (HUS), Orton Orthopaedic Hospital, Tampere University and Oslo University Hospital.
Implementation and future directions
The open R-source codes and example drug response data have been made openly available in GitHub. The new DSS calculations are additionally implemented in the Breeze data analysis platform maintained at FIMM and HiLIFE. The novel features implemented in the open-source software support the identification of targeted and non-toxic compounds for guiding clinical decision-making. We have also made available drug response profiles to 527 anticancer compounds tested in 10 healthy bone marrow samples as reference data for identification of cancer-selective drug responses and to de-prioritize drugs in preclinical testing that show cytotoxicity in the normal bone marrow cells. Since bone marrow toxicity is a common restrictive adverse effect among anticancer drugs, the healthy control bone marrow samples provide relevant control responses also for other cancer types than leukemia. We therefore hope that the selective DSS calculations will lead to more effective and safe therapy selections in many cancer types.
We and others are in the process of developing similar computational solutions also for co-culture assays, and for cell population-level drug testing assays, such as those based on imaging20,21 and flow cytometry22,23. Once tested at multiple sites and in large-scale screening, these methods are expected to provide us an improved means to identify patient-specific treatments that selectively inhibit cancer cells while avoiding inhibition of non-cancerous cells based on increasingly available ex vivo drug testing data from patients in real-world and clinical trial settings. For instance, in the VenEx trial, that is being implemented at HUS and FIMM, patient treatment optimization is based on flow cytometry DSS24. Beyond the monotherapy recommendations, ex vivo drug testing can be used also to guide the selection of drug combination treatments, as was shown, for instance, in patients with non-Hodgkin lymphoma25, as well as in solid tumors; e.g., across histological subtypes of ovarian carcinoma26.
References
- Kornauth, C. et al. Functional Precision Medicine Provides Clinical Benefit in Advanced Aggressive Hematologic Cancers and Identifies Exceptional Responders. Cancer Discov. 12, 372-387 (2022).
- Malani, D. et al. Implementing a Functional Precision Medicine Tumor Board for Acute Myeloid Leukemia. Cancer Discov. 12, 388-401 (2022).
- Letai, A., Bhola, P. & Welm, A.L. Functional precision oncology: Testing tumors with drugs to identify vulnerabilities and novel combinations. Cancer Cell 40, 26-35 (2022).
- Tognon, C.E., Sears, R.C., Mills, G.B., Gray, J.W. & Tyner, J.W. Ex Vivo Analysis of Primary Tumor Specimens for Evaluation of Cancer Therapeutics. Annu. Rev. Cancer Biol. 5, 39-57 (2021).
- Flobak, Å., Skånland, S.S., Hovig, E., Taskén, K. & Russnes, H.G. Functional precision cancer medicine: drug sensitivity screening enabled by cell culture models. Trends Pharmacol. Sci. 43, 973-985 (2022).
- Pemovska, T. et al. Individualized systems medicine strategy to tailor treatments for patients with chemorefractory acute myeloid leukemia. Cancer Discov. 3, 1416-1429 (2013).
- Tyner J.W. et al. Functional genomic landscape of acute myeloid leukaemia. Nature 562, 526-531 (2018).
- Yin, Y. et al. Functional Testing to Characterize and Stratify PI3K Inhibitor Responses in Chronic Lymphocytic Leukemia. Clin. Cancer Res. 28, 4444-4455 (2022).
- Bottomly, D. et al. Integrative analysis of drug response and clinical outcome in acute myeloid leukemia. Cancer Cell 40, 850-864 e859 (2022).
- Haibe-Kains, B. et al. Inconsistency in large pharmacogenomic studies. Nature 504, 389-393 (2013).
- Hatzis, C. et al. Enhancing reproducibility in cancer drug screening: how do we move forward? Cancer Res. 74, 4016-4023 (2014).
- Begley, C.G. & Ellis, L.M. Drug development: Raise standards for preclinical cancer research. Nature 483, 531-533 (2012).
- Errington, T.M. et al. Investigating the replicability of preclinical cancer biology. Elife 10, (2021).
- Rodgers, P. & Collings, A. Reproducibility in Cancer Biology: What have we learned? Elife 10 (2021).
- Mullard, A. Preclinical cancer research suffers another reproducibility blow. Nat. Rev. Drug Discov. 21, 89 (2022).
- Yadav, B. et al. Quantitative scoring of differential drug sensitivity for individually optimized anticancer therapies. Sci Rep 4, 5193 (2014).
- Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603-607 (2012).
- Garnett, M.J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570-575 (2012).
- Mpindi, J.P. et al. Consistency in drug response profiling. Nature 540, E5-E6 (2016).
- Heinemann, T. et al. Deep Morphology Learning Enhances Ex Vivo Drug Profiling-Based Precision Medicine. Blood Cancer Discov. 3, 502-515 (2022).
- Kropivsek, K. et al. Ex vivo drug response heterogeneity reveals personalized therapeutic strategies for patients with multiple myeloma. Nat Cancer. 4, 734-753 (2023).
- Kuusanmäki, H. et al. Phenotype-based drug screening reveals association between venetoclax response and differentiation stage in acute myeloid leukemia. Haematologica 105, 708-720 (2020).
- Ianevski, A. et al. Patient-tailored design for selective co-inhibition of leukemic cell subpopulations. Sci. Adv. 7, eabe4038 (2021).
- Kuusanmäki, H. et al. Ex vivo venetoclax sensitivity testing predicts treatment response in acute myeloid leukemia. Haematologica 108, 1768-1781 (2023).
- Goh, J. et al. An ex vivo platform to guide drug combination treatment in relapsed/refractory lymphoma. Sci. Transl. Med. 14, eabn7824 (2022).
- Murumagi, A. et al. Drug response profiles in patient-derived cancer cells across histological subtypes of ovarian cancer: real-time therapy tailoring for a patient with low-grade serous carcinoma. Br. J. Cancer 128, 678-690 (2023).
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in