Think simple
Published in Bioengineering & Biotechnology

To address the novel coronavirus SARS-CoV-2 that causes COVID-19, many types of assays are needed. There are PCR-based tests, there are immunoassays. And there have been robustness and other types of challenges with some assays as I reported in this story.
In a press conference today, Michael Ryan, executive director of the World Health Organization's Health Emergencies Programme pointed out that it can appear confusing now that a variety of tests are becoming available, including antibody tests and PCR-based tests. “Countries are grappling with what is the best choice for them,” said Ryan. PCR-based tests directly detect the virus and “are probably the most accurate way to determine whether someone is currently infected.”
Serology tests that detect antibodies and look at an immune response, for example the immunoglobulins IgM and IgG, detect if one has been infected recently or in the past, and it takes days or weeks to develop such a detectable response, he said.
Governments need to currently focus specifically on PCR-based tests or assays that detect active infection. But serologic tests “are extremely important for determining how many people have been infected in the population,” said Ryan. “Those data are very important to tell us where we are going in this epidemic.”
There are indeed hundreds of test-types available, also because the viral genome was sequenced and shared quickly, a number of labs could develop PCR-based kits. Maria van Kerkhove technical lead of the WHO Health Emergencies Programme said that she and her colleagues are working with collaborating labs and the non-profit Foundation for Innovative New Diagnostics (FIND) to evaluate these many assays “so we can see how well they perform when actually being tested on clinical samples.”
FIND is a WHO “Collaborating Centre for Laboratory Strengthening and Diagnostic Technology Evaluation.” It is headquartered in Geneva with hubs in India, Kenya, South Africa and Vietnam. The list of all molecular tests currently being assessed by FIND, 242 assays, is here.
The same process will be undertaken with serologic assays, says van Kerkhove.
The technical guidance on serologic-assays issued on 8 April is here.
“With the limited data now available, WHO does not currently recommend the use of antigen-detecting rapid diagnostic tests for patient care, although research into their performance and potential diagnostic utility is highly encouraged.”…
…”Based on current data, WHO does not recommend the use of antibody-detecting rapid diagnostic tests for patient care but encourages the continuation of work to establish their usefulness in disease surveillance and epidemiologic research.”
There is a lot on the market, van Kerkhove said, “this is a positive thing.” She and her colleagues want to assure these tests are evaluated and validated across multiple labs so that their performance in actual use can be determined.
As the palette of assay types grows, labs will find that some require quite specific hardware and in some cases supplies, such as reagents or the assays themselves, are not in ample supply for the pressing needs of a pandemic. That’s why a few labs are exploring a different tack: a simplified design for both PCR-based tests and immunoassays.
Paper-based assay
A research team at Purdue University has developed a paper-based immunoassay with which labs can detect nucleic acid amplicons. They published a paper about it in ACS Omega here. The assay uses dried reagents. “My lab specializes in taking assays from needing fancy equipment to being integrated into handheld detection devices,” says Purdue researcher Jacqueline Linnes. She and her team have developed a device for MERS-CoV. It uses dried solutions of hydrogen peroxide, diaminobenzidine and horseradish signal enhancers. She and her team would like to adapt this for COVID-19 and “ideally would be able to make it into a screening test that could be used for many different viruses,” she says.
Lateral flow assays, which are single channel microfluidic devices that use paper, are low-cost and easy to use but they run into issues: the delivered signal can be weak. To avoid that, a user would need to pipette different fluids into the assay at quite specific time-points, which is hard to do. For more complex types of reactions, labs have developed two-dimensional paper networks with several fluidic inputs and several channels. New types of chemical ligation assays can be used to amplify nucleic acid amplified needing enzymes. There are also CRISPR-based assays that follow this approach. But yields in these assays can be low when compared to PCR-based assays.
All of this led Linnes and her team to build an enzyme-linked immunosorbent assay (ELISA) into a 2D paper network for detecting nucleic acid amplification products. The chemistries, says Linnes, were developed by her colleagues at a company, CrossLife Technologies.
