The potential of biofilms in strain improvement
Published in Microbiology
The fermentation process has been applied for generations, aiming to reduce food spoilage and add flavor, texture etc. to the final product. The successful outcome of fermentation depends on the composition and activity of the microorganisms present. Back in time, fermentation relied on a random inoculum, which varied from batch to batch. Some products could only be produced in specific regions, such as Pajottenland in Belgium, which is still known for Lambic beers. The ability to isolate and proliferate single organisms did, however, enable the assembly of well-defined starter culture communities that warrant consistent taste, aroma, and consistency of the product. Paradoxically, despite the fact that many fermentation products require a range of different species that each concomitantly contribute to the process, the screening and improvement of starter culture candidates is conducted in a single strain manner.
In our lab we focus on community-intrinsic properties and we wondered whether starter cultures could be optimized by exposure other potential starter culture partner species in the optimization stage. If strains had already adapted to the presence of one another, they would perhaps perform better and potentially enable the production of new products? Inspired by a previous study, conducted in our group, where we showed that bacteria adapt differently when in the presence of other species, we aimed to test whether evolution of industrially relevant bacteria was impacted by interspecies interactions. We used two routinely used species in dairy production as a model system; Lactococcus lactis and Leuconostoc mesenteroides. These represent common dairy starter culture strains that are routinely refined using non-GMO methods i.e., experimental and adaptive evolution, and hence new knowledge could impact the current paradigm.

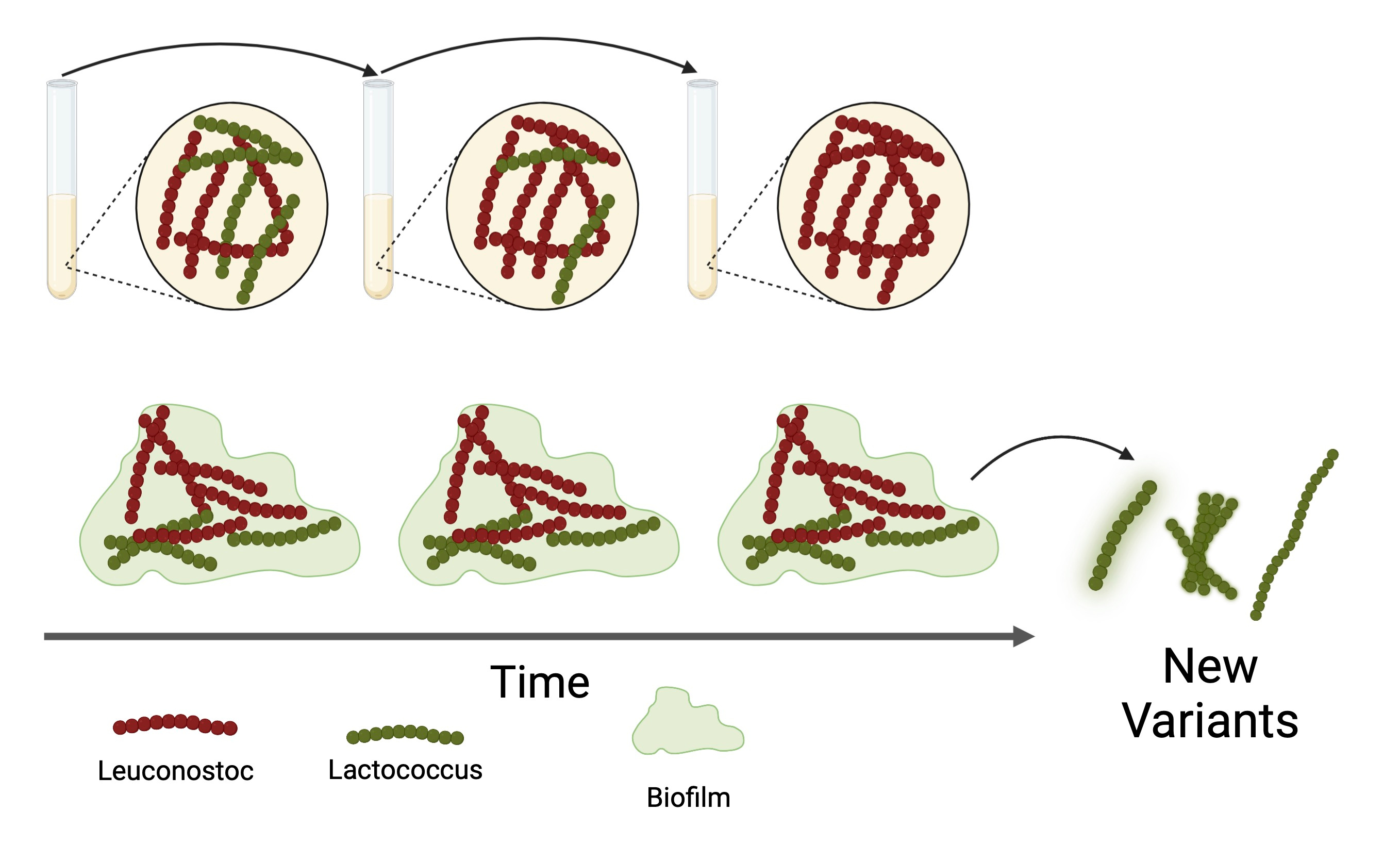
A common hurdle met when studying multispecies communities over time is that a single species often dominates due to competitive exclusion. The same was observed for our two bacteria when grown in liquid cultures. The simple addition of a glass surface facilitating bacterial attachment did, however, ensure stable coexistence. This is likely because cells now formed spatially structured biofilms on the glass beads, providing environmental heterogeneity and hence distinct niches for the two strains to occupy. With the ability to grow the species together, L. lactis was grown on glass beads with and without the presence of L. mesenteroides (Fig 1). After two weeks, 83 colonies were isolated and analyzed. Interestingly, variants isolated from the co-cultures grew significantly slower but reached higher biomass densities compared to the ancestor, which the mono-culture variants did not (Fig. 2AB). Both mono- and co-culture-grown variants formed more biofilm than the ancestor, with the co-culture variants forming more than the mono-culture variants (Fig. 2C).
