Tiny tundra plants and consistent trait-environment relationships
Published in Ecology & Evolution
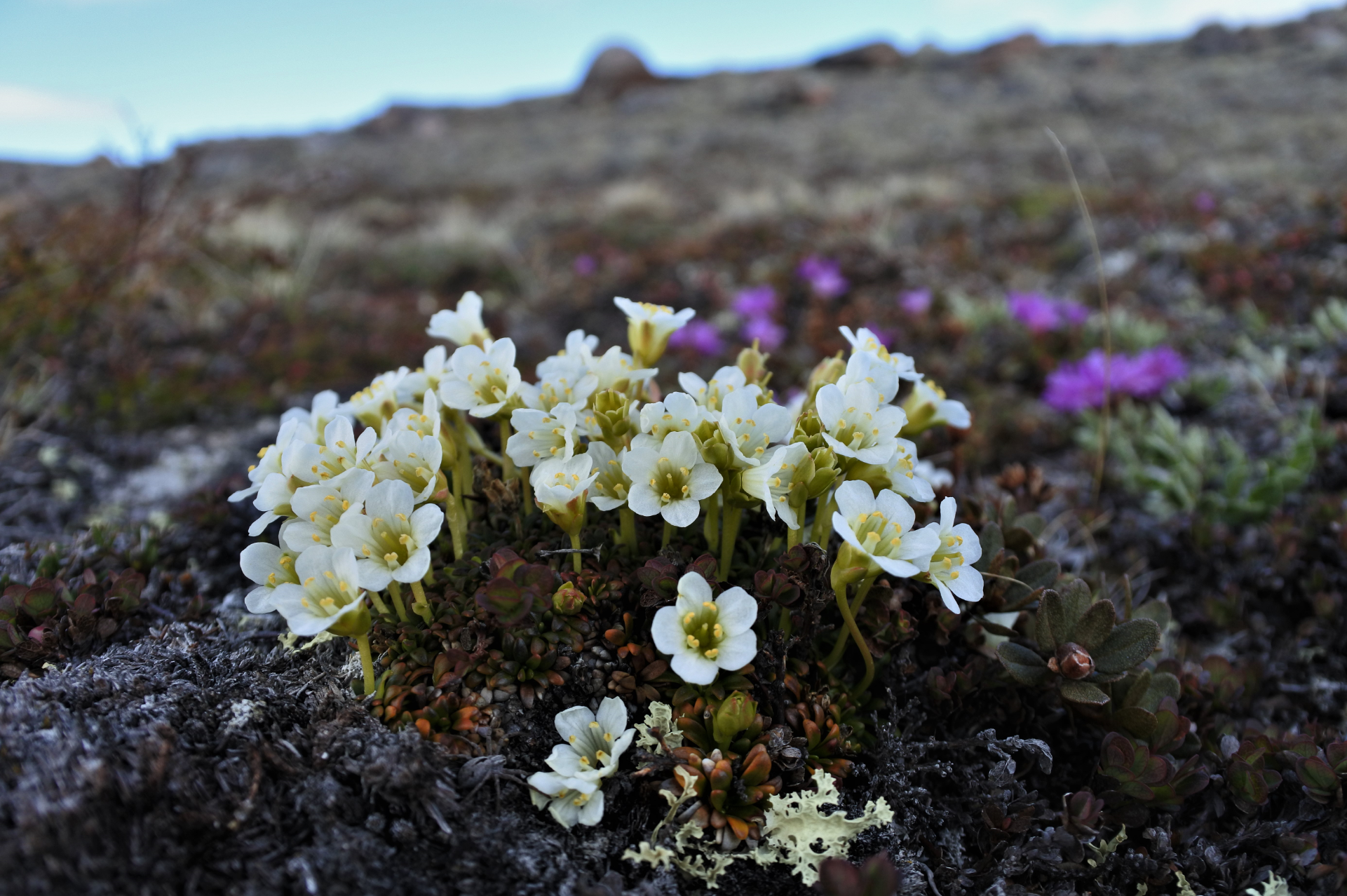

"See, the leaves are waxy and rounded", said Peter C. le Roux and explained me how to identify Vaccinium vitis-idaea among other tiny tundra plants. It was July 2013 and the weather was perfect for vegetation surveys on the slopes of Mount Saana, sub-Arctic Fennoscandia. It was my first day in the field and I fell for fells, flora and field work. I was a second-year geography student at the University of Helsinki, Finland, and I was working as a research assistant at the BiogeoClimate Modelling Lab led by Miska Luoto.
