The story of an analytical pipeline and the Atacama Desert
Published in Microbiology

The paper in Microbiome is here: go.nature.com/2P1ZWy3
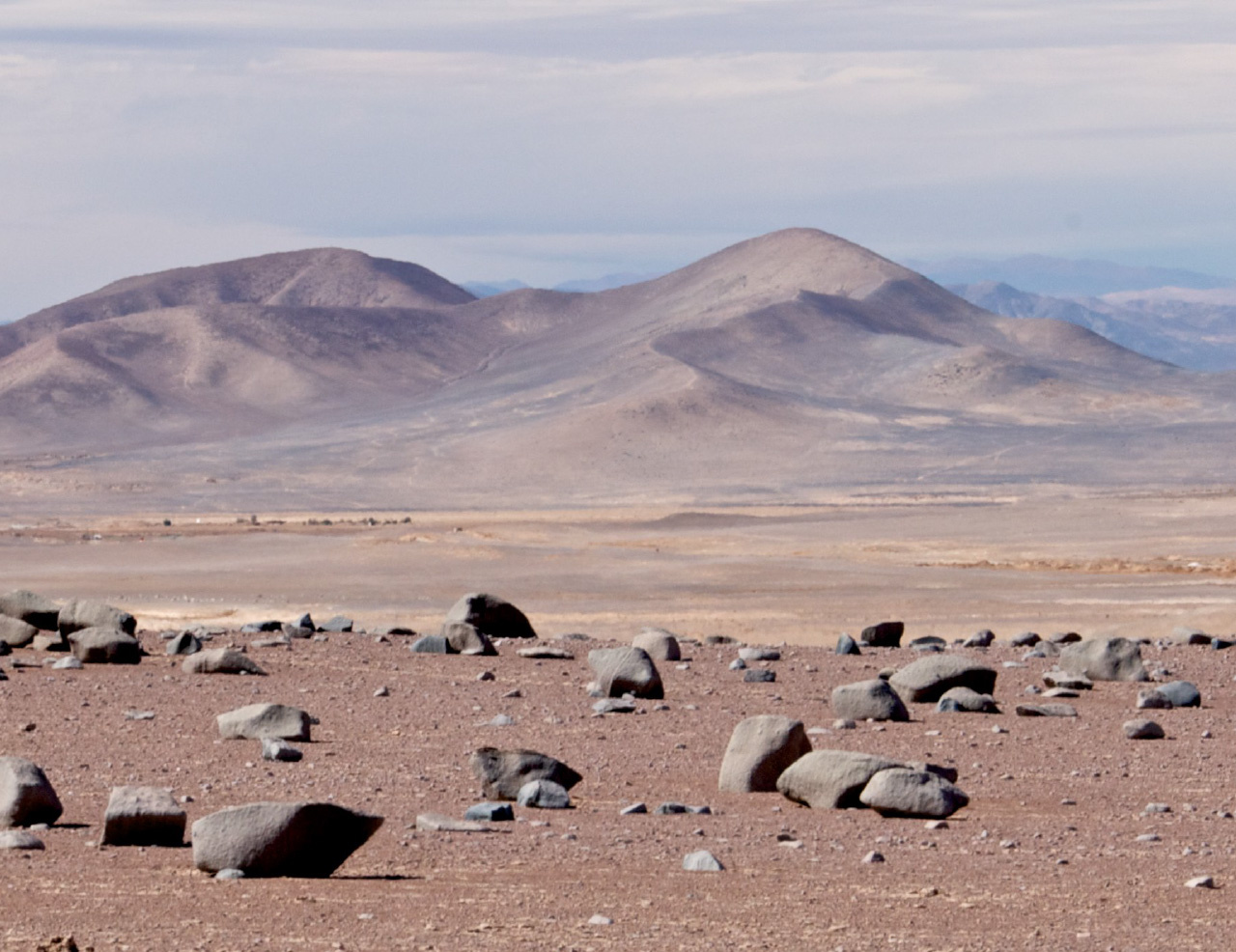
Gherman Uritskiy, a graduate student at Johns Hopkins University and the lead author of the paper, has been investigating the diversity and functional dynamics of halite microbiomes in the Atacama Desert, Chile. As such, Gherman and one of his graduate advisors, Dr. Jocelyne DiRuggiero, went on several field expeditions to the desert (Figure 1).