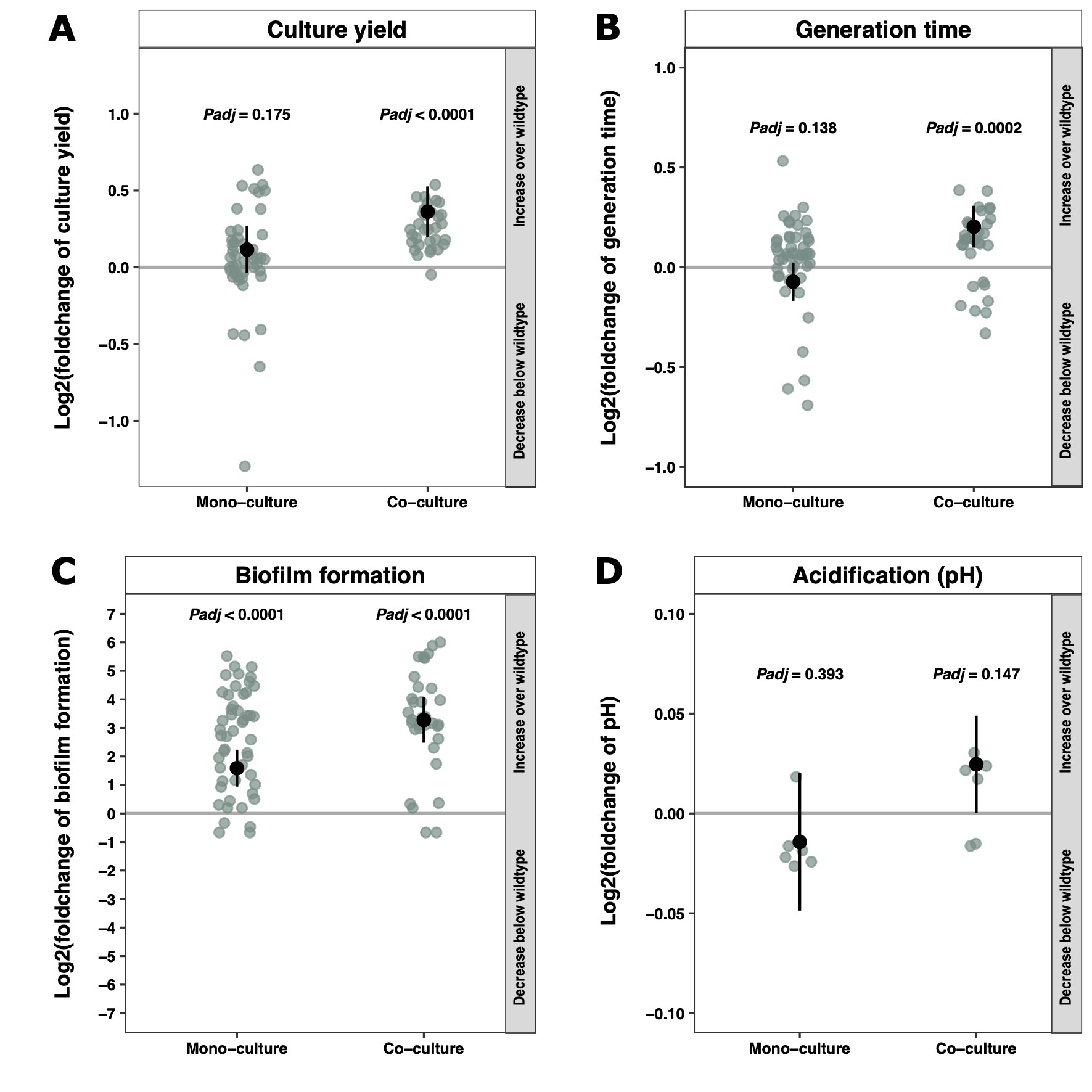
Among the isolates, variation in the colony morphology was observed, and some tended to entangle and sink to the bottom of cultures. With the use of mixed-effect linear models it was possible to dissect the effect of the morphotype on the tested phenotypes. It turned out that morphology was a significant parameter impacting generation time and biofilm formation.
Whole genome sequencing of selected variants did not reveal conserved genotypic adaptions associated with mono- or co-culture originating variants. However, variants from co-culture biofilms contained mutations in genes related to growth, such as genes encoding an ATP synthase or a DNA translocase. Also, a distinct, smaller, and darker colony morphology was associated with mutations in a gene encoding a Helix-turn-Helix transcriptional regulator.
Our study shows that small changes to the environment, selecting for biofilm formation, maintain and stabilize diversity. By these means, biofilms can function as evolutionary incubators and provide the possibility to study the coexistence of species that would otherwise outcompete one another.
The present findings exemplify the power of considering ecological principles in industrial bioproduction, and if you want to know more about the study, you can find the article here.
Mads Frederik Hansen, Mette Burmølle & Nathalie Nina Suhr Eiris Henriksen
Follow the Topic
-
npj Biofilms and Microbiomes
The aim of this journal is to serve as a comprehensive platform to promote biofilms and microbiomes research across a wide spectrum of scientific disciplines.
Related Collections
With Collections, you can get published faster and increase your visibility.
Microbial endocrinology
Publishing Model: Open Access
Deadline: Oct 21, 2026
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in