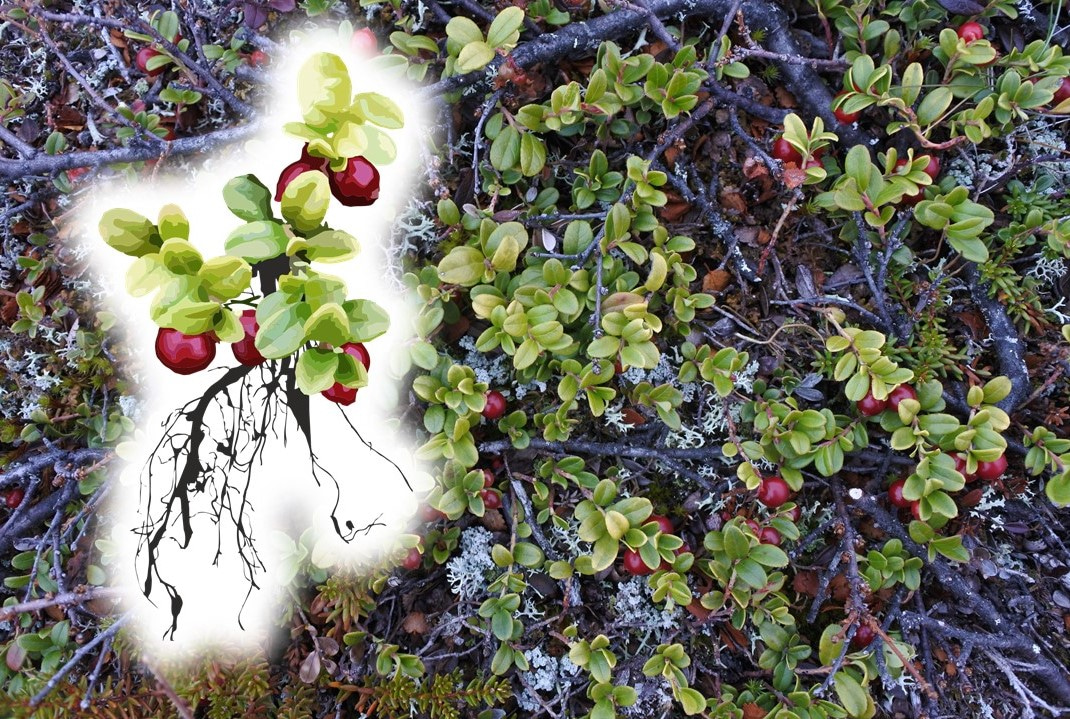

Couple of years prior Miska and Peter had established a vegetation survey methodology for maximising the breath of local environmental gradients that are sampled. The locations of the study grids are based on topographic and vegetation gradients. This means that each of the 21 grids at the sub-Arctic site captures local variation in mesotopography and also in vegetation cover and composition. In turn, each grid has range of microclimatic and soil chemistry conditions, and this is perfect for exploring fine-scale plant-environment relationships. One grid consists of 160 study plots, as the grid is 20 meters long and 8 meters wide, and this is perfect for high efficiency in the field.