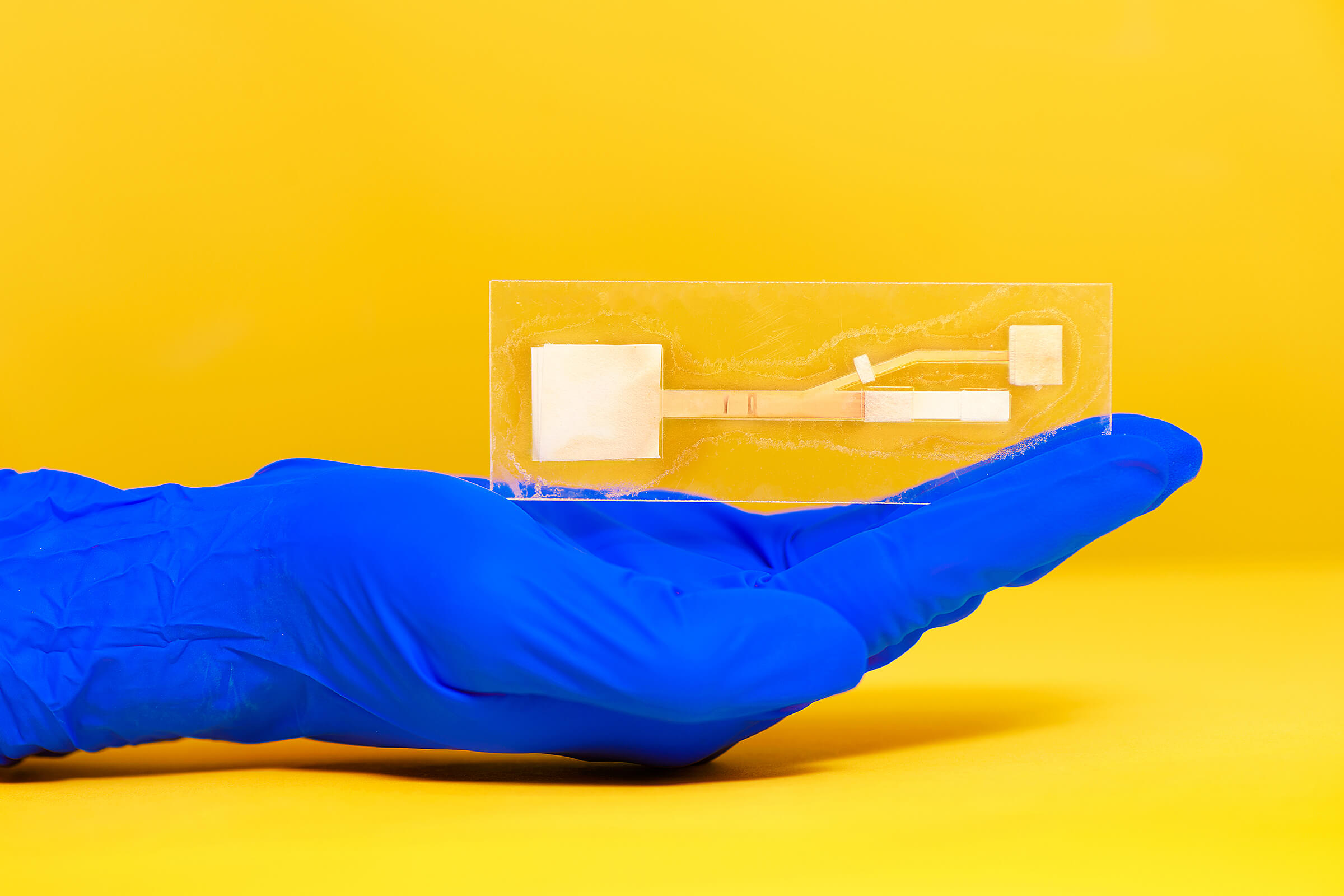

An assay made of paper developed at the Purdue University lab of Jacqueline Linnes. The lab would like to ready it for applications related to SARS-CoV-2 and other viruses, she says. (Credit R. McElhoe, Purdue University)
To use the assay, a scientist adds sample, which could be a swab from a person, to a tube with lysis buffer to break open the virus. The template assisted rapid amplification (TARA) assay uses the virus’s RNA to bring together two different synthetic nucleic acid strands that each match to side-by-side sections of the RNA sequence. One sequence carries a biotin tag and the other a fluorescein tag.