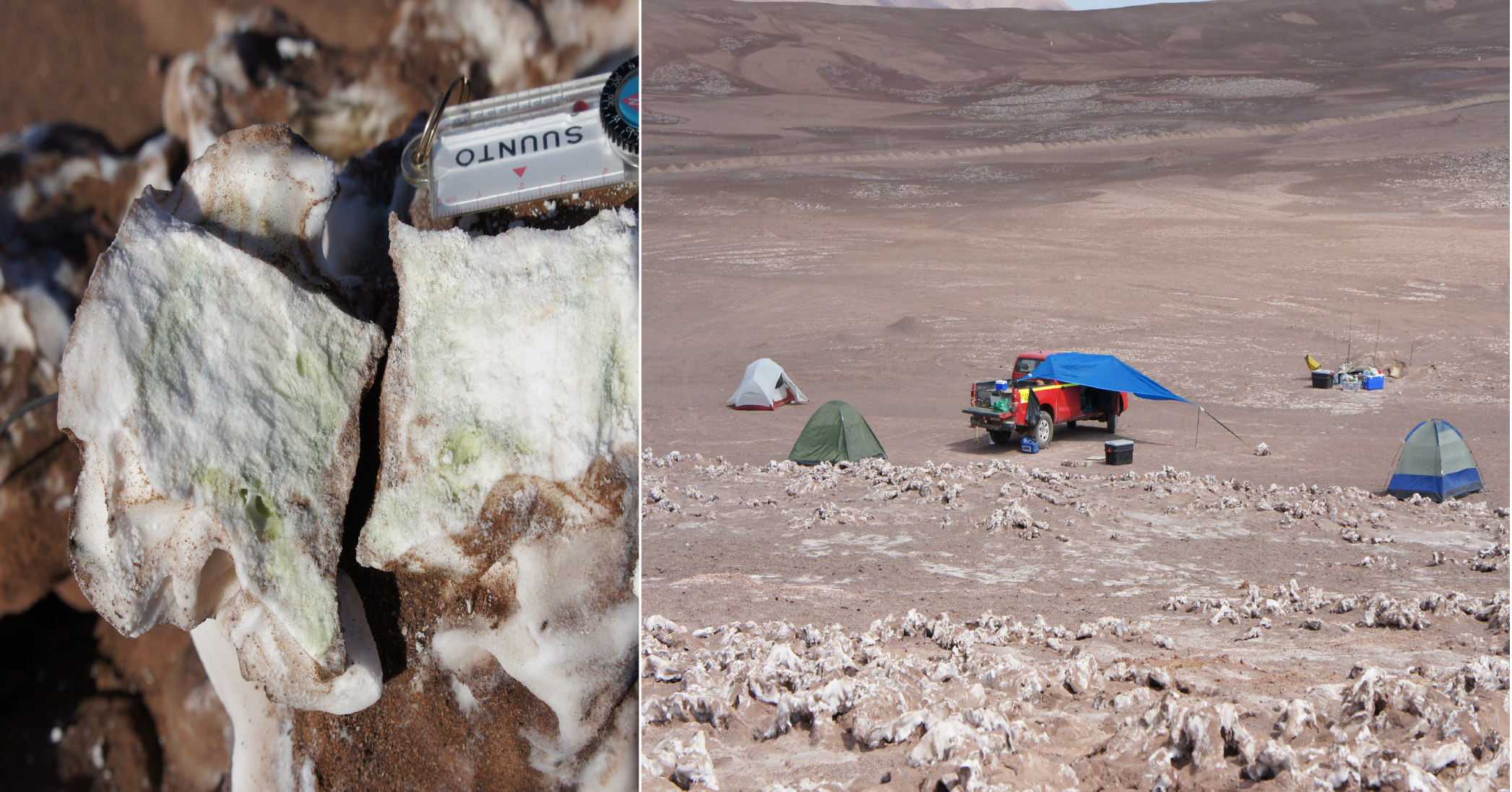

Fig. 1. Left: section of a halite nodule with a green colonization zone indicating the presence of photosynthetic microorganisms. Right: Atacama Desert fieldwork with our camp and the lab on the bed of a truck.