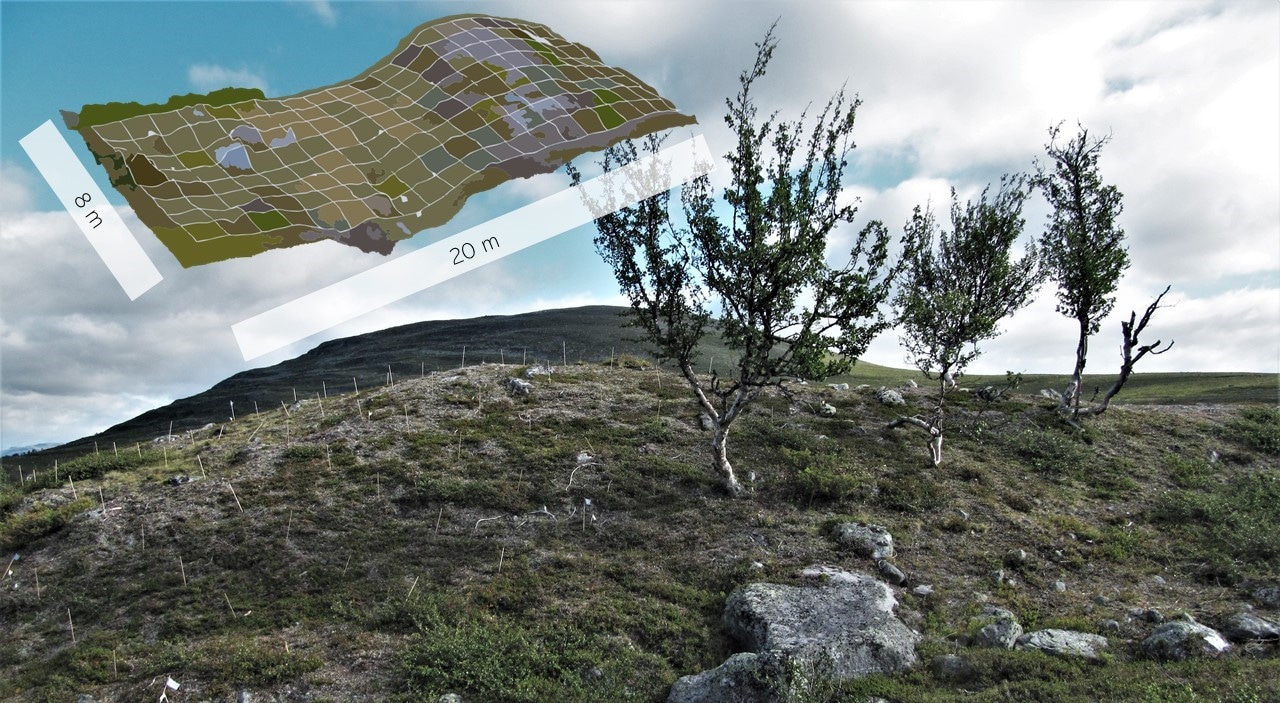

In 2013, we added more of these grids on Mount Saana and collected data on vascular plant species occurrences, soil moisture, soil temperature, slope, aspect and soil pH among many other things. Later that summer Peter, Juha Aalto and Miska published the grids in Global Change Biology (le Roux et al. 2013) and Peter returned back to the University of Pretoria, South Africa, and together with Mia Momberg established nine more grids on Marion Island, a sub-Antarctic island. For the grids, these were the opening shots for high-impact journals and comparing plant communities across tundra regions.
In summer 2018, me and Pekka Niittynen were touring the Arctic. We established six grids on the highland of Ammalortup Nunaa, low-Arctic Greenland and another six grids on the slopes of Adventdalen and Endalen valleys, high-Arctic Svalbard. I focused on identifying V. vitis-idaea and the rest of the vascular plants, while Pekka focused on mosses and lichens, on which we published in Nature Climate Change (Niittynen et al. 2020). On Svalbard, we also took the Plant functional traits course, on which Brian J. Enquist, Vigdis Vanvik, Aud H. Halbitter and Brian Maitner taught me that there is so much more to plants than just identifying their species.
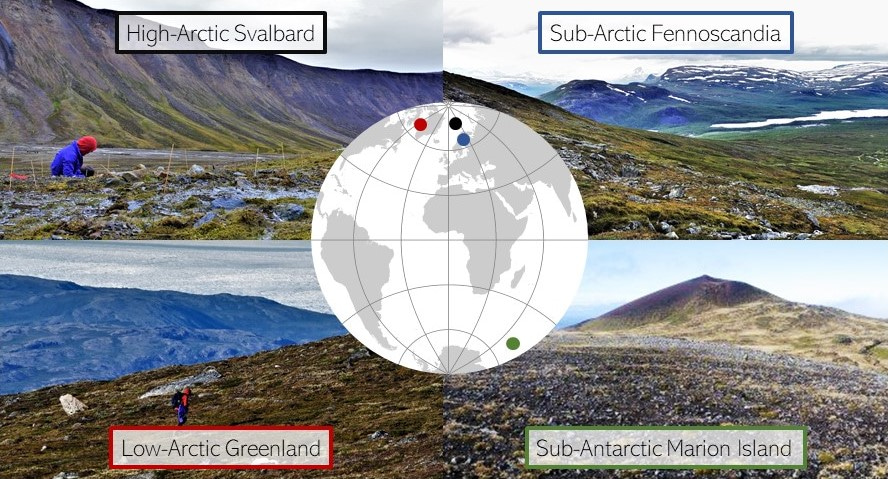
In autumn 2019, Helena Rautakoski had just finished soil pH analyses for the high-Arctic and low-Arctic sites, and I was exploring the data to find out if water as a resource, stress and disturbance influenced plant communities across the grids similarly as they did at the sub-Arctic site (Kemppinen et al. 2019). None of the 217 species were present on all four sites, but luckily plant functional traits provided a common currency. Pekka extracted over 76000 trait observations for seven plant functional traits for each species from three databases to calculate the community weighted means for the grid data (Kattge et al. 2011; Maitner et al. 2017; Bjorkman et al. 2018).
