
The paper in Scientific Reports can be found here: https://go.nature.com/2IBohqq
This is my first Behind the Paper submission, and I am excited to share my work in this forum. The posts in this channel span disciplines, but one theme is clear - we enjoy our science. I am no exception; this article lets me talk about one of my favorite subjects: deep-sea shipwreck microbiomes. As a topic, it sounds like someone dared me to propose an improbable place to do my work, and I blurted out shipwrecks! In reality, these are logical places to study microbial life in the deep-sea, and ideal to monitor the impacts of oil spills over time.
I had been a deep-sea microbial ecologist since 2004, studying microorgnaisms in hydrocarbon containing continental shelf sediments. How sediment microbiomes arrive in their muddy setting is a unifying theme that connects my research and bridges sediment, spill and shipwreck. Historic shipwrecks to be precise – ones that are greater than 50 years old, and protected under the National Historic Preservation Act on the outer continental shelf of the United States. Studying how the Deepwater Horizon spill of 2010 impacted historic shipwrecks in the northern Gulf of Mexico became the focus of our work, and our current study in Scientific Reports.
There are ~2000 historic shipwrecks in the Gulf of Mexico spanning 500 years of cultural heritage, and maritime and wartime history. Aside from cultural relevance, they contribute to contemporary life in the deep sea, by forming artificial reefs. These reefs are established by microorganisms when they form biofilms on surfaces. Once that occurs, a shipwreck begins a journey from maritime history to artificial reef. Our work under the Shipwreck Corrosion, Hydrocarbon Exposure, Microbiology and Archaeology project (SCHEMA) addressed how the spill impacted shipwrecks, how we could use microbiomes to monitor impacts and recovery, and how shipwrecks themselves shape seafloor microbiomes.
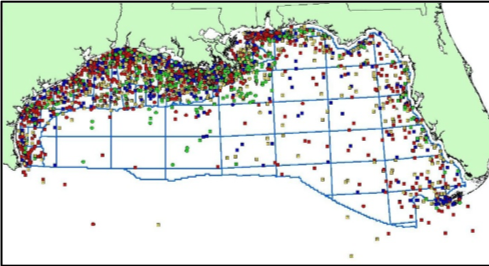
During two remotely operated vehicle (ROV) missions in 2014 on the UNOLS Research Vessel Pelican we visited seven historic shipwrecks at depths between 140m (459ft) to 1500m (4900ft). We used push corers which require a robotic arm to drive a tube into sediment to recover a column of mud approximately 20 cm deep.