A large number of samples were collected and processed, resulting in a considerable amount of metagenomics data. Gherman learnt to analyzed the data with the support of his co-advisor, Dr. James Taylor, but found that the learning curve for metagenomic data analysis was rather steep and that there was a distinct lack of a unifying platform to perform these analyses. By the end of the year, Gherman’s analytical pipeline had evolved into a complex web of interdependent software and custom programs. Seeing the promising results of the pipeline, we thought that many others in the microbial ecology field would benefit if such wrapper software became widely accessible - hence, metaWRAP was born. MetaWRAP came about because of necessity and, with a lot of support and help from the community, the pipeline continues to evolve and improve. New modules and features are being added every month based on requests from metaWRAP users. To date, the pipeline has been tremendously successful, with hundreds of unique users from around the world (Figure 2), and over 1000 views on its GitHub page each week (https://github.com/bxlab/metaWRAP).
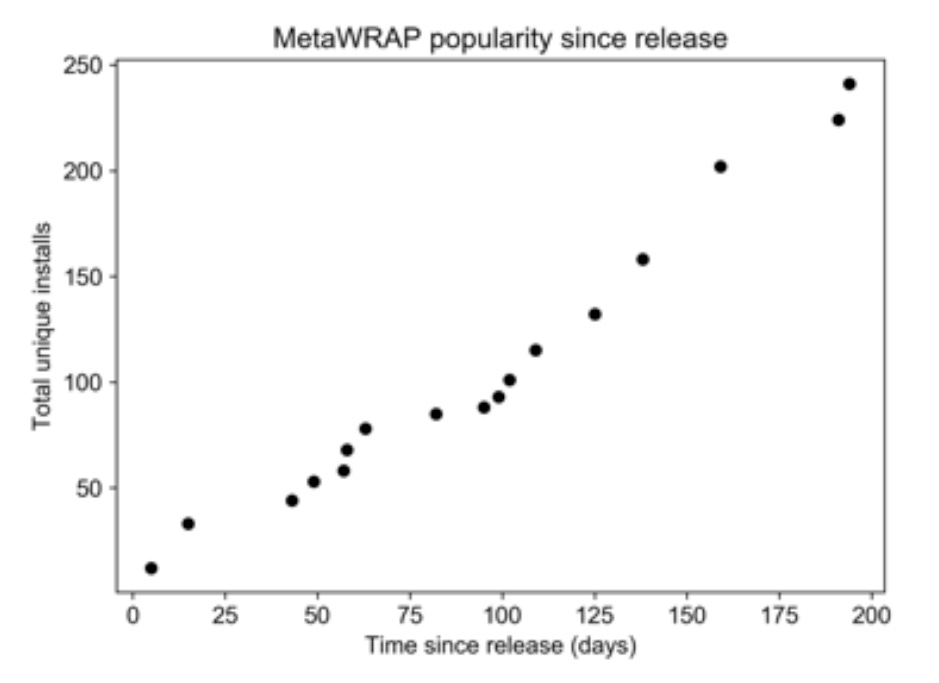
Fig. 2: Total installs of metaWRAP over time
Now, coming back to those microbial communities inhabiting halite nodules, we have now completed the study and are preparing to submit a manuscript for review. MetaWRAP allowed us to investigate this complex community at a level previously unattainable, yielding some fascinating results – it is a very exciting story, so stay posted!
And for those who wonder, no there is no shower and no bathroom where we go in the field. We carry with us enough water and food for 4 to 5 days at a time and rough it out in one of the most beautiful places on Earth.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Quite useful tool and it saves a lot of time installing so many softwares....Just read your newly published ISME J. Resilience of extremifiles under disturbance: https://www.nature.com/articles/s41396-019-0468-y