When the strands move close to one another, a transferase reaction occurs: the tag on one strand is transferred to the other strand. And the other strand then has both tags. “TARA is great because it can be done at room temperature within 15 minutes,” says Linnes. However, it is not super-efficient at amplifying the signal. The assay is not as sensitive as PCR or LAMP or other enzymatic tests, she says, referring to loop-mediated isothermal amplification-based assays.
Her lab worked with CrossLife Technologies to increase the signal intensity of the test on paper-based immunoassays, of which pregnancy tests are one example. The team developed a method that handles the sequences as follows: the sequence with the biotin tag is bound to a streptavidin-coated gold nanoparticle and flows down the paper strip.
The other end of the nucleic acid with the FAM binds to the anti-FAM antibody in the test line. “Extra streptavidin-coated nanoparticles get bound by a second control line with anti-streptavidin antibodies to prove that the test runs properly,” she says.
But without the other enzymes and reagents in the test, the readout-line is so faint that it can’t be seen with the naked eye, says Linnes, which is why they add not only streptavidin but also horseradish peroxidase (HRP) to the nanoparticles
With HRP, one can flow a substrate, diaminobenzidine (DAB) in hydrogen peroxide, past the test and control lines. In so doing the DAB is converted from colorless to a dark brown precipitate that gets caught in the membrane at the site of the bound nanoparticles. “When you do this, the test and control lines get much darker and it is possible to improve the signal intensity of the tests so that they’re visible by eye,” says Linnes. The team purchased DNA strands that already had both biotin and FAM attached so the team knew the exact concentrations.
By packaging all of these different chemistries as dried reagents on one side of the card, and the flow strips on the other side, a user can add the TARA samples and water to the reagent side and then fold the card to run the detection. “This way, you don’t have to do multiple steps like adding the sample, waiting 5 minutes, then adding the rinse, waiting 10 more minutes, then adding the H202 before you can see the results,” she says.
Building this took a lot longer than expected, she says. It worked great with liquid reagents and then they had to start over with dried reagents given the different flow rates. Anna Bird, now at University of Cambridge, modeled the flow of the various reagents in the material, says Linnes. They found it hard to dry the hydrogen peroxide, a volatile chemical that evaporates when wet. K Byers in the lab figured out that they could just crush up tablets of urea-H2O2 that are used in ELISA kits and stick them to the adhesive of the test card.
Before this is ready for commercialization, the assay and readout combination need a bit more work, says Linnes. The lab is working on ways to enhance the signal, refine the detection limit of the device, which can now only detect high concentrations of the virus.
For a sensitive test, a very low limit of detection is needed to also capture all concentrations, also so that people with little or no symptoms could be assessed. “Additional next steps are also to integrate then entire TARA assay onto the paper card so that we can make it a truly one-step test,” says Linnes.
With improvements and by adapting the methods for SARS-CoV-2, it would be possible to use as a point-of-care test so that when someone visits the doctor or takes part in a drive-through test, she says. People could get the results within an hour.
No purification
At the University of Oxford, chemical engineer Zhanfeng Cui and Wei Huang developed a rapid test for detecting SARS-Co-V-2 viral RNA and RNA fragments from swab samples. It’s basically a cell-lysate based PCR test with primers, says Cui, but he did not offer further detail. No instrument is needed for this assay beyond a heatblock that maintains a constant temperature for RNA reverse transcription and DNA amplification. The results can be seen with the naked eye.
