The genome and proteome of the first Nitrospira isolate: How the pandemic forced me to get creative
Published in Microbiology

I am a marine microbiologist with a passion for nitrifying microorganisms, which play an important role in the marine nitrogen cycle by oxidizing ammonia via nitrite to nitrate. Coming from Austria, a landlocked country - where working with marine microbes has some obvious challenges - I was looking forward to starting my postdoc at the University of California Santa Barbara, which is situated right next to the sea. Full of excitement to explore this new opportunity, to collect samples from the Santa Barbara Basin and immediately process them in the lab, I was faced with an unforeseen obstacle – a global pandemic.
Luckily, my postdoctoral advisor, Alyson Santoro, had accumulated a number of “orphan projects” over the past years, which were in need of “re-homing” and ready for me to work on. Among them was a genomic and proteomic dataset of the nitrite-oxidizing bacterium Nitrospira marina, which was an ideal project to work on during the lock-down since it did not require laboratory access. N. marina was isolated from a water sample collected from the Gulf of Maine at a depth of 206 meters during a research cruise in the Atlantic Ocean in 1967, and the species was officially described nineteen years later by Stanley Watson and co-authors in 1986 1.

The water sample which later led to the isolation of Nitrospira marina was collected in March 1967 onboard the research vessel Chain. © Woods Hole Oceanographic Institution
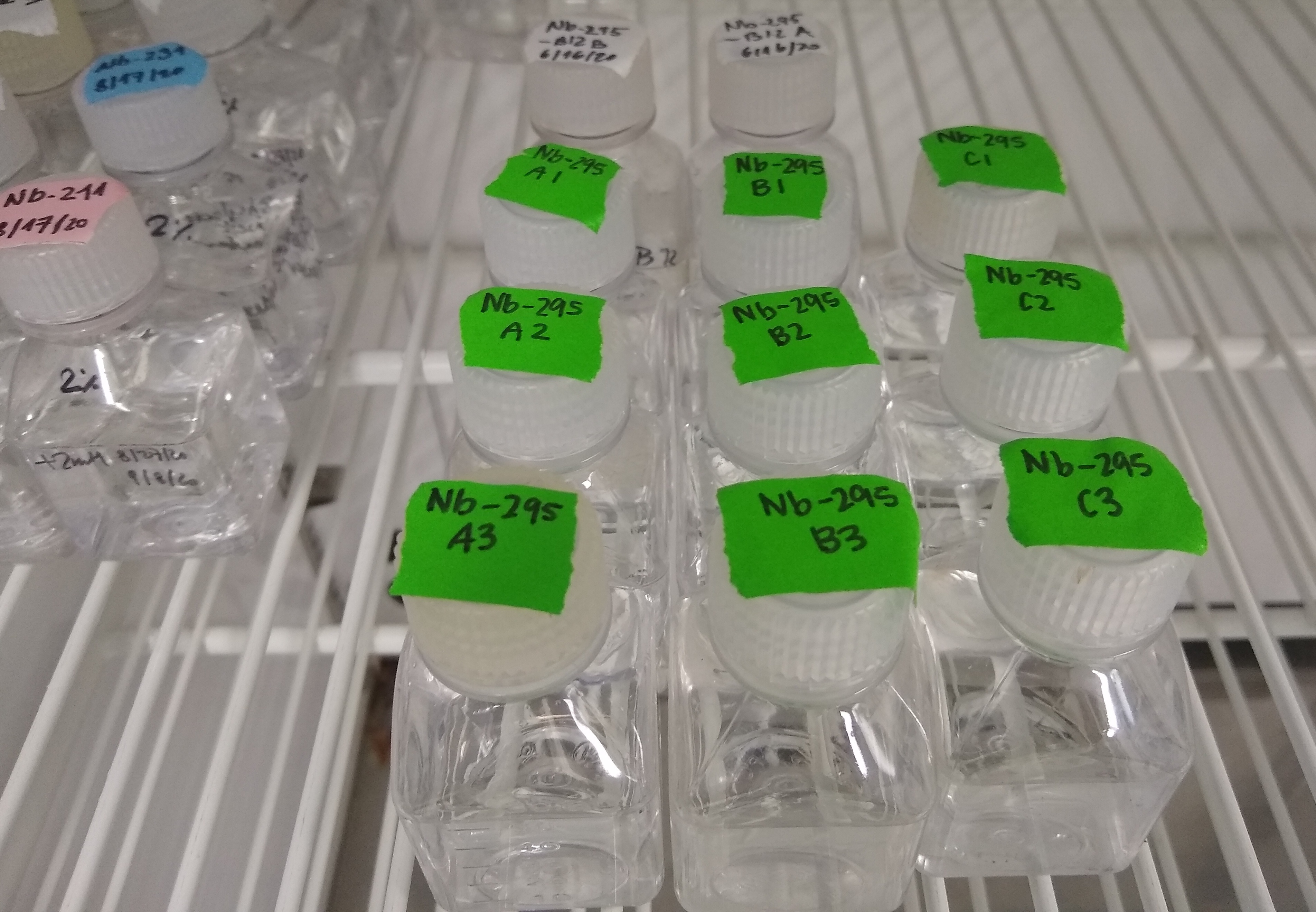
Although originally deposited to a culture collection, N. marina was not maintained in a public culture repository due to the many difficulties associated with cultivating and maintaining nitrifying microbes. Fortunately, Frederica Valois and John B. Waterbury at the Woods Hole Oceanographic Institution (WHOI) have maintained N. marina in liquid culture for the past 35 years in order to keep this important microbial resource alive and available for further research.
Since 1986, the availability of additional isolates and genomes of the genus Nitrospira has led to many new discoveries, illustrating its high metabolic flexibility beyond nitrite oxidation 2. However, despite the ubiquitous presence of this genus in many natural and engineered ecosystems, N. marina still represents the only Nitrospira isolate from the oceanic water column. I was curious to look into the genome of this special microbe to find out more about its metabolic capabilities and how it differs from other marine nitrite oxidizers. Additionally, the genomic information was used for proteome analyses through a collaboration with Mak A. Saito and his team, also at WHOI, which enabled us to look at the abundances of individual proteins in N. marina. When the lock-down was lifted, I got the opportunity to test some of the newly generated hypotheses on living cultures of N. marina in the laboratory.

