The evolutionary mystery of orphan genes
Published in Ecology & Evolution
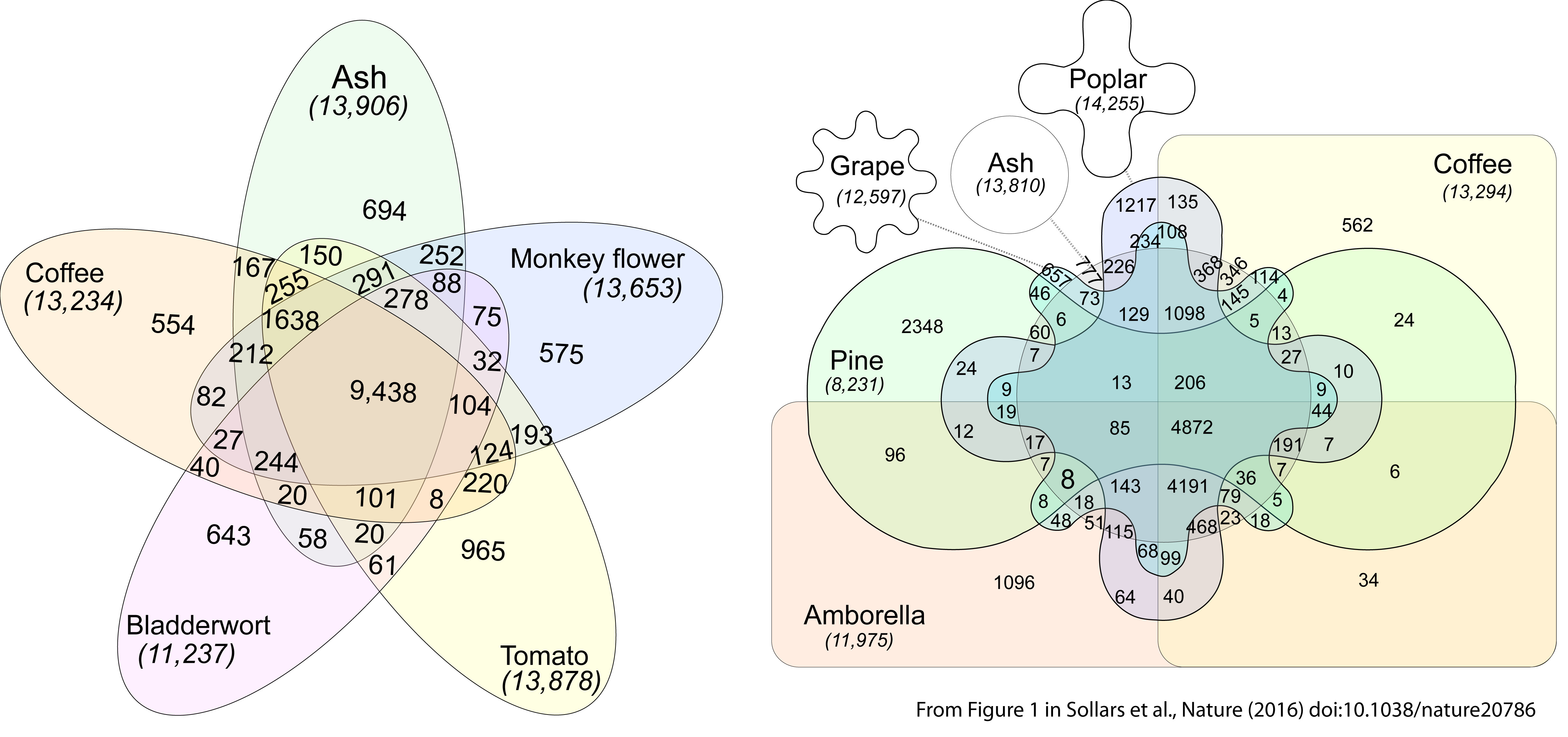
This week in Nature I and my co-authors published the ash tree genome. Within it we found 38,852 protein-coding genes. Of these one quarter (9,604) were unique to ash. On the basis of our research so far, I cannot suggest shared evolutionary ancestry for these genes with those in ten other plants we compared ash to: coffee, grape, loblolly pine, monkey flower, poplar, tomato, Amborella, Arabidopsis, barrel medic, and bladderwort. This is despite the fact that monkey flower and bladderwort are in the same taxonomic order (Lamiales) as ash.
Such genes are often known as "orphan genes" – orphans because they appear to be lacking evolutionary parents. A more precise term is "taxonomically restricted genes" as this allows us to specify the taxonomic level at which they are unique. Some of the 9,604 genes we found to be unique to ash will be unique to the species, some unique to the genus, and some to the family. On further research, some may turn out not to be orphans at all. In future we plan to conduct detailed comparisons with other ash species and with olive, which is in the same family as ash.
Orphan genes are found every time a new genome is sequenced. Their ubiquity has been one of the biggest surprises of genomics over the last 20 years. Many researchers had hypothesised that the number of orphan genes found would steadily diminish as more and more genomes were sequenced – but this is not the case. Orphan genes continue to comprise a sizeable proportion of each new genome sequenced. I and Paul Nelson reviewed this topic in a chapter of a book published this year by Cambridge University Press: "Next Generation Systematics".
Orphan genes are "the hard problem" for evolutionary genomics. Because we can't find other genes similar to them in other species, we can't build family trees for them. We cannot hypothesise their gradual evolution; instead they seem to appear out of nowhere. Various attempts have been made at explaining their origins but – as Paul and I describe in our book chapter – the problem remains unsolved.
Give their ubiquity in all genome sequences orphan genes receive comparatively little attention from the research community. I suspect this is partly because they are such a difficult problem. Science is "the art of the soluble". It may be that little funding finds its way to the origin of orphan genes because it appears to be an insoluble problem.
Popularisers and communicators of science have also had surprisingly little to say about orphan genes. This is a pity: what can be more interesting and more inspiring than an unsolved mystery? Who could choose to ignore a lost orphan?






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
The orphan gene problem is a fascinating conundrum. It is stunning that so little research into the biochemical pathways leading to these genes has been conducted. The theoretical explanations are so far merely speculations. I hope the author of this paper and his colleagues garner much funding and interest to answer the questions the orphan gene problem raises. As it stands now, the orphan gene problem is empirical evidence contesting the common descent model.