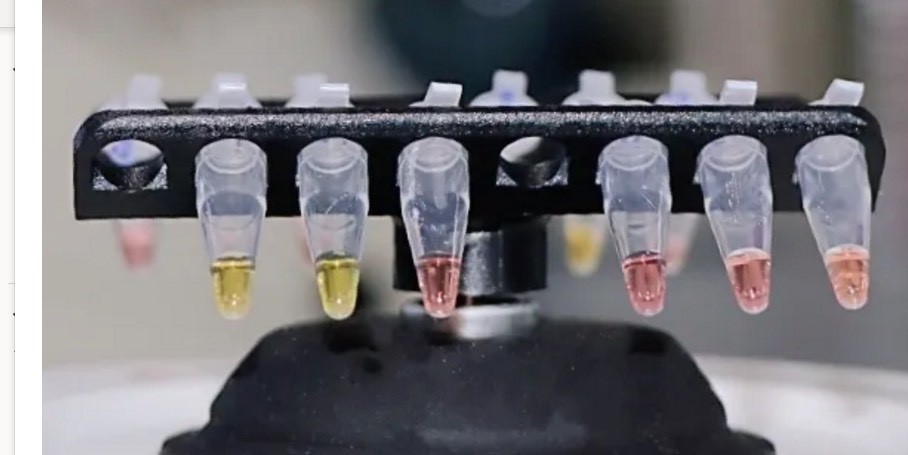

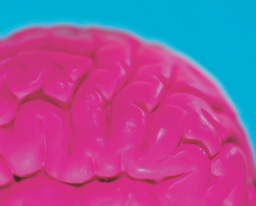
A color change indicates presence or absence of the virus in this assay developed at Oxford University. Each test uses three vials that contain different primers. According to the the university's information, a change from pink to yellow is a positive result. A positive test turns two vials yellow and leave one pink, which functions as a negative control. (Credit: University of Oxford)
The assay has internal controls to show false positive and negative readouts and the team also addressed issues of transport and storage. “We have done limited number of validation using clinical RNA samples,” says Cui. “A parallel test with RT-PCR would be the ‘standard.’” The clinical samples were from Shenzhen Luohou People’s Hospital in China where the assay has been applied to some patient samples.
The team is working on developing an integrated test, potentially for rural clinics or home use. The project involves a University of Oxford research center in Suzhou Industrial Park.
Commenting on this work, Linnes says she is intrigued and would be keen to see more detail, “but I am impressed that they performed the tests on real patient samples. Another plus is that the readout can be detected by eye. If the lab can ensure reagents can be stored at room temperature., such as by drying them, then they could be distributed to sites that need them.
She and her team are working on other, more sensitive, tests using enzymatic nucleic acid amplification for detection. They have not yet applied their paper-based assay to COVID-19 but have used it to test for HIV in blood samples.
A team at FDA’s Center for Drug Evaluation and Research developed and published, in Scientific Reports, a simplified way to perform RT-qPCR with cell lysates and avoids RNA purification. They first used a commercial reagent and noted that despite the appeal of simplicity afforded by these commercial cell-lysis reagents, the cost can be problematic. So they tried a simple buffer with a non-ionic detergent and found it worked well for generating cell lysates for their RT-qPCR influenza test.
“It's a good idea if it can be made to work, we'd like to do that with sequencing if possible,” says Nicholas Loman, a researcher at the University of Birmingham. “We routinely use crude cell lysates for gene expression studies,” says Jo Vandesompele, a researcher at Ghent University. He and his colleagues have published a paper on gene expression quantification using RT-qPCR with cell lysates in Scientific Reports, here. "When validated, it is not a problem to use such a sample prep method.” What’s important, he says is to test the methods on the intended biomaterial. If for example if the method was validated only for blood, then it cannot be used for example on nasal swabs. “It has to be demonstrated "fit-for-purpose” on nasal swabs,” he says.
Alongside his academic position, Vandesompele is chief scientific officer and co-founder of Biogazelle. His company, along with J&J, GSK ThermoFisher and in collaboration with clinical labs and universities is responding to a request by the Belgian government to significantly increase PCR- testing capacity for the novel coronavirus in Belgium.
PCR, smaller
There are a number of miniaturized PCR instruments on the market and some instrument makers and academic labs are pushing to make these available for testing for COVID-19. I wrote about some of the instruments here in a story called 'PCR heads into the field'.
And thank you for the heads up, @coregenomics, on this subject. Speaking of nudges…
DnaNudge says it is working with the UK National Health Service to trial what it calls a Nudge Cartridge for rapid COVID-19 testing.