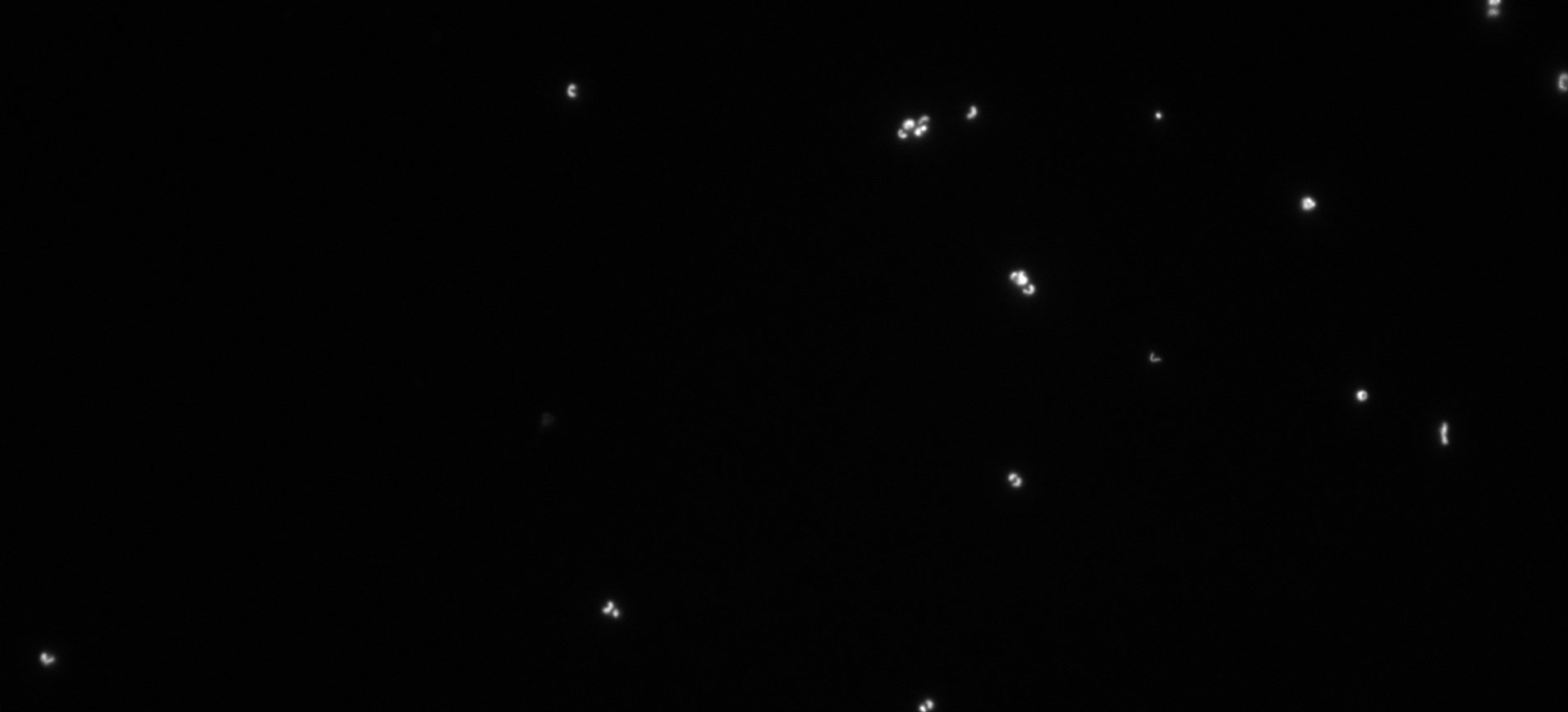
If you want to know more about what makes N. marina special, where you might be able to find it in the ocean, and how it responds to different environmental conditions, check out our paper.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
As somebody who has flipped pages of Stan Watson’s cruise and isolate log books on AOB and NOB and who understands the connection between their page numbers and the labels assigned to the original isolated, both of which are carefully preserved at WHOI, I enjoyed your story immensely! Keep up the good work!
Martin