Push Core collection near the Ewing Bank Shipwreck from Leila Hamdan on Vimeo. Video image obtained on the DSSI Global Explorer ROV, provided courtesy of BOEM.
The rate of sedimentation in the northern Gulf of Mexico, has been studied in the last two decades. Knowing the expected rate of sedimentation in the areas where our shipwrecks are found is ~0.09 cm/year, we assumed a 20cm deep core captures ~300 year of seafloor history, written in sediment geology, chemistry and microbiology. That is significant to the study, since our work focused on metal-hulled shipwrecks that sank during WWII (early 1940s), and wooden-hulled, copper-clad shipwrecks with origin in the 19th century.
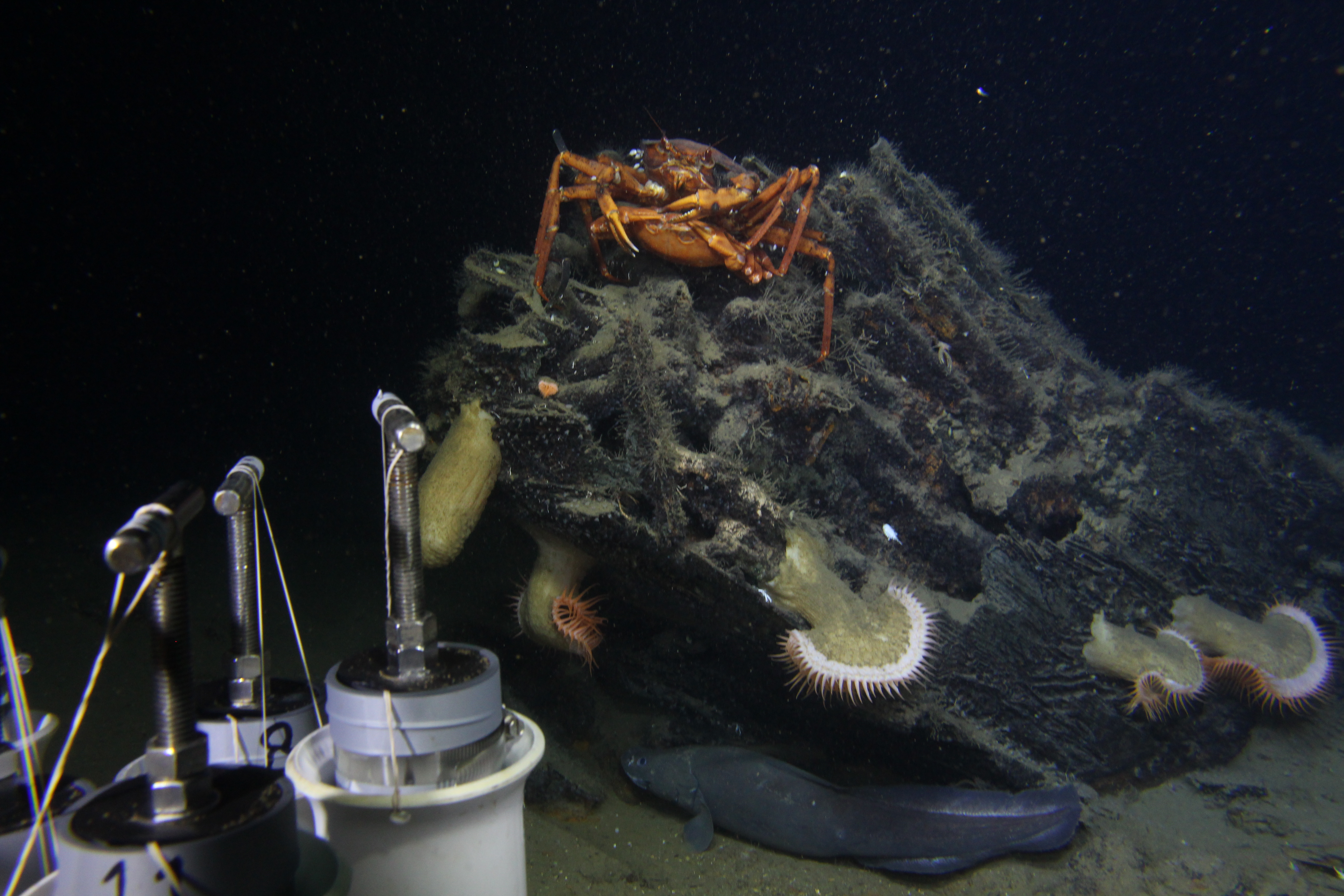
Sediment was used to compare the biological, chemical and physical conditions at heavily impacted shipwrecks near (<20km) the Macondo wellhead, the epicenter of the Deepwater Horizon spill, at moderately impacted sites 50-100km from the wellhead, and outside of the spill impact area (150km away). Sediment was collected within 2m of each shipwreck. This gave a front row view of the shipwrecks while we sampled, and conducted video, photo and sonar surveys. This provided our favorite view of the Ewing Bank shipwreck, with all of its reef inhabitants on display. At the Mardi Gras shipwreck, a wooden-hulled 19th century vessel located within the area impacted by the spill, we interrupted a private interlude between two deep-sea crabs. Sediment cores were also collected away from shipwrecks,where no debris was present (100-200m away).


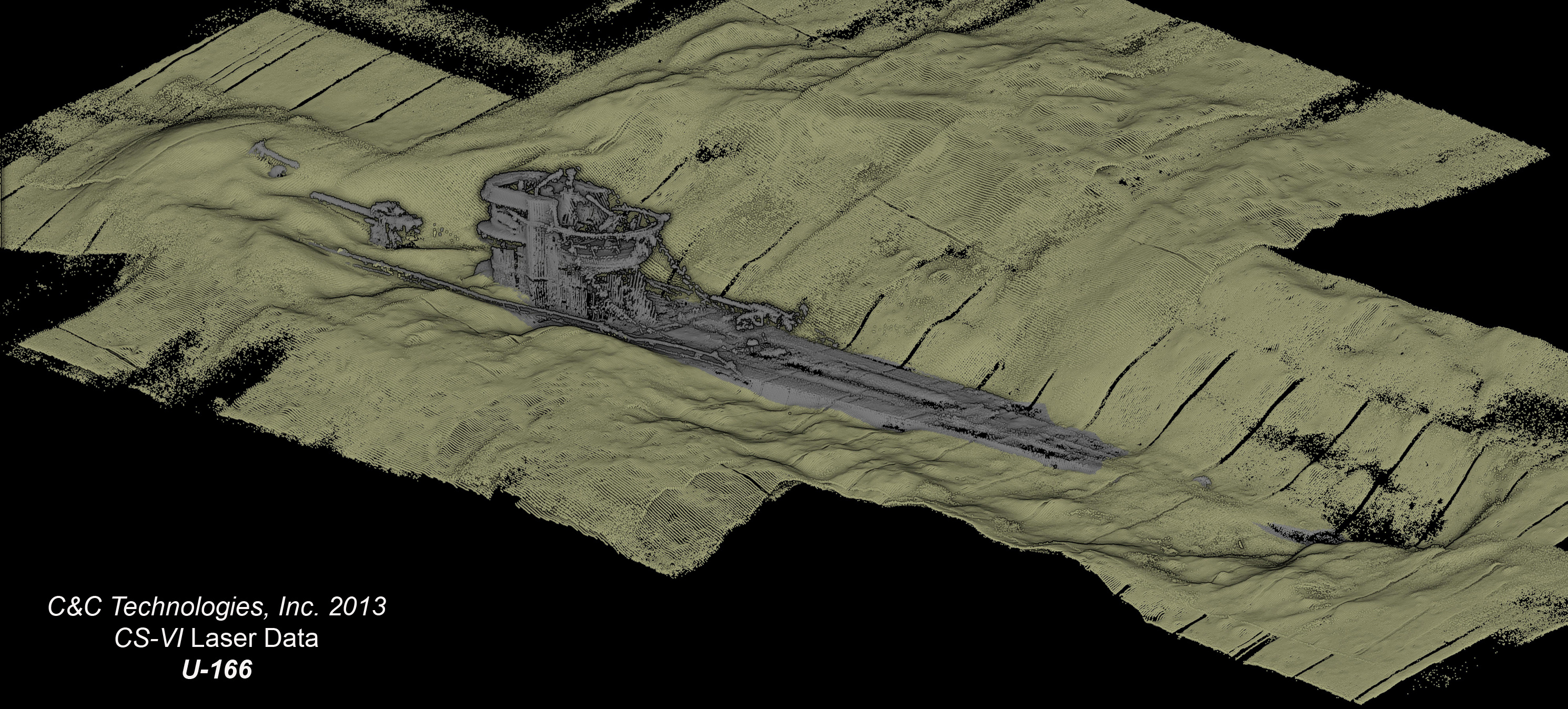
We discovered that the German submarine U-166, ~8km from the wellhead, had evidence of an oiling event. We used the Lead-210 approach to measure sediment accumulation rates. At U-166, that rate was 0.63 cm/year, about 7 times what is expected for this area of the Gulf of Mexico. Radiocarbon natural abundance at most sites showed a progression of newer sediment over older, aged material. This was not the case at U-166, where we observed aged material through the entire core. Both observations suggest a fallout event of oil snow associated with the spill. There was physical evidence for that event - the upper 6 cm of sediment had a reddish-brown color, and a slimy texture, consistent with oil snow deposition. The porosity of these sediments (above 90%) was anomalously high for a location 1500m deep.
