
Pollinators are the keystone for global biodiversity, with approximately 87.5% of flowering plants (30,8000 species) being pollinated by insect and vertebrate pollinators1. The mutualisms between pollinators and plants are generally considered to crucially drive the evolution of both species groups via reciprocal adaptation, which results in adaptive radiation and speciation. Therefore, it is of great importance to unravel the underlying molecular mechanisms of this mutualism type, as they not only improve our understanding of pollinator-plant coevolution but also are useful for predicting the fate of existing species in the ongoing global changes. However, at present, such mechanisms have not been unraveled even in a single pollinator-plant mutualism. This is mainly because (1) plant traits attracting pollinators are not well defined and often have multiple functions, and (2) mutualistic pollinators can be natural enemies to eat the plants at larvae phases.
The obligate pollinator-plant mutualisms with a long coevolutionary history and apparently matched traits provide an ideal system for uncovering molecular mechanisms underlying mutualism. Ficus species (producing figs) and their host-specific pollinating wasps (Agaonidae) comprise a classic example of obligate mutualism. Each Ficus species relies on only one or a few pollinating wasp species, and the pollinating wasps can only lay eggs into the ovules (galled ovules) of host plants where their larvae develop at the cost of plant nutrients2,3. Based on their tightly matched life history, two major connections between figs and pollinating wasps are: (1) host figs emit specific volatile compounds to attract pollinating wasps; and (2) larvae of pollinating wasps adapt to the environments especially the plant chemical defences inside galled ovules.
Ficus pumila var. pumila is widely distributed in China and was mentioned in the ancient Chinese poems in ‘The songs of Chu’ (c. 2200 years ago). As our research team has studied the phylogeography and population genetics of F. pumila var. pumila and its specific pollinator wasp Wiebesia pumilae (Figure 1) for more than 16 years, we decided to choose this fig-pollinator mutualism to decipher the genetic codes used by the two mutualists. Previous studies have found the specific attractants in some other fig-pollinator mutualisms4. However, due to lacking genomic and transcriptomic background, the molecular processes causing temporal-specific emission of attractants and pollinators’ recognition remains unexplored. The secondary metabolites associated with chemical defences (SMCDs) inside galled ovules have not been measured previously, and metabolism is competent to describe and the profiles of these chemicals. We thus designed to use multi-omics to solve the scientific questions.

Figure 1. Adult male (a) and female (b) individuals of W. pumilae and its adult females searching the entrance of host figs on a F. pumila var. pumila tree for either pollination or oviposition (c). The two photos of W. pumilae are provided by Dr. Jean-Yves Rasplus, and Yang Yang offers the other photo.
We then discussed the research plan with our collaborators. They presented constructive suggestions for electrophysiological experiments to identify the attractant(s) for the pollinator and functional validation of key regulatory genes. After improving the research plan, we launched our study in spring 2017.
During the period from 2017 to 2020, we combined multi-omics with validation experiments and finally revealed some molecular mechanisms underlying our target fig-pollinator mutualism. For F. pumila var. pumila (the plant), we identified the specific attractant (decanal) and repellent (nonanal) for pollinators and validated the function of several key genes in regulating temporal-specific biosynthesis of the attractant (Figure 2a). As to W. pumilae (the pollinator), we found that a highly reduced number of odorant-binding protein (OBP) genes to explain the pollinator’s specialized olfaction and that one OBP binds specifically to the attractant and initializes the pollination (Figure 2a).
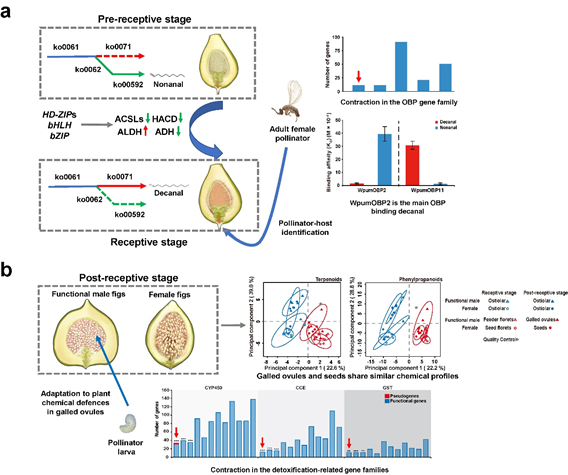
Figure 2. The molecular mechanisms causing specific fig-pollinator identification (a) and the adaptation of pollinator larvae to plant chemical defences in galled ovules (b).
When the two species are at the antagonistic stage (i.e., pollinator larvae utilize nutrients from host plants inside galled ovules), we found similar chemical profiles of SMCDs and turnovers throughout the development of galled ovules and seeds (Figure 2b). These findings suggest that galled ovules develop like seeds, i.e. seed mimicry, by which pollinator larvae can effectively avoid chemical sanction from host plants and feed on plant nutrients. Furthermore, a significant contraction of detoxification-related gene families in the pollinator genome (Figure 2b) indicates the specialized detoxification ability, probably reflecting that pollinating wasps spend most life span at larval stage in such a single environment.
So far, we have taken one step forward to reveal some molecular mechanisms of reciprocal adaptation between Ficus species and their specific pollinating wasps. Nevertheless, to further explore the factors driving the coevolution of both sides of mutualists, we are still facing challenges, for example, the incentives causing differential expression of the key transcription factors regulating the biosynthesis of attractant(s); the neurological processes leading to the preference of pollinating wasps to the attractant(s); and the molecular mechanisms forming galled ovules and the related genes in both mutualists. Future studies will continue to seek the interactive genes in fig-pollinator mutualisms integrating innovative techniques from molecular biology and neuroscience, and reconstruct the coevolution history based on these genes. Our study provides an example of gene-for-gene coadaptation, and the trait-based multi-omics can be applied to more diffuse pollinator-plant mutualisms.
Find out more on our paper: Wang et al. Molecular mechanisms of mutualistic and antagonistic interactions in a plant-pollinator association. Nat Ecol Evol (2021). https://www.nature.com/articles/s41559-021-01469-1
References:
- Ollerton, J., Winfree, R. & Tarrant, S. How many flowering plants are pollinated by animals? Oikos 120, 321‒326 (2011).
- Cook, J. M. & Rasplus, J.-Y. Mutualists with attitude: coevolving fig wasps and figs. Trends Ecol. Evol. 18, 241–248 (2003).
- Herre, E. A., Jandér, K. C. & Machado, C. A. Evolutionary ecology of figs and their associates: recent progress and outstanding puzzles. Annu. Rev. Ecol. Evol. Syst. 39, 439‒458 (2008).
- Chen, C. et al. Private channel: a single unusual compound assures specific pollinator attraction in Ficus semicordata. Funct. Ecol. 23, 941–950 (2009).
(Contributors: Rong Wang, Yuan-Ye Zhang)
Follow the Topic
-
Nature Ecology & Evolution
This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in