"I just read an inspirational article about hierarchical generalized additive models (Pedersen et al. 2019), you should try those" said my office mate Konsta Happonen. So we wrote a script for HGAM, and just to try it out we added "the basics" as explanatory variables: soil moisture, soil temperature, potential solar radiation and soil pH. I was hoping to find a general pattern between the different water aspects and traits. Turned out, there was a general pattern between all the explanatory variables and traits. It was a weird moment when we realized that we had just stumbled upon strong, consistent trait–environment relationships across Arctic and Antarctic tundra plant communities.
The manuscript was the last chapter of my PhD thesis (Kemppinen 2020) and a year later it was published in Nature Ecology and Evolution (Kemppinen et al. 2021) and also featured on the cover of the April issue. It was definitely a full circle moment for many years of hard work. Little did I know what a journey would come out of identifying those tiny tundra plants.

Kemppinen, Niittynen, le Roux, Momberg, Happonen, Aalto, Rautakoski, Enquist, Vandvik, Halbritter, Maitner & Luoto (2021). Consistent trait–environment relationships within and across tundra plant communities. Nature Ecology & Evolution.
Author contributions
JK conceived the research together with ML. JK, PN, PCLR, JA, and ML designed the study setting. JK, PN, PCLR, MM, JA, and ML performed the field research and HR laboratory analyses. JK, with support from KH and BJE, analysed the data. JK wrote the first version of the paper, with support, comments, and input from PN, PCLR, MM, KH, JA, HR, BJE, VV, AHH, BM, and ML. JK, PN, PCLR, MM, KH, JA, HR, BJE, VV, AHH, BM, and ML revised the paper based on peer review comments.
Acknowledgements
We thank the past and present members of the BioGeoClimate Modelling Lab and the le Roux lab for their hard work collecting the field data. We also thank the laboratory personnel at the University of Helsinki and University of Pretoria, as well as the staff at The University Centre in Svalbard, Kangerlussuaq International Support Services, Kilpisjärvi Biological research station, and Marion Island field assistants (specifically Elana Mostert, Nothando Mhlongo, Jenna van Berkel, and Janine Schoombie). We are also grateful to Pernille Eidesen for helping with the temperature loggers and Eric Pedersen for his help regarding HGAM. We thank Henri Riihimäki for providing the drone image composition, on which the study grid vector is based on in Figure 1. JK was funded by the Doctoral Programme in Geosciences at the University of Helsinki, PN by the Kone Foundation, MM by the National Research Foundation via the SANAP programme, and KH by the Doctoral Programme in Wildlife Biology Research at the University of Helsinki. The field campaigns were funded by the Academy of Finland (project numbers 307761 and 286950) and the National Research Foundation’s South African National Antarctic Program (unique grant numbers 93077 and 110726). We acknowledge the funding by the Finnish Ministry of Education and Culture (The FinCEAL Plus BRIDGES coordinated by the Finnish University Partnership for International Development).
Permissions
Permission to carry out fieldwork was granted by the Governor of Svalbard for the high-Arctic site, the Government of Greenland for the low-Arctic site, Metsähallitus for the sub-Arctic site, and the Prince Edward Islands Management Committee (permit PEIMC1/2013) for the sub-Antarctic site.
Follow the Topic
-
Nature Ecology & Evolution
This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in