Conning tower of U-166 from Leila Hamdan on Vimeo. Video images obtained on the DSSI Global Explorer ROV, provided courtesy of BOEM.
Surface sediments from three sites in the area heavily impacted by the spill were enriched with bacteria related to the Gammaproteobacteria, common in marine sediments. Up to 16% of those bacteria in the upper 5 cm at heavily impacted sites were dominated by one member of the Piscirickettsiaceae family, most closely related to obligate polycyclic aromatic hydrocarbon (PAH) degraders. At U-166, the Piscirickettsiaceae sequence was uniformly abundant in the thick layer of reddish brown oil snow deposits.
The study revealed that microbiome diversity is higher near shipwrecks than the surrounding seafloor, except for heavily impacted sites. This leads to two conclusions: 1) Shipwreck proximity influences seafloor microbiomes, increasing local biodiversity, and producing an island effect, similar to what as has been seen for macro-organisms, and 2) Spill contaminants on the seafloor mask the appearance of that island effect.
Our study was the first to examine the sediment microbiomes of deep-sea historic shipwrecks. That alone was exciting and provided building blocks for our next line of inquiry into how human structures shape the seafloor microbial landscape. We also conclude that the U-166, a historically significant shipwreck, war grave, and ecological resource was impacted by the Deepwater Horizon spill.
Our study involved scientists from disciplines that rarely intersect: marine archaeology and microbial ecology. This allowed us to look at the seafloor from different scales and perspective to understand the role of historic shipwrecks as deep-water ecosystems. We collaborated, and cooperated in scientific discovery, and as a result, we learned new things about the ecosystems that allow us to enjoy our science. It has been fascinating to go to archaeology conferences and share microbiome studies. I have seen the improbability of shipwreck microbiology register on the faces of archaeologists, and then enjoyed watching them accept the value of studying what is invisible, but consequential to charismatic seafloor features.

The paper in Scientific Reports can be found here: https://go.nature.com/2IBohqq

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in