The company has developed a DNA-focused ‘wearable.’ A user takes a cheek swab, inserts it to a DnaCartridge and placed into what it called its NudgeBox where DNA is extracted and a SNP related to “nutrition-related health conditions” is analyzed.
An hour later, the results are uploaded to an app as an iDNA report and the clinical material itself is destroyed to maintain “privacy and confidentiality.” A user can scan groceries and a flashing red or green light will “tell you if it it’s a good match for your biology.”
Justin O’Grady from the University of East Anglia and The Quadram Institute, a network of multiple institutions, and in collaboration with Jonathan Edgeworth at St Thomas have used a portable qPCR machine from Ubiquitome to do a CoV test in 50 minutes. The rapid cycle RT-qPCR handles 16 samples per run. The readout is through a phone app. O’Grady says it was a three-minute extraction process using Arcis Biotechnology’s platform.
O’Grady posted his trial run and the Twitter discussion about this method is here
Amplyus, a startup founded by molecular biologist Ezequiel Alvarez Saavedra, developed miniPCR, which can run with current from an electrical outlet or on batteries.
As miniPCR co-founder Sebastian Kraves explains, the company itself does not work on diagnostics itself but “several groups are looking to use our gear for field-based testing for SARS-CoV-2.”
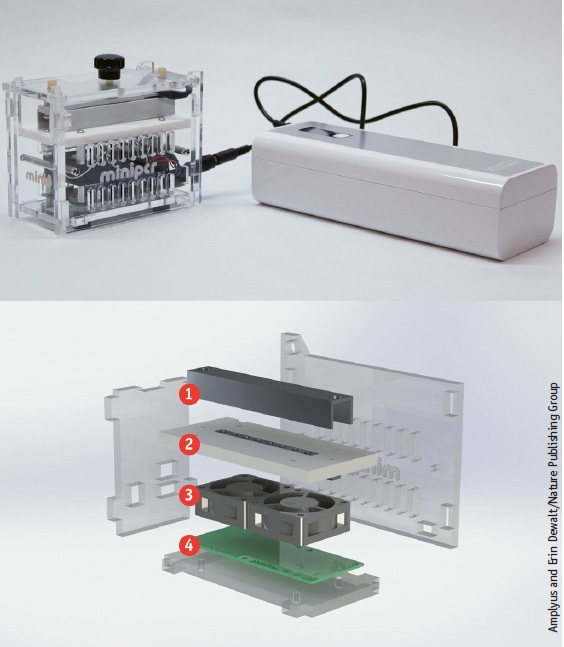

Amplyus designed and built miniPCR. Here, the layers show the heating lid (1), heating block for eight DNA samples (2), high-speed cooling fan (3) and open-source electronics (4).
For example, says Kraves, a team in Singapore has submitted a protocol for regulatory approval that applies the system with intended deployment targeting remote towns in Southeast Asia.
At Smith College Steven Williams, who developed Backpack PCR for detecting Brugia parasites in mosquitoes that cause the parasitic disease lymphatic filariasis, is working to combine the miniPCR system with lateral flow detection
A University of California, Santa Barbara team is using the miniPCR system to validate Cas13-mediated COVID-diagnostics for deployment in the field in Latin America.
Groups are refining colorimetric LAMP detection of SARS-CoV-2 with clinical samples, such as a team at the Wuhan Institute of Virology, the University of the Chinese Academy of Science and New England Biolabs. They published their work on medRxiv here. They tested and validated five sets of LAMP primers targeting two fragments of the SARS-CoV-2 genome.
A team at Centro de Biotecnología-FEMSA, Tecnologico de Monterrey in Nuévo Leon, Mexico has used the miniPCR to amplify samples combined with a DNA intercalating reagent to read fluorescence after amplification with a commercial 96-well plate reader. They published their work on medRxiv here. They used primers directed at three different regions of the genome encoding for the N protein of SARS-CoV-2.
As the team reports, the scientists found that the amplification proceeds with sufficient quality to allow proper visualization of the amplification products in electrophoresis gels, even at low concentrations of the nucleic acid. They used the miniPCR’s electrophoretic unit which is not only portable it its size limits reagent usage, they needed 15 mL agarose gel and 25 mL buffer). The team also liked the built-in power supply provided enough power for visualization of band separation in real time, which shortens the electrophoretic time by up to five minutes.
In their experience, amplification proceeds with sufficient quality to allow proper visualization of the amplification products in electrophoresis gels, even at low concentrations of the nucleic acid. They used the miniPCR’s electrophoretic unit, which is portable and limited in reagent usage: they needed 15 mL agarose gel and 25 mL buffer.
The team also liked the built-in power supply provided enough power for visualization of band separation in real time, which shortens the electrophoretic time by up to five minutes. The exposure to ethidium bromide and UV light is completely avoided by using of GelGreen dye and detection with blue light.
Band separation involved a typical processing time of 35 minutes to 60 minutes from the loading of the amplification product to final documentation of the result. They used a 96-well plate reader to quantitatively assess the amplification product by applying an intercalating agent during amplification in the miniPCR instrument. They used a 96-well plate reader to quantitatively assess the amplification product by applying an intercalating agent during amplification in the miniPCR instrument.
Overall, they note that the accuracy and simplicity of this diagnostics strategy may provide a cost-efficient and reliable alternative for COVID-19 pandemic testing, particularly in underdeveloped regions where RT-QPCR instrument availability may be limited.”
Startup GNA Biosolutions in Germany is developing a portable PCR platform that uses pulse-controlled amplification. It’s intended to be a portable COVID-19 testing platform that will be released this spring, says Anastasia Liapis, who handles partnering and marketing for the company, a resident in IZB Planegg-Martinsried. That's a startup incubator near some Max Planck Institutes and Ludwig Maximilian University.
Instead of a classic thermocycler, the instrument amplifies nucleic acids with microcyclers in which microsecond-timed electrical pulses guide the PCR-cycles of heating and cooling.
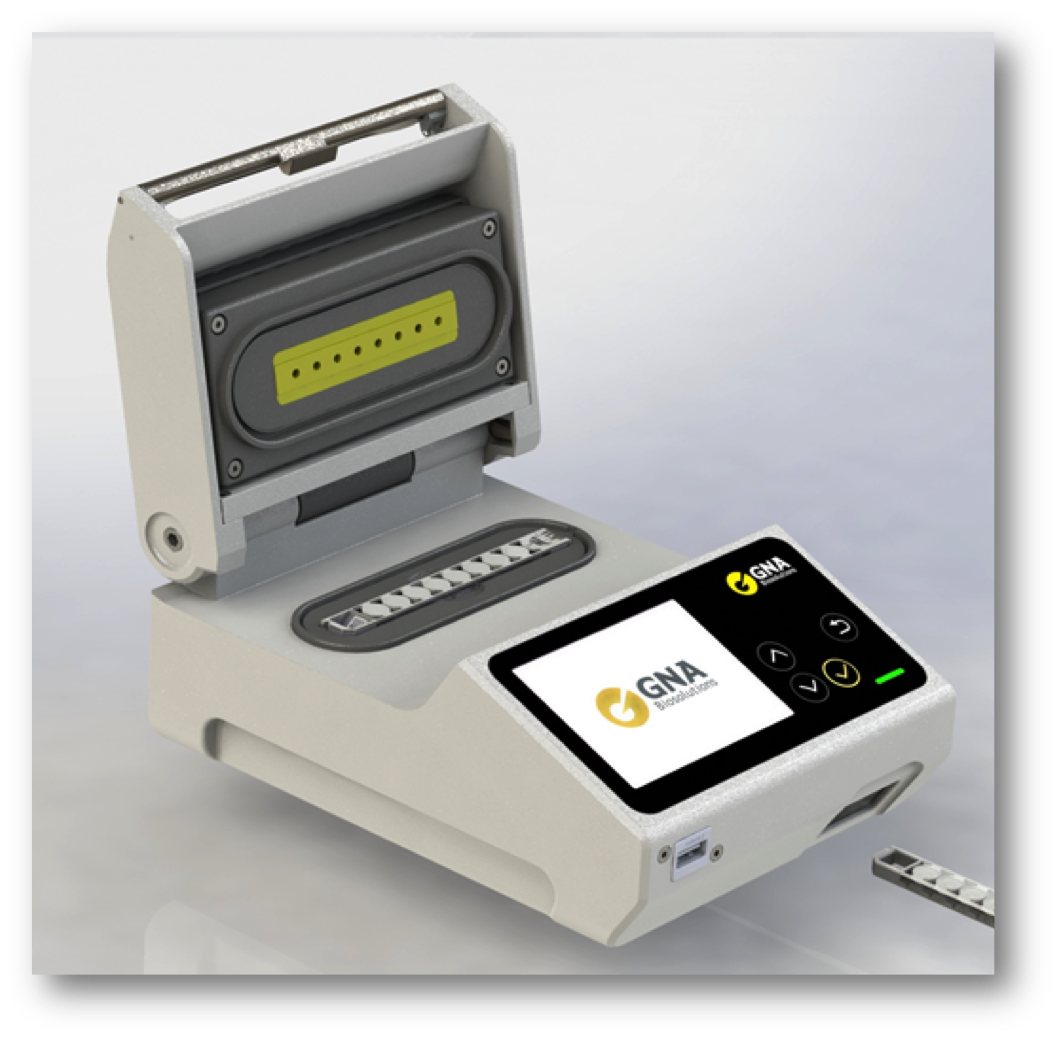

Startup GNA Biosolutions has developed a portable system that applies pulse-controlled amplification and is intended for SARS-CoV-2 detection. (Credit: GNA Biosolutions)
The instrument generates results in around 15 minutes, says Liapis. The approach can be scaled and the instrument can also be battery-operated. Among the company’s investors are Bosch, and a number of investment banks and venture capital firms, such as Mey Capital Matrix, btov and Greybird Ventures.
Other small PCR systems include OpenPCR's build-it-yourself thermocycler.
Biomeme’s mobile qPCR thermocycler is for performing PCR, RT-PCR, qPCR and isothermal tests. One hand-held device is called Franklin. It's battery-operated and can perform real-time detection of up to 27 targets. The instrument works with freeze-dried consumables that the company developed.
From BentoLab comes a platform that is a portable centrifuge, PCR and gel set-up.
Azura Genomics has developed the MyGo Mini S Real-time PCR instrument.
Finding reagents and consumables
One issue that affects both PCR-based assays as well as immunoassays, is the need for authorized use in each country if it is to be applied to patient samples and for, ultimately, diagnostic purposes. In the US that authorization comes from the Food and Drug Administration. “We’re only going to use a test that has emergency use authorization, we’re going to use tests that fit in with our workflow and for which we have a platform to run the tests” says Jennifer Rakeman, who heads the New York City Public Health Laboratory. The FDA is now giving emergency authorization use for many assay types. Such emergency steps are true in other countries, too.
As Luigi Basilio, applications specialist at Zymo Research points out, some emergency authorizations in the US for commercial manufacturers are for those who develop and distribute their own tests, of which Roche’s RT-PCR-based test--the cobas SARS-CoV-2 Test--is one example.
Then there are lab-developed tests, that CLIA labs--labs certified under the Clinical Laboratory Improvement Amendments (CLIA) of 1988-- are allowed to perform. And these labs may have assays of their own, which need to be authorized by federal or state authorities.
For commercial manufacturers to develop a product for research or clinical use, “they need to validate the entire workflow from collection, sample processing, and detection” he says. All deviation from that protocol requires additional validation and approval by the FDA. A limiting factor in all is supply, he says.
Smaller assays use lower amounts of reagents and consumables. But there is no way getting around the fact that, given the pandemic, for the many assays being run, disposable-type supplies and instruments are in short supply. “One thing we are worrying about is there is a shortage of lysis buffer and RT enzymes from certain suppliers, which is going to put a crimp on testing,” says Loman. Vandesompele echoes this concern. There is widespread back-ordering of chemicals, which include lysis reagents, binding reagents, washing reagents, and elution reagents.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in